Desulfovibrio desulfuricans: Difference between revisions
Rsmolinski (talk | contribs) |
Rsmolinski (talk | contribs) |
||
| Line 24: | Line 24: | ||
''D. desulfuricans'' ND132 is a strain of wildtype ''D. desulfuricans'' with the property to produce methylmercury. It has been fully sequenced by Brown et al., 2011. Sequence data is available online at GenBank at https://www.ncbi.nlm.nih.gov/nuccore/CP003220. Its genome is circular, ~3.8 Mb in length, codes for 3454 proteins, has a GC content of 65.21%, and has a coding density of 88.41%, which is based off of averages of ~40x and ~122X genome coverage for 454 and Illumina data respectively (Figure 3) (Brown et al., 2011). | ''D. desulfuricans'' ND132 is a strain of wildtype ''D. desulfuricans'' with the property to produce methylmercury. It has been fully sequenced by Brown et al., 2011. Sequence data is available online at GenBank at https://www.ncbi.nlm.nih.gov/nuccore/CP003220. Its genome is circular, ~3.8 Mb in length, codes for 3454 proteins, has a GC content of 65.21%, and has a coding density of 88.41%, which is based off of averages of ~40x and ~122X genome coverage for 454 and Illumina data respectively (Figure 3) (Brown et al., 2011). | ||
'''FIG. 3.''' There are 43 mean spacer HisKA-ATPase domains, 39 min spacer HisKA-ATPase domains, and 55 max spacer HisKA-ATPase domains present. This genome contains 77 response regulators; NtrC (15), OmpR (4), NarL (2), LytTR (2), CheB (2), RpfG (3), CheV (4), PleD_VieA (1), PleD (3), CheY (28), TrxB (1), VieB (1), and 11 unclassified. | '''FIG. 3.''' There are 43 mean spacer HisKA-ATPase domains, 39 min spacer HisKA-ATPase domains, and 55 max spacer HisKA-ATPase domains present. This genome contains 77 response regulators; NtrC (15), OmpR (4), NarL (2), LytTR (2), CheB (2), RpfG (3), CheV (4), PleD_VieA (1), PleD (3), CheY (28), TrxB (1), VieB (1), and 11 unclassified. From ''Desulfovibrio desulfuricans'' ND132, 2016. | ||
From ''Desulfovibrio desulfuricans'' ND132, 2016. | |||
=Cell structure and metabolism= | =Cell structure and metabolism= | ||
Revision as of 20:31, 2 December 2016
Classification
Higher order taxa
Domain Bacteria; Phylum Proteobacteria; Class Deltaproteobacteria; Order Desulfovibrionales; Family Desulfovibrionaceae; Genus Desulfovibrio (Madigan, et al., 2012).
Species
Desulfovibrio desulfuricans (also known as strain ND 132)
Description and significance
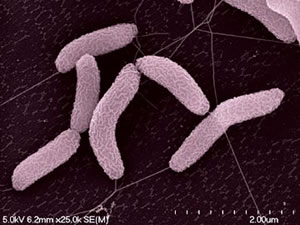
Desulfovibrio desulfuricans strain ND 132 was isolated from Chesapeake Bay sediments in May 1985 (Gilmour et al., 2011). Its cells are gram-negative, slightly curved rods (Gilmour et al., 2011) (Figure 1). D. desulfuricans ND 132 is an anaerobic sulfate-reducing bacterium which has the ability to produce methylmercury (MeHg), a powerful neurotoxin in humans (Brown et al., 2011). Sulfate-reducing bacteria are found in oxic-anoxic zones of sediments and soils where methylation rates are very high (Jay et al., 2016). MeHg production has many implications as having powerful bioaccumulating abilities throughout food webs, causing much damage to ecosystems, in addition to being a neurotoxin.
16S Ribosomal RNA Gene Information
16S Ribosomal RNA Gene Information
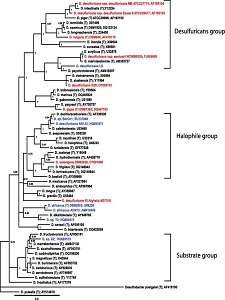
The 16S rRNA sequencing of Desulfovibrio desulfuricans ND132 is currently designated within the family of desulfuricans (Figure 2), however, there is much similarity between this strain and strains of D. dechloroacetivorans (98-99%) indicating proof its phylogenetic placement should be reconsidered (Brown et al., 2011).
Genome Structure (if the genome exists)
D. desulfuricans ND132 is a strain of wildtype D. desulfuricans with the property to produce methylmercury. It has been fully sequenced by Brown et al., 2011. Sequence data is available online at GenBank at https://www.ncbi.nlm.nih.gov/nuccore/CP003220. Its genome is circular, ~3.8 Mb in length, codes for 3454 proteins, has a GC content of 65.21%, and has a coding density of 88.41%, which is based off of averages of ~40x and ~122X genome coverage for 454 and Illumina data respectively (Figure 3) (Brown et al., 2011).
FIG. 3. There are 43 mean spacer HisKA-ATPase domains, 39 min spacer HisKA-ATPase domains, and 55 max spacer HisKA-ATPase domains present. This genome contains 77 response regulators; NtrC (15), OmpR (4), NarL (2), LytTR (2), CheB (2), RpfG (3), CheV (4), PleD_VieA (1), PleD (3), CheY (28), TrxB (1), VieB (1), and 11 unclassified. From Desulfovibrio desulfuricans ND132, 2016.
Cell structure and metabolism
D. desulfuricans strain ND 132 is an anaerobic dissimilatory sulfate-reducing bacterium. This strain ND132 can be isolated from estuarine sediment and anoxic saltmarsh sediment (Compeau and Bartha, 1985). Due to high production of methylmercury that occurs during the sulfate reduction, the production of mercury is linked to Hg-specific uptake pathway and biochemistry of Hg binding which depends on enzyme methyltransferase (Gilmour et. al., 2011).
Ecology and Pathogenesis
Sulfate reducers thrive in marine systems, owing to the high concentration of sulfate ions in seawater. They can be found in various environments such as oil field production waters, acid mine drainage waters, sewage, and soil. However the genetic structures of the sulfate-reducing bacteria differ in different environments (Verstreken et al., 2012). This proves that Desulfovibrio desulfuricans has an excellent ability to adapt to its surroundings while maintaining an optimal cell structure for nutrient uptake and allows for the anaerobic production of methylmercury. D. desulfuricans are not known to be pathogenic, however an infection of this microbe into the human body can prove to be detrimental to one’s health. We see an example of this in the case of a 69 year old woman who had been hospitalized with complaints of stomach irritation, irregular bowel movements, diarrhea, and fever. The final diagnoses being D. desulfuricans bacteremia occurring in an immunocompromised host with CMV colitis (Voorduow et al., 1995). Doctors then incubated the strain revealing a resistance to piperacillin-tazobactam. Desulfovibrio desulfuricans culture under anoxic conditions is more likely to form colonies in free living cells than particle associated environments (Voorduow et al., 1995).
Current Research
Desulfovibro desulfuricans strain ND132 presents an excellent opportunity for the study of mercury methylation because while being a typical anaerobic mesophilic bacterium with wide tolerance of pH and salinity, it also grows well with fumurate as electron acceptor, which prevents sulfide inhibition of mercury methylation (Gilmour et al., 2011). Its genome sequencing allows comparative transcriptomic and proteomic study against other Desulfovibrio species (Gilmour, et al., 2011) Additional research includes the effects of large colony blooms in seawater and effects of infections in the human body by D. desulfuricans. One of the main research proposals currently being performed on D.desulfuricans is the study of how methylmercury production occurs with this organism’s cellular respiration. D.desulfuricans is the only organism in which pathways of its methylation have been somewhat defined (Gilmour et. al., 2011). This team of scientists used enriched stable mercury isotopes to show that ND132 simultaneously produces and degrades methylmercury (MeHg) during growth but does not produce elemental Hg. A large colony of strain ND132 may cause harm to someone who comes in contact with the colony as methylmercury is known to cause brain and nervous system damage. There is still much research that can be done on this organism which may prove beneficial to human society as we learn about new ways to prevent corrosion caused by ND132, or as we may be able to utilize their respiration in some wastewater treatment plants.
References
xBrown, S. D., Gilmour, C. C., Kucken, A. M., Wall, J. D., Elias, D. A., Brandt, C. C., … Palumbo, A. V. (2011). Genome Sequence of the Mercury-Methylating Strain Desulfovibrio desulfuricans ND132. J. Bacteriol, 193(8): 2078-2079.
xCompeau, G. C., & Bartha, R. (1985). Sulfate-reducing bacteria: principal methylators of mercury in anoxic estuarine sediment. Applied and environmental microbiology, 50(2), 498-502.
xDesulfovibrio desulfuricans ND132. (2016, December). Retrieved from http://www.p2cs.org/page.php?base=Desd2DB
xGilmour, C., Elias, D., Kucken, A., Brown, S., Palumbo, A., Schadt, C., & Wall, J. (2011). Sulfate-Reducing Bacterium Desulfovibrio desulfuricans ND132 as a Model for Understanding Bacterial Mercury Methylation. Applied and environmental microbiology, 77(12). doi:10.1128/AEM.02993-10.
Jay, J., Murray, K., Gilmour, C., Mason, R., Morel, F., Roberts, A., & Hemond, H. (2016). Mercury Methylation by Desulfovibrio desulfuricans ND132 in the Presence of Polysulfides. Applied and Environmental Microbiology, 82(20). Doi:10.1128/AEM.68.11.5741-5745.2002. http://www.nature.com/articles/srep12872
xMadigan, M. T., Martinko, J. M., Stahl, D. A., & Clark, D. P. (2012). Brock biology of microorganisms. Boston: B. Cummings.
Rani, A., Rockne, K. J., Drummond, J., Al-Hinai, M., & Ranjan, R. (2015). Geochemical influences and mercury methylation of a dental wastewater microbiome [Abstract]. Scientific Reports, 5(12872). doi:10.1038/srep12872
Steger, J., Vincent, C., Ballard, J., Krumholtz, L. (2016). Desulfovibrio sp. Genes involved in the Respiration of Sulfate during Metabolism of Hydrogen and Lactate. Applied and Environmental Microbiology, 82(20). Doi: 10.1128/AEM.68.4.1932-1937.2002.
xVoordouw, G. (1995). The Genus Desulfovibrio: The Centennial. Applied and Environmental Microbiology, 61(8). Pg 2813-2819. Retrieved from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1388543/pdf/hw2813.pdf
xVerstreken, I., Lalemanb, W., Wauters, G., & Verhaegen, J. (2012). Desulfovibrio desulfuricans bacteremia in an immunocompromized host with a liver graft abd ulcerative colitis. American Society for Microbiology. 54(11). doi:10.1128/JCM.00987-11.
Author
Page created by Cameron Garcia, Christina Lopardo, Luka Ndungu, Rachel Smolinski, students of Dr. Hidetoshi Urakawa at Florida Gulf Coast University.