Staphylococcus epidermidis: Difference between revisions
| Line 42: | Line 42: | ||
==Pathology== | ==Pathology== | ||
The increase use of intravascular catheters has caused a similar increase in S. epidermidis infections. However, S. epidermidis is resistant to methicillin and all penicillins, penems, carbapanems, and cephalosporins which are commonly used antibiotics (3). S. epidermidis has also been found to be more resistant to antibiotics than other species (4). | |||
Although there are no one specific viruelence factors of S. epidermidis, the ability to form biofilm is one of the virulence factors (5,6). The biofilm allows the bacteria cells to adhere to an inert or living areas (5). When a biofilm has formed it becomes harder to treat since the cells inside the biofilm are guarded from antibiotics and the immune system (5). Biofilm also release a host immune response to antigens which prevents the removal of the biofilm and may also result in tissue damage (9). The bacteria can be released into the blood from biofilms and start new infections by attachment to the medical devices, thus devices will be required to be removed (8). | |||
Some preventive strategies for infections are to provide prohylatic antibiotic thereapy to cover surgical insertions from temporary intravascular devices. There are also reports that warn to not use antibiotic prohylaxis, especially vancomycin for dialysis. Aseptic techniques should be used for catheter infections to prevent contamination. New techniques concentrate on physical electrical barriers for colonization and using biomaterials with antimicrobial agents already inside. However these new methods have not been tried in the clincal settings(9). | |||
==Application to Biotechnology== | ==Application to Biotechnology== | ||
Revision as of 18:38, 29 August 2007
A Microbial Biorealm page on the genus Staphylococcus epidermidis
Classification
Higher order taxa
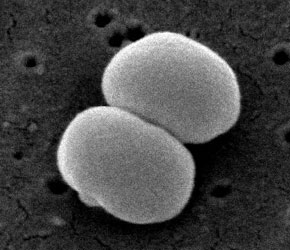
Bacteria; Firmicutes; Bacillales; Staphylococcus (1)
Species
Genus Species: Staphylococcus epidermidis
Other Names: Micrococcus epidermidis, Albococcus epidermidis, Staphylococcus epidermidis albus (2)
|
NCBI: Taxonomy |
Description and significance
Staphylococcus epidermidis is a gram-positive and coagulase-negative staphylococci (4). It typically lives on the human skin and mucosa and the most common infections on catheters and implants (5). S. epidermidis is one of five most common organisms that cause noscomial infections due to the increase in usage of biomaterials in the clinical environment (8). The nosocomial pathogen causes infections on prosthetic valves, cerebrospinal fluid shunts, joint prosthesis vascular prostheses, valves, and in postoperative wounds and the urinary tract. It is also the most freqent organism found in the blood of bone marrow transplant patients and on central venous catheters for patients of total parenteral nutition (4). A controversial issue on S. epidermidis argues whether or not all strains can equally cause disease and if a specific genotype determines a specific disease (6).
S. epidermidis strain from health adult nares show that there are many kinds of the organism in each individual (9). A common S. epidermidis strain is RP62a (ATCC 35984) was isolated in Memphis, Tennessee during 1979-1980 in a wide spread of intravascular catheter sepsis. RP62a is a strain that produces slime, grows collectively and forms biofilm. The treatment of S. epidermidis infections caused by biofilm with antibiotics is often ineffective and results in removal of the implants (5). Another strain is S. epidermidis ATCC 12228 which does not produce biofilm (7).
The ability to produce slime allows the pathogen to stay on biomaterials. Different strains of S. epidermidis can be distinguished by whether or not they produce slime. In clinical laboratories, S. epidermidis infections are studied by the use of antibiograms, biotyping, plasmid profiles, and slime production; whereas in research, phagetyping, serotyping, restrict enzyme analysis and DNA-DNA hybridization are used (4).
Genome structure
Random shotgun method was used to sequence S. epidermidis strain RP62a completely at The Institute for Genomic Research (TIGR) (7). RP62A’s chromosome length is 2,616,530bp, contains 32.10% of G+C content, 6 rRNA operons and plasmid with 28,080bp and 32% G+C content (11). Two plasmids were identified in the strains RP62a and ATCC 12228; vSe1 and vSe2 which have phrophage integrase genes. The plasmid vSe1 have genes for cadmium resistance whereas a second strain-specific sortase and two LPXTG surface attachment proteins is encoded by vSe2 of ATCC 12228 (7).
Cell structure and metabolism
When compared to other bacteria such as micrococcus, S. epidermidis’ cell wall is much stronger. The addition of lysostaphin can differentiate S. epidermidis from micrococcus. Micrococcus is more likely to lyse whereas the cell wall of S. epidermidis contain chemical properties of the peptidoglycan that prevent it from lysis. There are endopeptidases that cut the glycl-glycine bonds in the penta or hexpeptide crossbridge of the peptidoglycan of S. epidermidis. The strains that contain serine in the interpeptide bridges are more resistant to lysis (4).
The cell wall of Staphylococci contain teichoic acids which are connected to the peptidoglycan by covalent bonds. The teichoic acids are composed of either glycerol or ribitol which are connected by phosphodiester bonds. They are water soluble polymers made up of 30-50% of the dry components of the cell. S. aureus and S. epidermidis can be distinguished by having ribitol or glycerol. S. epidermidis has glycerol teichoic acid glucosyl residues while S. aureus has N-acetylglucosamine ribitol teichoic acid (4).
The metabolic characteristics of S. epidermidis consists of the capability to grow with glucose anaerobically, produce nitrate from nitrate, cannot create coagulase or ferment mannitol, and need biotin to grow. Most strains of S. epidermidis make acetoin, phosphatase and reduce nitrate. With oxygen, all strains can produce acid when exposed to glucose, fructose, maltose, sucrose, and glycerol and 70%-90% with galactose, mannose, and lactose. Acid cannot be formed from mannitol, trehalose, rhamnose, xylose, or arabinose (4).
Ecology
The natural environment of S. epidermidis is the human body and usually originates from disease. Since the bacteria usually lives on the skin and nares of all human beings and is a nosocomial pathogen, it is imporant to be able to identify the specific strains. S. epidermidis is the most common staphylococcus on the human skin. In addition, S. epidermidis also covers 90%-100% staphylococci from the nares when S. aureus is not present. When S. aureus is present the S. epidermidis amount decreases drastically (4).
Pathology
The increase use of intravascular catheters has caused a similar increase in S. epidermidis infections. However, S. epidermidis is resistant to methicillin and all penicillins, penems, carbapanems, and cephalosporins which are commonly used antibiotics (3). S. epidermidis has also been found to be more resistant to antibiotics than other species (4).
Although there are no one specific viruelence factors of S. epidermidis, the ability to form biofilm is one of the virulence factors (5,6). The biofilm allows the bacteria cells to adhere to an inert or living areas (5). When a biofilm has formed it becomes harder to treat since the cells inside the biofilm are guarded from antibiotics and the immune system (5). Biofilm also release a host immune response to antigens which prevents the removal of the biofilm and may also result in tissue damage (9). The bacteria can be released into the blood from biofilms and start new infections by attachment to the medical devices, thus devices will be required to be removed (8).
Some preventive strategies for infections are to provide prohylatic antibiotic thereapy to cover surgical insertions from temporary intravascular devices. There are also reports that warn to not use antibiotic prohylaxis, especially vancomycin for dialysis. Aseptic techniques should be used for catheter infections to prevent contamination. New techniques concentrate on physical electrical barriers for colonization and using biomaterials with antimicrobial agents already inside. However these new methods have not been tried in the clincal settings(9).
Application to Biotechnology
Does this organism produce any useful compounds or enzymes? What are they and how are they used?
Current Research
Enter summaries of the most recent research here--at least three required
References
Edited by Angela Cheng, student of Rachel Larsen at UCSD
