Deinococcus radiodurans: Difference between revisions
No edit summary |
No edit summary |
||
| Line 1: | Line 1: | ||
{{Biorealm Genus}} | {{Biorealm Genus}} | ||
[[Image:|frame|right|An epifluorescence image of stationary-phase ''D. radiodurans R1.'' From [http://www.nature.com/nrmicro/journal/v3/n11/full/nrmicro1264_fs.html Nature Reviews Microbiology.]]] | [[Image:D.radiodurans.r1dna.jpg|frame|right|An epifluorescence image of stationary-phase ''D. radiodurans R1.'' From [http://www.nature.com/nrmicro/journal/v3/n11/full/nrmicro1264_fs.html Nature Reviews Microbiology.]]] | ||
==Classification== | ==Classification== | ||
Revision as of 02:38, 3 May 2007
A Microbial Biorealm page on the genus Deinococcus radiodurans
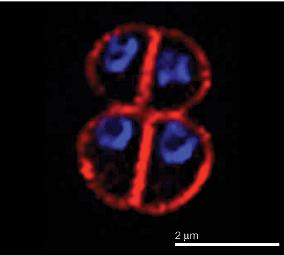
Classification
Higher order taxa
Bacteria; Deinococcus-Thermus; Deinococci; Deinococcales; Deinococcaceae; Deinococcus
Species
D. radiodurans, D. radiodurans R1
Description and significance
Genome structure
The genome of D. radiodurans consists of four major parts. The complete sequence of the R1 strain has 3,284,156 base pairs made up of two circular chromosomes (2,648,638 and 412,348 base pairs), a major plasmid (177,466 base pairs), and a small plasmid (45,704 base pairs). No current research shows whether or not these plasmids contribute specifically to functionality or virality. However, it is known that multiple copies of each gene are found on all the chromosomes and plasmids, which most likely contributes to its amazing repair capabilities associated with its radiation resistance.
Cell structure and metabolism
Ecology
Application to Biotechnology
Current Research
References
Edited by Edwin Cook, student of Rachel Larsen and Kit Pogliano
