Sphingomonas
A Microbial Biorealm page on the genus Sphingomonas
Classification
Higher order taxa:
Bacteria; Proteobacteria; Alphaproteobacteria; Sphingomonadales; Sphingomonadaceae
Species:
Sphingomonas abaci, S. adhaesiva, S. aerolata, S. aquatilis, S. asaccharolytica, S. aurantiaca, S. azotifigens, S. cloacae, S. dokdonensis, S. echinoides, S. elodea, S. faeni, S. genosp. 1, S. genosp. 2, S. koreensis, S. mali, S. melonis, S. mucosissima, S. oligophenolica, S. panni, S. parapaucimobilis, S. paucimobilis, S. phyllosphaerae, S. pituitosa, S. pruni, S. rhizogenes, S. roseiflava, S. sanguinis, S. suberifaciens, S. taejonensis, S. trueperi, S. ursincola, S. wittichii, S. xenophaga, S. yabuuchiae, S. yunnanensis, Sphingomonas sp.
Description and Significance
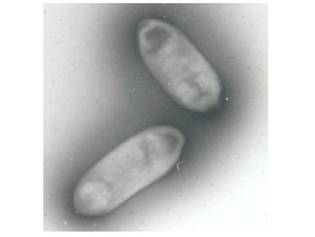
Sphingomonas is a Gram-negative, nonsporeforming, chemoheterotrophic, strictly aerobic bacterium that typically produces yellow- or off-white-pigmented colonies. It is unique in its possession of ubiquinone 10 as its major respiratory quinone, and of glycosphingolipids (GDLs) instead of lipopolysaccharide in their cell envelopes. The bacterium is metabolically versatile, which means it can utilize a wide range of naturally occuring compounds as well as some types of environmental contaminants. Therefore, studies have been held to further explore its metabolic mechanisms for more application in biotechnology, in addition to its current utilization in biomeditation and in the food technology. Also some species of Sphingomonas (especially S. paucimobilis) cause minor infection in humans and damage some other organisms.
Genome Structure
Sphingomonas is distinguishable from the other three genera (Sphingobium, Novosphingobium, and Sphingopyxis) by its lipid composition (major fatty acids [18:1, 16:1, and/or 17:1] and major 2-OH fatty acids [2-OH 14:0 or 2-OH 15:0]), utilization of homospermidine as the major polyamine, and signature nucleotide bases at certain positions within the 16S rRNA gene (Balkwill et al). Its genome sequence (incomplete) has about 3,948,000 Bp and 70 structural RNAs. Also, it contains 3914 proteins.
Cell Structure and Metabolism
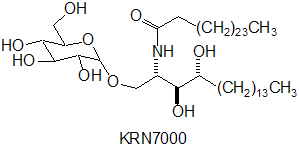
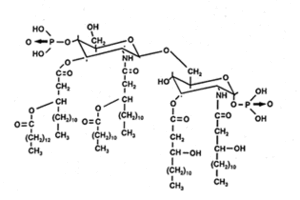
Sphingomonas is a rod-shaped bacterium (0.3-0.8 * 1.0-2.7µm) that has yellow or white color. Sphingomonas completely lacks lipopolysaccharides (LPS) that can carry endotoxins, which makes the bacterium almost unique among Gram-negative bacteria. Instead, it has a cell membrane that consists of proteins, phospholipids, and respiratory quinones and an outer membrane that contains glycosphingolipids (GSLs) (Balkwill et al). The GSLs occupy a position similar to that of LPS in other Gram-negative bacteria and appear to have many of the same functions, such as a barrier to batericidal substances. Also, because the carbohydrate part of a GSL is shorter than that of LPS, the cell surface of Sphingomonas is more hydrophobic than that of other Gram-negative bacteria. This is the probable reason for the bacterium's ability to degrade hydrophobic polycyclic aromatic hydrocarbons and its susceptibility to hydrophobic antibiotics.
Glucose is primarily utilized for growth, but a wide variety of other sugars such as arabinose, fucose, glactose, lactose, mannose, melibiose, sucrose, trehalose, and xylose also are frequently assimilated. Also, not only it can degrade mono- and disaccharides, Sphigomonas can also degrade polysaccharides. In addition, many of the strains discovered in contaminated environments were able to utilize one or more of the contaminants as their carbon sources, which demonstrated the versatility of the organism.
Ecology
Sphingomonas has been found in various environments, including both aqueous (both fresh- and seawater) and terrestrial habitals, plant root systems, clinical specimens, and others. Its widespread distribution in the environment is due to its ability to utilize a wide range of organic compounds and to grow and survive under low-nutrient conditions. Although many strains of Sphingomonas have been isolated from relatively clean environments, a great portion of them also had been discovered in contaminated locations which included toxic compounds such as PCBs, creosote, pentachlorophenol, herbicides, etc. In one study by Nishiyama et al, a strain of S. paucimobilis that degrades hexachlorocylohexane survived and grew in contaminated soil, especially inside of soil aggregates less than 0.025 mm in diameter, despite its surviving poorly in an unamended control soil. Also, the bacterium maintained higher population densities in the presence of the contaminant than in its absence. These discoveries demonstrate that Sphingomonas can utilize contaminants as a growth and energy source and that it can compete successfully with the indigenous organisms in various environments.
Pathology
Some species of Sphingomonas, especially S. paucimobilis, are found in equipment in hospitals and various types of clinical specimens, including respiratory therapy items, humidifiers, water, air, bedside water bottles, sinks, and temperature probes. Although infections by S. paucimobilis are rarely serious and could be effectively treated with antibiotics, it is capable of causing active infections in humans. For example, in 1991, at least 18 cases of infections caused by the bacterium were reported, which included six cases of bacteremia (bacterial infections of the bloodstream), two cases of leg ulcers, four cases of peritonitis (in patients undergoing chronic ambulatory dialysis), a case of brain abscess, a case of cervical adenopathy, a case of splenic abscess, a case of respiratory infection, a case of urinary infection, and a case of meningitis (Balkwill et al).

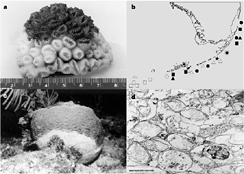
Also, an unidentified species of Sphingomonas is held responsible for coral plague. It was first discovered on the Florida Reef Tract in June, 1995, and is was reported that the star coral showed an unusually rapid rate of tissue degredation (up to 2 centimeters per day). The symptoms included a sharp line between healthy and diseased tissue, which was found in a wider variety of species than the other typical diseases (Richarson et al). It was further reported that, within four months of its initial discovery, the disease had spead about 200 km along the Florida Keys, and the disease continued to spread until around 1997 when it gradually disappeared.
In addition, recent study conducted by Scott T. Kelley et al (2004) showed that Sphingomonas is present in the shower curtain biofilms. The study stated that the "soap scum" of shower curtains is in fact a lush bed of micorbes, full of bacteria that are frequently isolated in wet environments. The authors of the report concluded that shower curtains can threaten immune-compromised or ill patients, and that consistent exposure to source of infection such as shower curtains can lead to health problems (Kelley et al).
References
M. Nishiyama, K. Senoo, H. Wada, and S. Matsumoto; 1992. "Identification of soil micro-habitats for growth, death and survival of a bacterium, g-1,2,3,4,5,6-hexachlorocyclohexane-assimilating Sphingomonas paucimobilis, by fractionation of soil" FEMS Microbiol. Ecol., vol. 101, pp. 145–150
