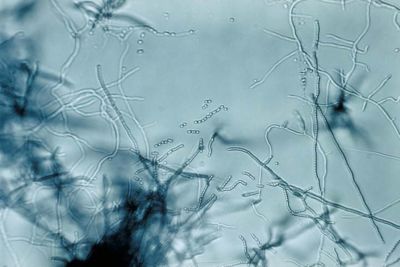
Streptomyces lividans
A Microbial Biorealm page on the genus Streptomyces lividans
Classification
Domain: Bacteria
Phylum: Actinobacteria
Class: Actinobacteria
Subclass: Actinobacteridae
Order: Actinomycetales
Suborder: Streptomycineae
Family: Streptomycetaceae
Strains: Streptomyces lividans
Species
|
NCBI: Taxonomy |
Genus species: Streptomyces lividans
Description and significance
Streptomyces lividans is a gram-positive, filamentous, soil bacterium. With its high G-C content, this species is known for its genomic plasticity, which consists of large deletions and extensive amplifications (1). Responsible for the breakdown of organic soil material, Streptomyces are non-pathogenic bacteria that live in colonies. Because of their natural habitat, along with the non-pathogenic character of the species in the genus, S. lividans is used as hosts for the synthesis and secretion of homologous and heterologous proteins (3). Bacteria belonging to the genus Streptomyces have mycelial growth, undergoing a complex biochemical and morphological change before forming exospore chains (2). In short, the complex life cycle of the genus comprises of the germination of spores to form a branched, vegetative mycelium, which then produces an aerial mycelium, and finally, spores (1). The Streptomyces lividans has been extensively studied in laboratories for possible therapeutic uses because of its abundance in multi-drug resistant genes. The Streptomyces genus is responsible for producing a majority of the antibiotics, some immunosuppressants, and anti-tumor agents used today (9).
Genome structure
Streptomyces lividans has a linear structure, and compared to Streptomyces coelicolor, they have common structures with identical ordering of the cosmid sequence (1). The central core region contains the essential genes, while the arms carry conditionally adaptive genes and species-specific DNA (13). Present in multiple copies in the vegetative and aerial mycelium, the chromosome is apparent only as a single copy in the spores. Physical map of the S. lividans genome was done by using restriction endonucleases AseI and DraI, cutting the linear chromosome into 16 and 7 fragments, respectively. The 8000 kb or so linear genome has an approximate 70% G+C content (1). Unlike most eubacteria, Streptomyces species usually contain linear chromosomes and plasmids. The linear plasmids are 12-1700 kb long. With inverted repeat sequences in their telomeres, replication of Streptomyces linear plasmids starts at centrally located loci and continues bidirectionally towards the telomeres. This leaves a single strand overhand of approximately 280 nt. as an intermediate. However, when the telomeres are deleted, the centrally located loci of Streptomyces linear plasmids can also maintain the circular mode. The replication modes and functions of different Streptomyces species has been highly conserved (7).
While replication modes remain conserved within the Streptomyces genus, there are considerable phenotypic variations observed across many genera of actinomycetes. Because of the high frequency of deletions, insertions, amplifications, and rearrangements, there are high levels of chromosomal instability. Also, horizontal gene transfer account for further genome instability of the genome (10).
Cell structure and metabolism
Gram-positive bacteria of the genus Streptomyces, S. lividans has a linear structure and undergoes dramatic changes in its life cycle. In short, the complex life cycle of the genus comprises of the germination of spores to form a branched, vegetative mycelium, which then produces an aerial mycelium, and finally, spores (1). The life cycle of Streptomycyes resembles that of fungi. In the vegetative growth stage, DNA replication occurs without cellular division. This creates the filamentous structure. Following the period of vegetative growth is the formation of spores, called conidia, which are produced in aerial filaments called sporophores.
Perhaps the most significant characteristic of the genus Streptomyces is that they produce and secrete large quantities of proteins. Often used as a host for secretory production of heterologous proteins, S. lividans has more than one gene for type 1 signal peptidases, 4 of which are located chromosomally. Transcription of S. lividans signal peptidase genes occur in the shape of polycistronic messengers. Their ability to export makes them a strong candidate for the study of drug resistant antibiotics.
Ecology
The major contribution of the Streptomyces genome is for the environment. Living in soil, they interact with various terrestrial environments. Certain strains such as Streptomyces scabies and Streptomyces ipomoeae effect vegetables (9, 14, 15).
Pathology
Streptomyces are non-pathogenic so it does not cause disease in animals, plants, and humans, although some species are plan pathogens. For example, Streptomyces scabies causes potato scabs and Streptomyces ipomoeae causes disease in sweet potatoes (14, 15).
Application to Biotechnology
Because Streptomyces lividans produces a large amount of secretory proteins, it has been widely used in the production and secretion of proteins on industrial interest (2). The Streptomyces genus are soil-dwelling filamentous bacteria, and their natural habitat has allowed them to produce most of the known natural antibiotics, as well as many other secondary metabolites and secreted enzymes important for economic and industrial use (10).
Current Research
In a current study conducted at the University of Wisconsin, capreomycin (CMN), part of a family of antibiotics, is studied for the treatment of multidrug-resistant tuberculosis (MDR-TB). CMN targets the ribosomes of sensitive bacteria and disrupts the function of both subunits. Strepotmyces lividans is genetically altered in this experiment to produce CMN. The altered plasmid inserted into the S. lividans genome conferred resistance to all four CMN derivatives, suggesting that additional resistance genes were encoded within the bacterium genome (6).
Further studies pertaining to the multidrug resistance genes of the Streptomyces genome has been recently conducted. MDR genes are abundant in bacteria, and they encode many membrane-bound protein pumps that export a wide variety of drugs from the cells. Driven by ATP hydrolysis and proton-motive force of the transmembrane proton gradient, these pumps seem to focus on the removal of specific toxic compounds endogenously synthesized during cellular growth rather than the removal of extrinsic drugs. Even though this species has a wide variety of drug export mechanisms, laboratory studies show its sensitivity to most of the drugs tested. However, spontaneous mutants show and elevated efflux of the drugs. One class of spontaneous resistant mutant was isolated from Streptomyces lividans: ethidium bromide (EB) resistance. The protein products of the EB resistant gene appear to be transmembrane proteins that resemble other MDR proteins. The functions and purposes of EB resistant genes, along with other MDR genes, continue to be studied (9).
Current studies have found the complete sequence of a genomic island (GI) named SLG from Streptomyces lividans 66. With an overall G+C content of 67.8%, a DNA S-modification system was determined. Of the 85 open reading frames in SLG, 22 of them showed little homology with previously known proteins. Because of the mosaic structure of the SLG, it seems that there possible recombination events occurred during its formation, including possible spontaneous excision and invasion by a mobile element. The diverse morphology of the SLG could be an example of the successful capture of genetic information by host bacteria (10).
References
Edited by Linnea Shen, student of Rachel Larsen
Edited by KLB