Pseudomonas putida’s Role In The Bioremediation Of Plastic: Difference between revisions
| Line 31: | Line 31: | ||
=Bioremediation of Styrene= | =Bioremediation of Styrene= | ||
It is suggested that ''P. putida'' has relatively relaxed substrate specificity; this is further supported by a study showing that when a strain of ''P. putida'' is isolated from conditions where [http://en.wikipedia.org/wiki/Cumene isopropylbenzene] is normally the only carbon source, it is able to grow using alternate aromatic source [[#References|[14]]]. | It is suggested that ''P.putida'' has relatively relaxed substrate specificity; this is further supported by a study showing that when a strain of ''P.putida'' is isolated from conditions where [http://en.wikipedia.org/wiki/Cumene isopropylbenzene] is normally the only carbon source, it is able to grow using alternate aromatic source [[#References|[14]]]. | ||
''P.putida'' is also found to be capable of biodegrading styrene in two different pathways via two modes of initial attack of styrene from the isolation of [[#References|[15]]]. The isolation of two different oxidation products, 2-phenyl-2-propen-1-nol and 1,2-dihydroxy-3-isopropenyl-3-cyclohexene suggest different initial steps of styrene metabolism by ''P.putida'' [[#References|[14]]]. The first being oxidation of the vinyl side chain as the predominant pathway under aerobic conditions; the second involves the direct oxidation of the aromatic ring [[#References|[9]]][[#References|[12]]][[#References|[14]]][[#References|[15]]][[#References|[16]]]. | |||
=Metabolism= | =Metabolism= | ||
Revision as of 02:36, 30 November 2013
Overview
With the rapid industrial development in recent decades, hazardous substance emission is causing more concern. One of the most common materials used in synthetic plastic and rubber is polystyrene, commonly known as styrofoam. This rigid polymer is made of an aromatic monomer phenylethene or styrene. Polystyrene is found to be very recalcitrant and persistent in the environment and the inhalation of polystyrene in the air could cause problems in the central nervous system [1]. The production of polystyrene is also found to be able to cause air pollution by altering the composition of the stratosphere and the troposphere depending on the process used [2]. Therefore it is of great interest to study microorganisms capable of the degradation of styrene such as Pseudomonas putida.
Pseudomonas putida
Domain: Bacteria
Phylum: Proteobacteria
Class: Gamma proteobacteria
Order: Pseudomonadales
Family: Pseudomonadaceae
Genus: Pseudomonas
Species: Pseudomonas putida
Pseudomonas putida is a gram-negative, rod-shaped bacteria which undergoes aerobic metabolism. It is normally found in terrestrial and aquatic environments where oxygen is abundant. P.putida is well known for being one of the most metabolically diverse organisms capable of breaking down both natural organic molecules such as vanillin, limonene and camphor [5] as well as industrial compounds such as toluene and styrene [4]. There is great interest in the genomic analysis of P.putida by a joint genome sequencing project by several institutes from the U.S and Germany with one of the most well studied strains being the P.putida K2440 strain. The genome of this particular strain contains 6181 kbp with 5420 ORFs. Out of all these ORFs, 3571 ORFs were assigned putative roles through comparisons against a non-redundant protein database [6]. Although these ORFs may not all be translated, the abundance of ORFs suggest that the bacteria is capable of various metabolic activities. In addition, to achieve the metabolic versatility of the bacteria requires the ability to cope with numerous abiotic stresses [7]. This is further supported by the genomic analysis of KT2440 where an unusually ample amount of transporters, oxidoreductases, mono-oxygenases, dioxygenases, ferredoxins, sulphur metabolism proteins, dehydrogenases, and cytochromes are found to be encoded in the genome [6]. Aside from P. putida, several other bacteria are also being looked into for bioremediation purposes.
Properties of Polystyrene
Huge amounts of potentially toxic styrene are produced each year, making the removal of the pollutant from the environment of critical importance. There are few microbes capable of metabolizing styrene as a sole source of carbon; these bacteria are able to transform the recalcitrant and persistent styrene into another molecule that’s easier to break down such as polyhydroxyalkanoate or PHA [10]. Styrene is the simplest alkenyl benzene. It is colourless, oil-like and not very soluble in water. The polymerization of styrene into polystyrene is a spontaneous process that can occur at room temperature [9]. Though less toxic than benzene or polycyclic aromatic hydrocarbons, styrene is still potentially carcinogenic. It also affects the central nervous system if inhaled. Chronic exposure to styrene could result in memory loss, difficulties in concentration and learning, and cancer [11]. Most industries deal with styrene pollutants through combustion or land injection. Both methods are flawed as soil styrene rapidly degrades at first but starts to persist subsequently [13]. It is also observed that aerosol styrene created from combustion is capable of damaging the ozone by reacting with O3 to form highly unstable and reactive ozonide [3][9].
Bioremediation of Styrene
It is suggested that P.putida has relatively relaxed substrate specificity; this is further supported by a study showing that when a strain of P.putida is isolated from conditions where isopropylbenzene is normally the only carbon source, it is able to grow using alternate aromatic source [14].
P.putida is also found to be capable of biodegrading styrene in two different pathways via two modes of initial attack of styrene from the isolation of [15]. The isolation of two different oxidation products, 2-phenyl-2-propen-1-nol and 1,2-dihydroxy-3-isopropenyl-3-cyclohexene suggest different initial steps of styrene metabolism by P.putida [14]. The first being oxidation of the vinyl side chain as the predominant pathway under aerobic conditions; the second involves the direct oxidation of the aromatic ring [9][12][14][15][16].
Metabolism
Ignicoccus species are chemolithoautotrophs that use molecular hydrogen as the inorganic electron donor and elemental sulphur as the inorganic terminal electron acceptor[1] . The reduction of the elemental sulphur results in the production of hydrogen sulphide gas.
Ignicoccus are autotrophs in that they fix their own carbon dioxide into organic molecules. The carbon dioxide fixation process they use is a novel process called a dicarboxylate/4-hydroxybutyrate autotrophic carbon assimilation cycle that involves 14 different enzymes[8] .
Members of the Ignicoccus genus are able to use ammonium as a nitrogen source.
Growth Conditions
Because members of the Ignicoccus genus are hyperthermophiles and obligate anaerobes, it is not surprising that their growth conditions are very complex. They are grown in a liquid medium known as ½ SME Ignicoccus which is a solution of synthetic sea water which is then made anaerobic.
Grown in this media at their optimal growth temperature of 90C, the members of the Ignicoccus genus typically reach a cell density of ~4x107cells/mL[1] .
The addition of yeast extract to the ½ SME media has been shown to stimulate the growth and increase maximum cell density achieved. The mechanism by which this is achieved is not known[1] .
Symbiosis
Ignicoccus hospitalis is the only member of the genus Ignicoccus that has been shown to have an extensive symbiotic relationship with another organism.
Ignicoccus hospitalis has been shown to engage in symbiosis with Nanoarchaeum equitans . Nanoarchaeum equitans is a very small coccoid species with a cell diameter of 0.4 µm[9] . Genome analysis has provided much of the known information about this species.
To further complicate the symbiotic relationship between both species, it’s been observed that the presence of Nanoarchaeum equitans on the surface of Ignicoccus hospitalis somehow inhibits the cell replication of Ignicoccus hospitalis . How or why this occurs has not yet been elucidated[3] .
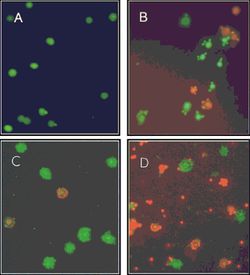
Nanoarchaeum equitans
Nanoarchaeum equitans has the smallest non-viral genome ever sequenced at 491kb[9] . Analysis of the genome sequence indicates that 95% of the predicted proteins and stable RNA molecules are somehow involved in repair and replication of the cell and its genome[3] .
Analysis of the genome also showed that Nanoarchaeum equitans lacks nearly all genes known to be required in amino acid, nucleotide, cofactor and lipid metabolism. This is partially supported by the evidence that Nanoarchaeum equitans has been shown to derive its cell membrane from its host Ignicoccus hospitalis cell membrane. The direct contact observed between Nanoarchaeum equitans and Ignicoccus hospitalis is hypothesized to form a pore between the two organisms in order to exchange metabolites or substrates (likely from Ignicoccus hospitalis towards Nanoarchaeum equitans due to the parasitic relationship). The exchange of periplasmic vesicles is not thought to be involved in metabolite or substrate exchange despite the presence of vesicles in the periplasm of Ignicoccus hospitalis .
These analyses of the Nanoarchaeum equitans genome support the fact of the extensive symbiotic relationship between Nanoarchaeum equitans and Ignicoccus hospitalis. However, it has not yet been proven that it is a strictly parasitic relationship and further research may prove that there is a commensal relationship between the two species.
References
(1) Burggraf S., Huber H., Mayer T., Rachel R., Stetter K.O. and Wyschkony I. ” Ignicoccus gen. nov., a novel genus of hyperthermophilic, chemolithoautotrophic Archaea, represented by two new species, Ignicoccus islandicus sp. nov. and Ignicoccus pacificus sp. nov.” International Journal of Systematic and Evolutionary Microbiology, 2000, Volume 50.
(2) Naether D.J. and Rachel R. “The outer membrane of the hyperthermophilic archaeon Ignicoccus: dynamics, ultrastructure and composition.” Biochemical Society Transactions, 2004, Volume 32, part 2.
(3) Giannone R.J., Heimerl T., Hettich R.L., Huber H., Karpinets T., Keller M., Kueper U., Podar M. and Rachel R. “Proteomic Characterization of Cellular and Molecular Processes that Enable the Nanoarchaeum equitans- Ignicoccus hospitalis Relationship.” PLoS ONE, 2011, Volume 6, Issue 8.
(4) Eisenreich W., Gallenberger M., Huber H., Jahn U., Junglas B., Paper W., Rachel R. and Stetter K.O. “Nanoarchaeum equitans and Ignicoccus hospitalis: New Insights into a Unique, Intimate Association of Two Archaea.” Journal of Bacteriology, 2008, DOI: 10.1128/JB.01731-07.
(5) Grosjean E., Huber H., Jahn U., Sturt H, and Summons R. “Composition of the lipids of Nanoarchaeum equitans and their origin from its host Ignicoccus sp. strain KIN4/I.” Arch Microbiol, 2004, DOI: 10.1007/s00203-004-0725-x.
(6) Briegel A., Burghardt T., Huber H., Junglas B., Rachel R., Walther P. and Wirth R. “Ignicoccus hospitalis and Nanoarchaeum equitans: ultrastructure, cell–cell interaction, and 3D reconstruction from serial sections of freeze-substituted cells and by electron cryotomography.” Arch Microbiol, 2008, DOI 10.1007/s00203-008-0402-6.
(7) Burghardt T., Huber H., Junglas B., Naether D.J. and Rachel R. “The dominating outer membrane protein of the hyperthermophilic Archaeum Ignicoccus hospitalis: a novel pore-forming complex.” Molecular Microbiology, 2007, Volume 63.
(8) Berg I.A., Eisenreich W., Eylert E., Fuchs G., Gallenberger M., Huber H.,Jahn U. and Kockelkorn D. “A dicarboxylate/4-hydroxybutyrate autotrophic carbon assimilation cycle in the hyperthermophilic Archaeum Ignicoccus hospitalis.” PNAS, 2008, Volume 105, issue 22.
(9) Brochier C., Gribaldo S., Zivanovic Y., Confalonieri F. and Forterre P. “Nanoarchaea: representatives of a novel archaeal phylum or a fast-evolving euryarchaeal lineage related to Thermococcales?” Genome Biology 2005, DOI:10.1186/gb-2005-6-5-r42.
(10) Huber H., Rachel R., Riehl S. and Wyschkony I. “The ultrastructure of Ignicoccus: Evidence for a novel outer membrane and for intracellular vesicle budding in an archaeon.” Archaea, 2002, Volume 1.