User:GillenK
Targeting of proteins to different cellular compartments in E. coli. From MicrobeWiki, the student-edited microbiology resource Jump to: navigation, search Contents [hide]
Introduction
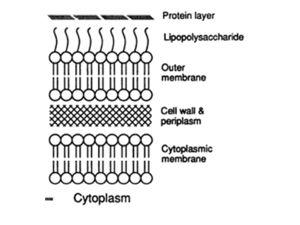
Sorting proteins to their correct cellular location is of critical importance to cells. Mis-localized proteins are non-functional and can lead to cell death. Protein sorting to the various organelles in eukaryotic cells is widely studied and resulted in a Nobel Prize to Blobel for the recognition of the signal sequence that initially targets secreted and membrane proteins to the endoplasmic reticulum. While it has been long assumed that prokaryotes lack the sophistical intracellular architecture of eukaryotic cells, more recent research has shown that prokaryotes do indeed have cytoplasmic organization with various proteins restricted to certain areas. In addition to the cytoplasm, Gram-negative bacteria must selectively localize proteins to the inner membrane (IM), periplasm, outer membrane (OM), and extracellular environment (Figure 1). This page will summarize how proteins are targeted to various compartments in Gram-negative cells, taking most examples from the well-studied Escherichia coli (E. coli). Beyond to contributing to basic biological research, studies of protein sorting in bacteria have biotechnology and medical applications (Cormelius, P., 2000). Purification of recombinant proteins in E. coli is aided by targeting the proteins to particular cell locations and studies of bacterial toxin secretion may lead to novel therapeutic agents.
Cytoplasm
Proteins with uniform cytoplasmic concentrations do not need specialized mechanisms for localization. Proteins are produced by ribosomes located in the cytoplasm and can diffuse away from their point of synthesis to reach homogenous cell concentrations. However, many cytoplasmic proteins do not exhibit uniform distribution and instead are restricted to particular subcellular regions. This restricted localization does require mechanisms to insure correct spatial distribution. Shapiro et al. (2009) reviewed the types of proteins that are targeted to specific cytoplasmic locations and discussed possible mechanisms for localization. For example, in E. coli FtsZ forms a ring at the midpoint of dividing cells due to the polar localization of an FtsZ polymerization inhibitor protein. Shapiro et al. document that protein localization is likely due to regions of the targeted proteins that form complexes with previously localized proteins in a process they term “diffuse and capture”. The initial protein localization may occur through spatial constraints of cell architecture. Recently, protein localization due to localization of bacterial mRNA has been reported. In an elegant study, Nevo- Dinur et al. (2011) found that the mRNA for a membrane protein BglF, was associated with membranes even without translation of the protein. mRNA localization in multicellular eukaryotes has been appreciated since the 1980’s (Holt and Bullock, 2009) but the localization of mRNAs in bacteria is an emerging field (Ranamurthi, K. 2011) with much to be discovered.
Inner Membrane
Many proteins are found at the Gram-negative inner membrane including lactose permease, ATP synthase and multidrug efflux pumps. The orientation of over one hundred inner membrane proteins has been described and is curated at the University of Michigan OPM database (http://opm.phar.umich.edu/localization.php?localization=Bacterial%20gram-negative%20inner%20membrane). Most inner membrane proteins localize to and insert in the inner membrane due to the presence of an amino terminus signal sequence that consists of amino acids with hydrophobic side chains. The signal sequence interacts with a signal recognition particle (SRP), a ribonucleoprotein that in E. coli is composed of the protein Ffh and a 4.5 S RNA. The SRP nascent protein ribosome complex associates with the SRP receptor (SR). FtsY is the E. coli SR. FtsY is homologous with a subunit of the eukaryotic SR that is found in the endoplasmic reticulum membrane (Angelini, et al. 2005). Ataide et al. (2011) recently reported the crystal structure of the SRP FtsY interaction using proteins from E. coli . Since the signal recognition particle-signal receptor interaction is conserved across domains, this structure will also provide insight into eukaryotic complexes. FtsY is located at the inner membrane due to interactions with the SecYEG translocon ((Angelini, et al. 2005). Using an energy source, proteins are translocated across the inner membrane. Integral inner membrane proteins contain long stretches of amino acids with hydrophobic side chains that will anchor the proteins in the membrane. While most inner membrane proteins take the route outlined above, there are examples of proteins inserting into the inner membrane without interacting with the signal recognition particle (de Gier, et al. 1998).
Periplasm
Include some current research, with at least one figure showing data.
Outer membrane
Include some current research, with at least one figure showing data. Conclusion
Extracellular
In addition to targeting to various cellular locations, some proteins are destined to leave the cell entirely and enter the extracellular environment. Examples include hemolysins exotoxins that lyse host cells to release nutrients and pili that mediate attachment to substrates. In gram-negative cells secreted proteins must cross both the inner and outer membranes. Secretion across the outer membranes differs from secretion across the inner membrane in a number of respects. In contrast to the IM Sec proteins through which proteins pass in an unfolded state, proteins may partially fold in the periplasm prior to secretion through the OM. (Thanassi and Hultgren 2000).
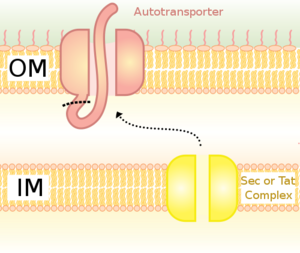
Thanassi and Hultgren (2000) reviewed the different pathways proteins may take on their voyage outside the Gram-negative cell. They discuss six different pathways to protein secretion, four of which are Sec dependent and two that are Sec independent. Proteins using the Sec dependent pathways, as the name suggests, cross the inner membrane via the Sec pathway described above. They then rely on different paths to extracellular secretion. The simplest path is taken by autotransporters, proteins that form their own pore in the OM. Autotransporter proteins contain three domains, the N-terminal signal sequence for Sec targeting, a carboxy-terminal beta-barrel that forms a pore in the OM, and a passenger domain. The passenger domain passes through the pore and can be cleaved to be released extracellularly. There is also a single accessory factor pathway that encodes the beta-barrel OM pore as a separate protein. The beta-barrels of the autotransporters and single accessory factors are not homologous and appear to have evolved separately. The chaperone/usher pathway is a second route outside the cell.
References
Angelini, et al. 2005. FtsY, the bacterial signal-recognition particle receptor, interacts functionally and physically with the SecYEG translocon. EMBO reports 6, 5, 476–481.
Ataide, S. et al. 2011. The Crystal Structure of the Signal Recognition Particle in Complex with its Receptor. Science 311: 881-885.
Cornelis, P. 2000. Expressing genes in different Escherichia coli compartments. Current Opinion in Biotechnology 2000;11(5):450-4
de Gier et al. 1998. Differential use of the signal recognition particle translocase targeting pathway for inner membrane protein assembly in Escherichia coli. PNAS 95: 14646-14651
Holt and Bullock. 2009. Subcellular mRNA Localization in Animal Cells and Why it Matters. Science 326: 1212-1215.
Nevo-Dinur, et al. 2011. Translation-Independent Localization of mRNA in E. coli. Science 331: 1081-1084.
Pugsley, A. 1993. The Complete General Secretory Pathway in Gram-Negative Bacteria. Microbiological Reviews. 57: 50-108
Ranamurthi, K. 2011. mRNA delivers the Goods. Science 331: 1021-1022.
Shapiro, L. et al. 2009. Why and How Bacteria Localize Proteins. Science 326: 1225- 1228.
Thanassi and Hultgren. 2000. Multiple pathways allow protein secretion across the bacterial outer membrane. Current Opinion in Cell Biology 12:420-430.
Edited by student of Joan Slonczewski for BIOL 238 Microbiology, 2010, Kenyon College.
Retrieved from "http://microbewiki.kenyon.edu/index.php/BIOL_238_Paper_2010"
Category: Pages edited by students of Joan Slonczewski at Kenyon College
