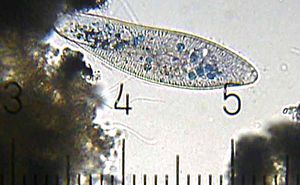
Paramecium
A Microbial Biorealm page on the genus Paramecium
Classification
Higher order taxa
Eukaryota; Alveolata; Ciliophora; Oligohymenophorea; Peniculida
Species
Paramecium aurelia
Paramecium polycarum
Paramecium woodruffi
|
NCBI: Taxonomy P. aurelia mitochondrion genome P. tetraurelia macronuclear genome |
Description and Significance
Paramecium are members of the phylum Ciliophora. They share many common characteristics with the rest of their phylum, but are also unique. For example, their shape is quite different from that of many other Ciliophora. They are also famous for their predator-prey relationship with Didinium. Paramecium are known for their avoidance behavior. If an encounters a negative stimiulus, it is capable of rotating up to 360 degrees to find an escape route.
Genome Structure
Macronuclear DNA in Paramecium has a very high gene density. The macronucleus can contain up to 800 copies of each gene.
Research on the genome structure of Paramecium is still largely incomplete. However, the genomes of some species are beginning to be sequenced. For example, the complete mitochondrion genome for Paramecium aurelia has been established. The complete macronuclear genome of Paramecium tetraurelia has also been sequenced.
Cell Structure and Metabolism
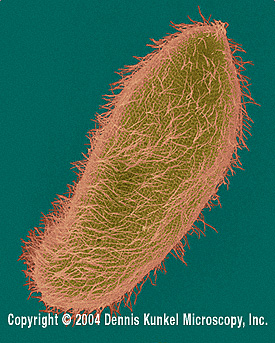
Paramecium are ciliated unicellular organisms. The cilia cover the entire body. Like other ciliates, they are multinucleated. Paramecium may eject trichocyts when they detect food, in order to better capture their prey. These trichocyts are filled with proteins. Trichocysts can also be deployed for self-defense.
Paramecium are heterotrophs. Their common form of prey is bacteria. A single organism has the ability to eat 5,000 bacteria a day. They are also known to feed on yeasts, algae, and small protozoa. Paramecium capture their prey through phagocytosis.
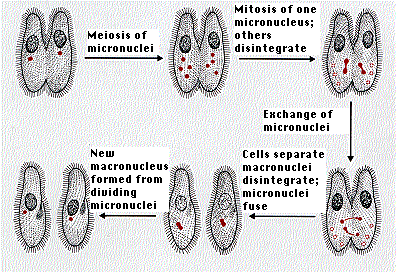
Paramecium are capable of both sexual and asexual reproduction. Asexual reproduction is the most common, and this is accomplished by the organism dividing transversely. The macronucleus elongates and splits. Under ideal conditions, Paramecium can reproduce asexually two or three tiems a day. Normally, Paramecium only reproduce sexually under stressful conditions. This occurs via conjugation, a process of gamete agglutination and fusion. Two Paramecium join together, forming a conjugation bridge. Each Paramecium has a diploid (2n) micronucleus that undergoes meiosis creating four haploid (1n) micronuclei Three of the resulting nuceli disintegrate, the fourth undergoes mitosis. Daughter nuclei fuse and the cells separate. The old macronucleus disintegrates and a new one is formed. This process is usually followed by asexual reproduction.
Ecology
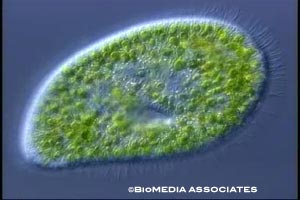
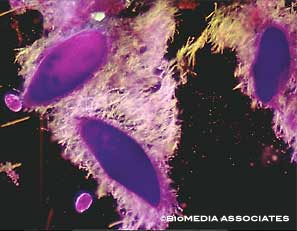
Paramecium live in aquatic environments, usually in stagnant, warm water. The species Paramecium bursaria forms symbiotic relationships with green algae. The algae live in its cytoplasm. Algal photosynthesis provides a food source for Paramecium. Some species form relationships with bacteria. For example, Paramecium caudatum hosts Holospora obtusa in its macronucleus. This bacteria is specific to the macronucleus of Paramecium caudatum; they cannot grow outside of this organism. This species acquires heat-shock resistance when infected with Holospora obtusa, which contributes to ciliary motion. Paramecium are also well known as prey for Didinium.
Paramecia play a role in the carbon cycle because the bacteria they eat are often found on decaying plants. Paramecium will eat the decaying plant matter in addition to the bacteria, further aiding decomposition.
Paramecia can be used as model organisms in research. Currently, they are being used a great deal in genetics research. For example, recent research involves inactivating Paramecium genes for studying functional analysis by homology-dependent gene silencing. They can also be used to study membrane excitability and the duplication of basal bodies.
References
BioMedia. "The Classics of Biology: Paramecium." 2004. Accessed 21 June 2005.
Haselton, Aaron. "Paramecium putrinum." The Connecticut River Homepage. Accessed 21 June 2005.
Kimball, John W. Ciliated Protozoans. 14 June 2003. Accessed 21 June 2005.
Samworth, Mike. "Paramecium." Microscopy UK. 1999. Accessed 21 June 2005.
Sperling, Linda. Paramecium Genomics. 17 April 2005. Accessed 21 June 2005.