The Burden of C. difficile and its Link to Nosocomial Infections: Difference between revisions
No edit summary |
|||
| Line 1: | Line 1: | ||
==Introduction== | ==Introduction== | ||
[[Image:C. difficile illustration.jpg|thumb|300px|right|Medical Illustration of C. difficile http://www.cdc.gov/hai/organisms/cdiff/Cdiff-patient.html]] | [[Image:C. difficile illustration.jpg|thumb|300px|right|Medical Illustration of C. difficile http://www.cdc.gov/hai/organisms/cdiff/Cdiff-patient.html]] | ||
<br>At right is a sample image insertion. It works for any image uploaded anywhere to MicrobeWiki. The insertion code consists of: | <br>At right is a sample image insertion. It works for any image uploaded anywhere to MicrobeWiki. The insertion code consists of: | ||
<br><b>Double brackets:</b> [[ | <br><b>Double brackets:</b> [[ | ||
| Line 24: | Line 19: | ||
<br> <br> | <br> <br> | ||
== | ==Types of HAI== | ||
<br>Include some current research in each topic, with at least one figure showing data.<br> | <br>Include some current research in each topic, with at least one figure showing data.<br> | ||
== | ==Clostridium difficile as a Leading Cause== | ||
<br>Include some current research in each topic, with at least one figure showing data.<br> | <br>Include some current research in each topic, with at least one figure showing data.<br> | ||
== | ==Toxin Production== | ||
<br>Include some current research in each topic, with at least one figure showing data. | |||
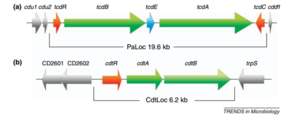
[[Image:PaLoc.png|thumb|300px|right|Genetic map of PaLoc and CDT locus in C. difficile. Arrows denote the direction of transcription. Toxin genes are shown in green, and regulator genes in red. | |||
http://ac.els-cdn.com/S0966842X1100196X/1-s2.0-S0966842X1100196X-main.pdf?_tid=77e5bb16-e7ad-11e4-8275-00000aab0f27&acdnat=1429569475_da85a2645d2046d4eb2278f4c889fe9a | |||
]]<br> | |||
==Strains and Identification== | |||
<br>Include some current research in each topic, with at least one figure showing data. | |||
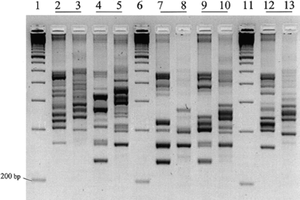
[[Image:PCR-ribotyping.png|thumb|300px|right|Comparison of PCR-ribotyping using newly developed primers that closely surround the rRNA operon, versus primers used in older literature. Lanes 1, 6, and 10 are 100-bp mass markers. Lanes 2, 4, 7, 9, and 12 show the results using new primers, next to Lanes 3, 5, 8, 10, and 13 which use primers from older literature (18). http://femsle.oxfordjournals.org/content/175/2/261.figures-only | |||
]]<br> | |||
==Treatment== | |||
<br>Include some current research in each topic, with at least one figure showing data.<br> | <br>Include some current research in each topic, with at least one figure showing data.<br> | ||
== | ==Prevention== | ||
<br>Overall paper length should be 3,000 words, with at least 3 figures.<br> | <br>Overall paper length should be 3,000 words, with at least 3 figures.<br> | ||
Revision as of 20:23, 22 April 2015
Introduction
At right is a sample image insertion. It works for any image uploaded anywhere to MicrobeWiki. The insertion code consists of:
Double brackets: [[
Filename: PHIL_1181_lores.jpg
Thumbnail status: |thumb|
Pixel size: |300px|
Placement on page: |right|
Legend/credit: Electron micrograph of the Ebola Zaire virus. This was the first photo ever taken of the virus, on 10/13/1976. By Dr. F.A. Murphy, now at U.C. Davis, then at the CDC.
Closed double brackets: ]]
Other examples:
Bold
Italic
Subscript: H2O
Superscript: Fe3+
Types of HAI
Include some current research in each topic, with at least one figure showing data.
Clostridium difficile as a Leading Cause
Include some current research in each topic, with at least one figure showing data.
Toxin Production
Include some current research in each topic, with at least one figure showing data.
Strains and Identification
Include some current research in each topic, with at least one figure showing data.
Treatment
Include some current research in each topic, with at least one figure showing data.
Prevention
Overall paper length should be 3,000 words, with at least 3 figures.
References
Edited by student of Joan Slonczewski for BIOL 238 Microbiology, 2009, Kenyon College.
