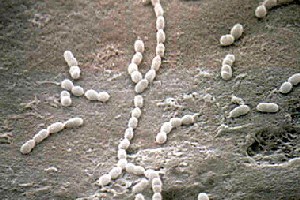
Streptococcus thermophilus
Classification
Higher order taxa:
Kingdom: Bacteria
Phylum: Firmicutes
Class: Bacilli
Order: Lactobacillales
Family: Streptococcaceae
Genus: Streptococcus
Species: Streptococcus thermophilus
|
NCBI: Taxonomy |
Description and Significance
Streptococcus thermophilus is an essential lactic acid bacterium used for commercial purposes, which includes the production of milk, cheese, and other dairy products. This organism is a thermophilic Gram-positive bacterium with an optimal growth rate at 45 °C. It is also capable of generating energy, in the form of adenosine triphosphate (ATP), by aerobic respiration with the presence of oxygen; however, without the presence of oxygen, it still can produce ATP through fermentation. S. thermophilus lacks cytochrome, oxidase, and catalase enzymes. It does not have motility and it does not form spores. Although S. thermophilus is closely related to other pathogenic streptococci (such as S. pneumoniae and S. pyogenes), S. thermophilus is classified as a non-pathogenic, alpha-hemolytic species that is part of the viridians group. The increasing consumer need for dairy products and booming manufacture of dairy products ($40 billion industry) led to the investigation and sequencing of S. thermophilus.
Genome Structure
Originally isolated for yogurt in 1974, the entire genome sequence of Streptococcus thermophilus LMG18311 has been sequenced[9]. The sequencing of S. thermophilus LMG18311 was accomplished by a genome shotgun method, which was consummated with the joint efforts of Biochemistry Unit and Genetics Unit from Université Catholique de Louvain. Also, a different strain, CNRZ1066, of S. thermophilus was sequenced.
The genomes of S. thermophilus CNRZ1066 and LMG18311 are approximately 1,800,000 base pairs in length and contain about 1900 coding regions. From these 1900 coding sequences, 1500 sequences are orthologus, genes from different species that are similar to each other, to various streptococcus genes, revealing that S. thermophilus share common physical and cellular properties with pathogenic streptococci. Both strains contain single circular chromosomes; however, they differ by 3,000 single nucleotide base pairs. The difference in genome sequence may account for difference in extracellular polysaccharide biosynthesis, bacteriocin synthesis, and immunities. S. thermophilus genes have low GC nucleotide base pairs (approximately 39%).
By sequencing the S. thermophilus genome, factors (which explain the non-pathogenic characteristics of S. thermophilus) have been observed. Nearly 10% of S. thermophilus genes are inactive or pseudogenes, which are caused by frame-shift, deletion, and or mutation of genes. Many virulent related genes (VRGs) that contribute to virulence of pathogenic streptococci are either not present or present as pseudogenes in S. thermophilus. [Virulence determinants such as pnemococcal surface protein A and C (PspA and C), pnemococcal manganese ABC transporter lipoprotein PsaA, IgA proteases, and choline binding proteins are inactivated in S. thermophilus.]
Cell Structure and Metabolism
Streptococcus thermophilus is a gram-positive bacterium; the cell wall is composed of N-acetylglucosamine (NAG) and N-acetylmuranic acid (NAM), which is bond by ether bonds. This unique structure allows S. thermophilus to endure elevated temperatures, which is useful for many industrial dairy fermentations requiring the process of milk at higher temperatures. It is non-motile, consists of coccus (circular) cell structure, produces no spores, and lives in pairs or chains (size of chain varies). S. thermophilus is a facultative anaerobe, which is an organism that is capable of generating ATP through both aerobic respiration and fermentation (depending on the presence or absence of oxygen). Furthermore, S. thermophilus lacks genes or contain pseudogenes expressing surface protein (excluding lipoproteins); pathogenic streptococci use these surface proteins to adhere to mucosal surfaces and evade host defense mechanisms. [Surface protein such as sortase-anchored surface proteins, an important virulence factor of pathogenic streptococci, is not present on the surface of S. thermophilus.]
The gene regions (epsE, epsF, epsG, and epsI) are responsible for synthesizing exopolysaccharide (EPS) in Streptococcus thermophilus, which consists of many functions that include: adhering to environmental stimuli, cleaving proteins (proteolysis), giving rise to pathogenic characteristics (excluding S. thermophilus), and etc[4]. EpsE encodes for phosphogalactosyltransferase, which is responsible for the translocation of galactose onto a lipophilic carrier; this initiates the synthesis of EPS. Gene epsG catalyzes the transference of α-N-acetylgalactosamine to β-galactoside. Genes epsF and epsI encode for galactosyltransferase and glycosyltransferase, respectively; these genes finalize the biosynthesis of EPS.
Lactose Transport:
Lactose is the primary source of carbon and energy for many lactic acid bacteria, including S. thermophilus. S. thermophilus uses a unique symport called lactose transporter LacS, which utilizes proton-gradient to collocate galactoside inside the cell[3]. The accumulation of galactose promotes a proton-neutral exchange of lactose for galactose; lactose is then metabolized and used for energy. The LacS transporter system is highly regulated by transcriptional and translocation activity of the cell, which requires lactose uptake and metabolism. LacS transporter consists of two parts: a carrier domain (embedded in the membrane) and a soluble IIAGlc domain (attached to the carrier domain via a flexible linker). This LacS-IIA complex is regulated by the phosphoenolpyruvate-phosphotransferase (PEP-PTS) system, which phosphorylates the IIA domain[5]. Phosphorylation of IIA domain activates the carrier domain, allowing the transport of galactoside and metabolism of lactose.
Proteolytic System:
The proteolytic mechanism degrades proteins by enzymes called proteases. S. thermophilus contains two unique peptidases, oligopeptidase and aminopeptidase PepS, which have multiple functions in bacterial growth[2]. Aminopeptidases are unique enzymes that function as exopeptidases, enzymes that catalyze the cleavage of specific amino acids from the end of a polypeptide. These essential enzymes incorporate many different functions, involving proliferation of cells: protein maturation, hydrolysis of regulatory proteins, regulation of gene expression, metabolism of nitrogen, etc. Animopeptidase PepS is a monomeric metallopeptadise that haz high affinity towards peptides with arginine or aromatic amino acids on the N-terminus of the peptide. The food industry uses lactic acid bacteria, such as S. thermophilus, for their proteolytic ability to grow and hydrolyze milk proteins (casein).
Ecology
S. thermophilus has become an essential bacterium for fermentation and production of dairy products over the past few years. The $40 billion food industry utilities the bacterium’s ability to hydrolyze milk proteins, casein, into nitrogen compounds. Enzymes in S. thermophilus, such as aminopeptidase, are used as starter cultures for food fermentation[6]. Exopolysaccharides created by these peptidases are essential to generate the texture of fermented milk products and organoleptic properties of dairy products. Food industries have high demands for an improved S. thermophilus strain, which can provide stable fermentation qualities, resilience to bacteriophage, and consistent flavor and texture of dairy products.
Research
“Application of Exopolysaccharide-Producing Cultures in Reduced-Fat Cheddar Cheese: Composition and Proteolysis”
Studies have shown that exopolysaccharide (EPS) producing bacteria are essential to produce reduced-fat dairy products with similar characteristics of regular (full-fat) products[7]. In this experiment, two different strains of EPS-producing bacteria were used to produce reduced-fat cheddar cheese: a “ropy” strain of Lactococcus lactis ssp. cremoris, a “capsule-forming non-ropy” and “ropy” strains of Streptococcus thermophilus. These bacteria have unique proteolysis abilities that allow them to breakdown protein, casein, in dairy products such as cheese into small peptides and amino acids. The small peptides and amino acids are crucial for the maturation of the texture and flavor of reduced-fat cheddar cheese. However, cheese made with Lactococcus lactis ssp. cremoris and Streptococcus thermophilus yield various types of cheeses.
Lactococcus lactis is shown to produce cheese with higher moisture levels compared to other reduced-fat cheeses. The high moisture level is due to the high chymosin activity induced by Lactococcus lactis. Chymosin is a protease enzyme that cleaves specific peptide bond, mainly between phenylalanine (Phe) and methionine (Met). In dairy products, casein proteins are linked with hydrophobic (para-casein group) and hydrophilic (acidic glycopeptide group) that is bound by Phe and Met. Cleavage of the peptide bond allows hydrophobic groups to bind to each other, forming precipitation and curd in cheese. The accumulation of hydrophobic peptides also gives the bitterness flavor of cheddar cheese. On the other hand, S. thermophilus produces low moisture cheddar cheese due to its low chymosin activity. Its ability to hydrolyze hydrophobic peptides decreases the bitterness in cheese. Therefore, applying both bacteria yields higher quality reduced-fat cheddar cheese with similar characteristics of regular, full-fat cheddar cheese.
References
- Bolotin, Quinquis, Renault, Sorokin. "Complete sequence and comparative genome analysis of the dairy bacterium Streptococcus thermophilus." Nature Biotechnology 22, 1554 – 1558. November 14 2004.
- Fernandez-Espla, Maria Dolores and Rul, Françoise. "PepS from Streptococcus thermophilus: A new member of the aminopeptidase T family of thermophilic bacteria." Eur. J. Biochem. 263, 502 – 510. FEBS 1999.
- Foucaud, C and Poolman, B. "Lactose transport system of Streptococcus thermophilus. Functional reconstitution of the protein and characterization of the kinetic mechanism of transport." J. Biol. Chem., Vol. 267, Issue 31, 22087-22094, 11, 1992.
- Francesca Stingele, John W. Newell, and Jean-Richard Neeser. "Unraveling the Function of Glycosyltransferases in Streptococcus thermophilus Sfi6." J Bacteriol. 1999 October; 181(20): 6354–6360.
- Geertsma, Duurkens, Poolman. "The Lactose Transport from Streptococcus thermophilus is Activated by Phosphorylated IIA." EU-FP6, E-Mep; 504601.
- Hutkins, Robert. Streptococcus thermophilus LMD-9. JGI Microbes. September 30, 2002.
- S Awad, AN Hassan, K Muthukumarappan. "Application of Exopolysaccharide-Producing Cultures in Reduced-Fat Cheddar Cheese: Composition and Proteolysis." Journal of Dairy Science, 2005 - Am Dairy Sci Assoc. Page 1. J. Dairy Sci. 88:4204–4213 American Dairy Science Association, 2005.
- Simonson, Lloyd G. Streptococcus thermophilus CNRZ1066. NCBI: Genome Project.
- Université Catholique de Louvain. Sequencing of Streptococcus thermophilus LMG18311. June 20, 2005.
KMG

