Zinc Finger Nuclease (ZFN)
Introduction

Zinc finger nucleases (ZFNs) are proteins composed of a DNA binding-domain and a DNA cleaving-domain. One zinc “finger” binding domain recognizes and binds to a three nucleotide sequence, one can thus increase the binding specificity of a ZFN by adding more zinc fingers. The DNA cleaving-domain (Fok 1 nuclease) can then be used to create a double-stranded break in DNA at the desired point (Fig. 1).
After a double stranded break, DNA is repaired by non-homologous end joining. This often causes small insertions or deletions in the DNA, resulting frame-shift mutations. These frame-shifts result in nonsense mutations or nonsense-mediated decay. These effects are therefore very useful for creating knockout genes.
Additionally, genetic material such as DNA plasmids can be integrated into a gene after a double stranded break. A desired donor plasmid will contain arms of DNA which are homologous to the ZFN cut site. The donor plasmid and ZFNs can then be added simultaneously to the cell, the genetic material will be successfully incorporated after a double stranded cut and subsequent homologous recombination. Though homologous recombination is a rare event in most cells, double stranded breaks from ZFNs greatly increase the frequency of such events, facilitating gene insertion.
This makes ZFNs very useful tools for creating transgenic organisms and genetic engineering.
History
While other gene targeting procedures have existed for quite some time, the success of earlier methods were dependent on several factors including homologous recombination. Also, while genetic changes were observed in fungi like Saccharomyces cerevisiae and bacteria, most methods did not work on eukaryotic organisms. However, when studies in the 80s revealed that double-stranded DNA breaks greatly increased the frequency of recombination and thus successful genetic modifiactions, tools such as ZFNs became favored. Previously, stem cells were required 5.
Using ZFNs to create knockout cells
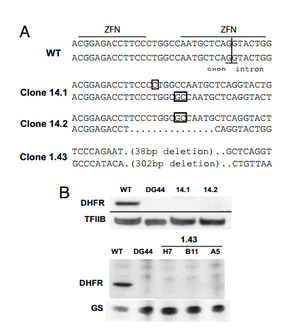
Santiago et. al tested the ability of ZFNs to create knockout mammalian cells. To create knockouts, the same principles described in the introduction were followed. That is, nonhomologous end joining imperfectly repairs the DNA after a double stranded break, leading to nonsense frame-shift mutations. The group knocked out the dihydrofolate reductase (DHFR) gene in the ovary cell line of a Chinese hamster (CHO).
A ZFN-based approach to knockout generation was found to be very feasible. The one-step technique much faster than other conventional methods. Double stranded breaks in DNA increase the frequency of homologous recombination by more than 1000. Thus, a high-frequency (2-3%) of mutants were reported after ZFN treatment. The knockout mutants could then be isolated out and cloned separately. When using methods where mutations are less frequent, the creation of knockout genes must be coupled with a selection method that selects for the low frequency mutants and fleshes them out. Because ZFNs directly affect the genome, gene knockouts are permanent and conserved in new generations. 6
Using ZFNs to insert genes into an existing genomic site
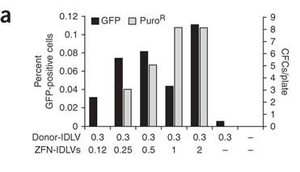
Lombardo et. al studied the insertion of genes into human stem cells after a ZFN mediated double stranded break. Lentiviral vectors were used to transport ZFNs and donor DNA to the targeted cells. This experiment had useful implications for curing genetic diseases in humans. Compared to older gene-replacement methods, it was discovered that gene-correction methods like those involving zinc finger nucleases and integrase-defective lentiviral vectors (ZFN-IDLVs) were safer and more effective for gene therapy. The group found that IDLVs successfully donate ZFNs and donor DNA to many type of human cells, and that once ZFNs and donor DNA have been supplied to the cell, insertion of the donor DNA into the desired site also occurred smoothly. 3
TALENs
Transcription activator-like effector nucleases (TALENs) are restriction enzymes comparable to ZFNs. Some believe that TALENs are emerging as the superior technology since they boast higher mutation frequencies and can bind to more sequences.
Conclusion
Proteins such as ZFNs and TALENs have massive potential in the world of genetic engineering and are already starting to see a lot of use. Their specificity and save a lot of time and effort, facilitating the research of transgenic organisms. These enzymes could potentially be used for gene therapy, combating pathogens, or making taller, faster growing crops.
