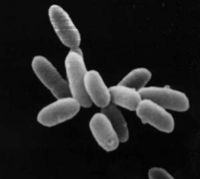
Halobacterium
A Microbial Biorealm page on the genus Halobacterium
See also Natronococcus
Classification
Higher order taxa:
Archaea; Euryarchaeota; Halobacteria; Halobacteriales; Halobacteriaceae
Species:
Halobacterium sp. NRC-1, H. salinarum
Description and Significance
Halobacteria are halophilic microorganisms, which means they grow in extremely high salinity environments. This archaeon can act as a good model for some aspects of eukaryotic biology, such as DNA replication, transcription, and translation. The major reason for this is that certain strains can be grown in laboratory conditions. Comparing a halophile genome to that of other prokaryotes should give insight into microbial adaptation to extreme conditions.
Genome Structure
The Halobacterium NRC-1 genome is 2,571,010 bp compiled into 3 circular replicons. More specifically, it is divided into one large chromosome with 2,014,239 bp and 2 small replicons pNRC100 (191,346 bp) and pNRC200 (365,425 bp). While much smaller than the large chromosome, the two plasmids account for most of the 91 insertion sequences and include genes for a DNA polymerase, seven transcription factors, genes in potassium and phosphate uptake, and cell division. The genome was discovered to contain a high G+C content at 67.9% on the large chromosome and 57.9% and 59.2% on the two plasmids. The genome also contained 91 insertion sequence elements comprised of 12 families, including 29 on pNRC100, 40 on pNRC200, and 22 on the large chromosome. This helps explain the genetic plasticity that has been observed in Halobacteria. Of the archaea, Halobacteria are viewed as being involved in the most lateral genetics (gene transfer between domains) and a proof that this transfer does take place.
Cell Structure and Metabolism
Halobacterium species are obligate aerobic, rod-shaped, archaea enveloped by a single lipid bilayer membrane surrounded by an S-layer made from the cell-surface glycoprotein. Halobacteria can grow on amino acids in aerobic conditions, however it has been found that they can also grow in an anaerobic environment given the correct conditions. These conditions are supported by amino acids such that arginine fermentation can occur. Although Halobacterium NRC-1 contains genes for glucose degradation as well as genes for enzymes of a fatty acid oxidation pathway, it does not seem able to use these as energy sources. Even though the cytoplasm retains an osmotic equilibrium with the hypersaline environment, the cell maintains a high potassium concentration. It does this by using many active transporters.
Many Halobacterium species possess proteinaceous organelles called gas vesicles. These vesicles are used to increase buoyancy and allow for taxis towards water-air interfaces. These bacteria are obligate aerobes that colonize water surfaces and harvest using membrane proteins, bacteriorhodopsin for metabolism. The gas vesicles, therefore, serve the function of allowing the Halobacterium to meet the obligate aerobe requirements.
Ecology
Halobacteria can be found in highly saline lakes such as the Great Salt Lake, the Dead Sea, and Lake Magadi. Halobacterium can be identified in bodies of water by the light-detecting pigment bacteriorhodopsin, which not only provides the archaeon with chemical energy, but gives it a reddish hue as well. An optimal temperature for growth has been observed at 37oC.
On an interesting note, however, Halobacteria are a candidate for a life form present on Mars. One of the problems associated with the survival on Mars is the destructive ultraviolet light. Halobacteria have an advantage here. These microorganisms develop a thin crust of salt that can moderate some of the ultraviolet light. Sodium chloride is the most common salt and chloride salts are opaque to short-wave ultraviolet. Their photosynthetic pigment, bacteriorhodopsin, is actually opaque to the longer wavelength ultraviolet (its red color). The obstacle Halobacteria would need to overcome is being able to grow at a low temperature during a presumably short time span when a pool of water could be liquid.
References
- Ng et al. 2000. Genome sequence of Halobacterium species NRC-1. PNAS vol 92, no 22; 12176-12181
- Walsby, A. E. (1994). Gas vesicles. Microbiol. Mol. Biol. Rev., 58(1), 94-144. Retrieved December 4, 2008, from http://mmbr.asm.org/cgi/content/abstract/58/1/94.
- Landis, Geoffrey A. 2001. Martian Water: Are There Extant Halobacteria on Mars?. Astrobiology vol. 1 no. 2: 161-164.
- Koonin, E.V., K.S. Makarova, and L. Aravind. 2001. Horizontal Gene Transfer in Prokaryotes: Quantification and Classification. Annu. Rev. Microbiol. 55:709-42.
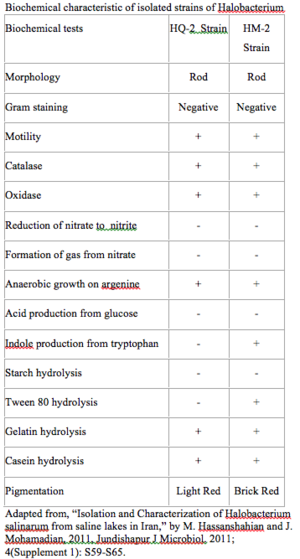
- Hassanshahian M, Mohamadian J. Isolation and characterization of Halobacterium salinarum from saline lakes in Iran. Jundishapur J Microbiol. 2011; 4(Supplement 1): S59-S65.