User:Liwoo92
Introduction
The use of antibiotics has become common in the livestock production around the world. The growth-promotic effects of antibiotics are undisputed, but the collateral and long-term effect are a cause for a heated debate and banning in the European Union in 2005(OMAFRA, 2005.) Antibiotics increase the efficiency of animal growth by inhibiting the growth of microbes in the gastrointestinal tract which triggers immune responses in the host (Gaskins et al., 2002). They have been shown to improve the health of animals raised in close quarters in conventional operations and also reduce microbes on the meat that cause foodborne illnesses (OMAFRA, 2005). However, there is much concern regarding the development of antibiotic resistance associated with the use of drugs. It is important to study the microbial system within the host organism to carry out further studies related to the controversy.
Host Microbial Community
All livestock harbor a intestinal microbes in a dense and highly diverse community, which are engaged in complex interactions with one another. Despite the diversity, specific animals have innate microbial communities. For example, in swine, major groups include Bacteroides, Peptostreptococcus, Bifidobacterium, Selenomonas, Clostridium, Butyrivibrio, and Escherichia (Moore et al., 1987). Most of the microbes are found in the large intestine because of slow digesta turnover. A low number of microbes occupy the small intestine because of low pH and the rapid digesta flow which results in bacterial washout (Gaskins, 2000). Gut flora benefits the host in a variety of ways including digestion of unutilized energy substrates, stimulating cell growth, repressing the growth of harmful microorganisms, training the immune system to respond to pathogens, and defending against some diseases (Guarner et al.,2003). The gut flora and the host form an important mutualistic relationship.
Immunological Interactions
The non-pathogenic microbes benefit their host by stimulating the development of immune responses. The host organism develops defensive responses such as the constant mucus production and high cell turnover of the GI tract (Gaskins et al., 2002). The immune responses result in bacterial washout, controlling the growth rate of the enteric bacteria (Gaskins et al., 2002). The washout leads to the prevention of the pathogen growth, defending against diseases. Also, indigenous bacteria are proposed to prevent the colonization of nonindigenous bacteria via competition for nutrients and mucosal attachment sites, or alteration of the growth environment by producing antimicrobial compounds and modified bile acids (Rolfe, 1997). Studies with germfree animals show that the absence of indigenous bacteria leads to an underdeveloped immune system and less effective response to pathogens (Gaskins, 2000). Thus, the normal intestinal microbes provide the host with important defense by outcompeting pathogenic bacteria and preventing enteric diseases (Nisbet, 1998).
Immunity vs. Growth efficiency
However, those innate immune responses are offered at the expense of the growth efficiency (Gaskins et al., 2002). Building the defense against the microbial community in the GI tract requires disproportionate amounts of energy and resources. For example, in swine, although the GI tissues only represent 5% of the total body weight, they receive 15 to 35% of the whole body oxygen consumption and protein turnover because of high metabolic rate (Gaskins, 2000). Germfree organisms do not have to develop such immune responses, and instead, they can make effective use of their energy by investing it in weight growth.
Efficacy and Mechanism of Antibiotics
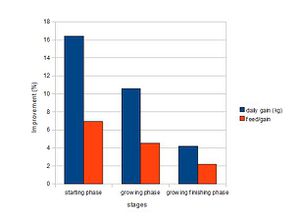
Numerous researches have shown that antibiotics increase the rate and efficiency of growth in animals. Figure 1 shows that antibiotics are effective in pigs of all growth stages, but they are the most effective when the pigs are young and weanling. The growth rate of pigs in starting phase (7-25kg) improved by an average of 16.4% and reduced the amount of feed required by 6.9%. However, the experiments are controlled and performed at clean and disease-free research facilities. Thus, the antibiotics are predicted to be more beneficial when used at the farm, where it is less clean, and thus, easier to catch diseases. (Cromwell, 2000). The exact mechanisms by which antibiotics favor growth are not known; however, researches propose that they possibly promote growth by depressing the growth of microbes that are toxic or steal nutrients from the host, leading to the increased nutrition utilization and reduced energy investment in maintaining immune responses in the GI tract (Gaskins et al., 2002).
Debate regarding Resistance
Since antibiotics have been introduced as growth promoters, there have been concerns about antimicrobial resistance. Frequent use of antibiotics leads to increasing resistance in enteric bacteria that can infect people or transfer their resistance to other pathogenic bacteria (OMAFRA, 2005). When infected by multi-drug resistant bacteria, treatment options are limited, recovery is slow, and treatment is more costly (WHO, 2012). The concerns especially involve resistance against penicillin and tetracycline because they are used in human medicine as well (Cromwell, 2000). Antibiotic resistance is controlled by decreasing the use of unnecessary antibiotics. For example, the use of antibiotics in the livestock and poultry industries is completely banned in the European Union (EU) (OMAFRA, 2005). However, there has been no concrete evidence that shows that use of antibiotics in animal food production poses a threat to human health, and studies are still under way to find the direct linkage (OMAFRA, 2005). The process involving the transfer of antibiotic resistance plasmid between animal and human bacteria is not well known yet (Cromwell, 2002). Also, there is a study which shows that the amount of antibiotic resistance transfer from resistant E.coli to E.coli in the GI tract is small, and animal strains are poor at colonizing the human intestinal tract (Smith, 1969). The question whether the use of antibiotics in animals presents a potential threat to human health will continue to be controversial.
Current Research
1. A study of the microbial community composition and functional genes in oil contaminated soil
In order to compare microbial functional diversity in different oil contaminated soils, and to find the relationship between the contamination and environmental factors, Liang et al. analyses soil samples from 5 different oil filed and used GeoChip to evaluate the microbial functional genes. Results showed that the samples were clustered by geographic locations and the contaminant degradations genes presented similar patterns under oil contaminant stress. Canonical analysis results also indicated that the local environmental variables significantly affect the microbial functional patterns (Liang et al. 2011).
2. A case study of the environmental condition after Exxon Valdez oil spill
In order to evaluate the long-term environmental effect of Exxon Valdez oil spill in 1989, a series of measurements of the background concentration of nutrients, dissolved oxygen (DO), and salinity were obtained from a contaminated beach. Results showed that both nutrients and DO are limiting factors for biodegradation. Also, the lowest nitrate and DO values were found in the oiled pits, implying that the microbial oil degradation was probably under anoxic conditions associated with denitrification (Boufadel et al. 2010).
3. The application of bioautmentation
The feasibility of a bioaugmentation strategy based on use of microbial formula tailored with selected native strains to remediate diesel contaminated site was assessed. The biodegradation process of diesel oil was assessed by monitoring the DO composition, CO2 evolution rate, microbial load and composition of the community by T-RFLP, physiological profile in Biolog® ECOplates and ecotoxicity. The mixture of microbes that combines 10 bacterial strains selected for resistance to heavy metals was found to efficiently facilated and speed up the bioremediation of diesel hydrocarbons and heavey metals (Alisi et al. 2009)
References
19. Prince R (1993) Petroleum spill bioremediation in marine environments. Crit Rev Microbiol 19:217-242
Edited by Yan Zhou, a student of Angela Kent at the University of Illinois at Urbana-Champaign.