Pelagibacterales (SAR11): Difference between revisions
| Line 22: | Line 22: | ||
==Subgroups== | ==Subgroups== | ||
Currently the SAR11 clade is divided into five subgroups: | Currently the SAR11 clade is divided into five subgroups: | ||
Subgroup Ia: Open ocean, crown group — includes Pelagibacter unique HTCC1062 | Subgroup Ia: Open ocean, crown group — includes Pelagibacter unique HTCC1062 | ||
| Line 41: | Line 41: | ||
Five SAR11 strains HTCC1062, HTCC1002, HTCC9565, HTCC7211, and HIMB5) form a group of closely related lineages (16S identity ≥ 98%; ANI [average nucleotide identity] ≥ 75%) within SAR11 subclade Ia, which is ubiquitous in geographic distribution (1, 2). Strain HIMB114 is more distantly related (88% 16S identity with HTCC1062) and is part of the subclade IIIa, which is a sister group to the freshwater SAR11 subclade IIIb/LD12 lineage (Vergin et al., submitted). The subclade Va strain HIMB59 is very distantly related (82% 16S identity with HTCC1062) but has been classified as a SAR11 strain based on monophyletic grouping with the other SAR11 strains using both 16S (Vergin et al., submitted) (Fig. 1) and concatenated protein phylogenies (6). Here we present a detailed comparative analysis of these seven SAR11 genomes that provides new insight into the genome features and genetic content of this diverse group of globally abundant organisms. | Five SAR11 strains HTCC1062, HTCC1002, HTCC9565, HTCC7211, and HIMB5) form a group of closely related lineages (16S identity ≥ 98%; ANI [average nucleotide identity] ≥ 75%) within SAR11 subclade Ia, which is ubiquitous in geographic distribution (1, 2). Strain HIMB114 is more distantly related (88% 16S identity with HTCC1062) and is part of the subclade IIIa, which is a sister group to the freshwater SAR11 subclade IIIb/LD12 lineage (Vergin et al., submitted). The subclade Va strain HIMB59 is very distantly related (82% 16S identity with HTCC1062) but has been classified as a SAR11 strain based on monophyletic grouping with the other SAR11 strains using both 16S (Vergin et al., submitted) (Fig. 1) and concatenated protein phylogenies (6). Here we present a detailed comparative analysis of these seven SAR11 genomes that provides new insight into the genome features and genetic content of this diverse group of globally abundant organisms. | ||
General genome features.The strains in this study were isolated from surface seawater of disparate origin: HTCC1062 and HTCC1002 from the temperate coastal Northeast Pacific (4), HTCC9565 from the temperate open ocean of the Northeast Pacific, HTCC7211 from the Sargasso Sea in the subtropical Atlantic (21), and HIMB5, HIMB114, and HIMB59 from the coastal tropical North Pacific (Table 1; see also Table S8 at http://giovannonilab.science.oregonstate.edu/publications). The genomes of HTCC1062, HTCC1002, HTCC7211, HIMB5 and HIMB59 are closed, while the genomes of HIMB114 and HTCC9565 consist of scaffolds with one and three contigs, respectively. Based on synteny with the other genomes of subclade Ia, the amount of missing information for the HTCC9565 genome is estimated to be from <1 to ca. 5.5 kbp. While the degree of completion of the HIMB114 genome is more difficult to estimate, a second recently sequenced subclade IIIa genome is complete at 1.285 Mbp (22), which is less than 50 kbp larger than the current HIMB114 sequence. The presence of a compact (mean genome size of 1.337 ± 0.08 Mbp), low G+C (28.6 to 32.3%) genome is a unifying characteristic of the SAR11 clade (Table 1). The genomes code for between 1,357 and 1,576 genes, one copy of the 5S, 16S, and 23S ribosomal RNA genes, and 30 to 35 tRNAs (see Table S1 at the above URL). No pseudogenes were identified in any of the strains. | |||
==Cell Structure, Metabolism and Life Cycle== | ==Cell Structure, Metabolism and Life Cycle== | ||
Revision as of 13:52, 15 May 2015
Classification
Domain: Bacteria
Phylum: Proteobacteria
Class: Alphaproteobacteria
Subclass: Rickettsidae
Order: Pelagibacterales
Description and Significance
Pelagibacterales (SAR11) is an order in the Alphaproteobacteria composed of free-living planktonic oligotrophic facultative photochemotroph bacteria. (4)(5) They are most abundant group of planktonic cells in marine systems and possibly the most numerous bacterium in the world. Typically accounting for ~25% of prokaryotic cells in seawater worldwide.(3)
The order was originally named SAR11 following its discovery in the Sargasso Sea in 1990 by Professor Stephen Giovannoni and colleagues, from Oregon State University.(2)(5) It was first placed in the order of Rickettsiales, but after rRNA gene-based phyogenetic analysis, in 2013 it was raised to the rank of order, and then placed as sister order to the Rickettsiales in the subclass Rickettsidae.(5)
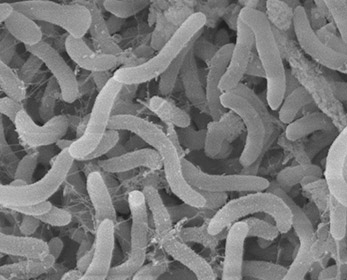
These gram negative, rod shaped bacteria are one of the smallest free cell living organisms, with a cell volume less than 1/500th the volume of E.coli. They have a high surface to volume ratio to better absorb nutrients from its oligotrophic environment. Unsurprisingly, it is also known to have one of the smallest genomes of free living cells.(7) Pelagibacterales have the complete biosynthetic pathway for all 20 amino acids and all but a few cofactors. There are also no pseudogenes, introns, transposons, or extrachromosomal elements yet observed for any cell. (7)
Subgroups
Currently the SAR11 clade is divided into five subgroups:
Subgroup Ia: Open ocean, crown group — includes Pelagibacter unique HTCC1062
Subgroup Ib, open ocean, sister clade to Ia
Subgroup II, coastal, basal to Ia + Ib
Subgroup III, brackish, basal to I + II along with its sister clade IV
Subgroup IV, also known as LD12 clade, freshwater[13]
Subgroup V, which includes alphaproteobacterium HIMB59, basal to the remainder
[[Image:SAR11Phylo.jpg|thumb|500px|FIGURE 3: 16S phylogenetic tree of the SAR11 clade (blue)]
SAR11 has three divergent phylogenetic lineages of the proposed family “Pelagibacteraceae” (Figure 2).
Five SAR11 strains HTCC1062, HTCC1002, HTCC9565, HTCC7211, and HIMB5) form a group of closely related lineages (16S identity ≥ 98%; ANI [average nucleotide identity] ≥ 75%) within SAR11 subclade Ia, which is ubiquitous in geographic distribution (1, 2). Strain HIMB114 is more distantly related (88% 16S identity with HTCC1062) and is part of the subclade IIIa, which is a sister group to the freshwater SAR11 subclade IIIb/LD12 lineage (Vergin et al., submitted). The subclade Va strain HIMB59 is very distantly related (82% 16S identity with HTCC1062) but has been classified as a SAR11 strain based on monophyletic grouping with the other SAR11 strains using both 16S (Vergin et al., submitted) (Fig. 1) and concatenated protein phylogenies (6). Here we present a detailed comparative analysis of these seven SAR11 genomes that provides new insight into the genome features and genetic content of this diverse group of globally abundant organisms.
General genome features.The strains in this study were isolated from surface seawater of disparate origin: HTCC1062 and HTCC1002 from the temperate coastal Northeast Pacific (4), HTCC9565 from the temperate open ocean of the Northeast Pacific, HTCC7211 from the Sargasso Sea in the subtropical Atlantic (21), and HIMB5, HIMB114, and HIMB59 from the coastal tropical North Pacific (Table 1; see also Table S8 at http://giovannonilab.science.oregonstate.edu/publications). The genomes of HTCC1062, HTCC1002, HTCC7211, HIMB5 and HIMB59 are closed, while the genomes of HIMB114 and HTCC9565 consist of scaffolds with one and three contigs, respectively. Based on synteny with the other genomes of subclade Ia, the amount of missing information for the HTCC9565 genome is estimated to be from <1 to ca. 5.5 kbp. While the degree of completion of the HIMB114 genome is more difficult to estimate, a second recently sequenced subclade IIIa genome is complete at 1.285 Mbp (22), which is less than 50 kbp larger than the current HIMB114 sequence. The presence of a compact (mean genome size of 1.337 ± 0.08 Mbp), low G+C (28.6 to 32.3%) genome is a unifying characteristic of the SAR11 clade (Table 1). The genomes code for between 1,357 and 1,576 genes, one copy of the 5S, 16S, and 23S ribosomal RNA genes, and 30 to 35 tRNAs (see Table S1 at the above URL). No pseudogenes were identified in any of the strains.
Cell Structure, Metabolism and Life Cycle
Oenococcus oeni is a facultative anaerobe. It is able to use oxygen for cellular respiration but can also gain energy through fermentation. It characteristically grows well in the environments of wine, being able to survive in acidic conditions below pH 3.0 and tolerant of ethanol levels above 10% (2). Optimal growth occurs on sugar and protein rich media, like grape or tomato juice. The cocci are ellipsoidal to spherical in shape, usually grow in chains or pairs, and are typically non-sporulating. Lactic acid bacteria, like Oenococcus oeni, perform malolactic fermentation (also known as malolatic conversion). It occurs after (or sometimes during) primary fermentation. The main function of malolatic fermentation is converting glucose and malic acid to lactic acid. This is occurs by the uptake of malate, decarboxylation of malate to L-lactic acid and carbon dioxide, and the export of end products. O. oeni is heterofermentative, meaning it can create multiple end products from fermenting the sugars. In O. oeni’s case, it produces carbon dioxide, ethanol, and acetate, as well as characteristic flavor molecules like diacetyl. Strain variation of Oenoccocus oeni cellular processes can have significant effects on the community dynamics, fermentation, and overall quality of wine. Strain variation exists in sugar utilization pathways, phosphototransferase enzyme II systems, bacteriophage integration, and cell wall exopolysaccharides (3).
Ecology
Oenococcus oeni stabilizes wine communities by consuming available nutrients and lowering potential growth of other microbes, but its malolactic fermentation can be beneficial or detrimental to the production of wine depending on grapes, climate, and style of wine. Variations between strains and fermentation conditions have the potential to impact general quality and production of wine. Industrial winemakers use a standardized strain of O. oeni, but the many external and environmental variables will dictate the success of the wine. O. oeni is not the only lactic acid bacteria that can perform secondary fermentation. There are a variety of lactic acid bacteria that can dominate bacterial community in response to temperature, nutrients, sulfur dioxide content, pH, ethanol levels, and inoculation densities. O. oeni commonly dominates secondary fermentation from its extreme tolerance to pH and ethanol levels. The molecule diacetyl is produced as a byproduct of lactic acid bacteria in secondary fermentation. In wine, the levels of diacetyl create buttery and caramel flavor notes. It is generated when there is little or no malic acid to be consumed so citric acid is used. This byproduct is sought out by some winemakers while it is avoided by others. Rarely do the diacetyl levels reach a point of spoiling the wine (10).
Because of its heterofermentive properties, Oenoccocus may be viewed as an ecosystem engineer. O. oeni plays a major role in establishing the environment for which other microbes will interact. Its end products and life strategies positively feedback into creating a more harsh environment for other microbes like yeasts and fungi while making the conditions more ideal for other lactic acid bacteria (5).
References
(1) "Rebounding bacteria". 2013
(2) “SAR11 bacteria thrive — despite viruses”.“Nature”
(6) A war without end - with Earth’s carbon cycle held in the balance
(9) phys.org “Raise your glass to Oenococcus oeni, a real wine bug”. “Science X Network”. 2014.
Author
Page authored by Digvinder Singh Mavi student of Prof. Katherine Mcmahon at University of Wisconsin-Madison.