Computer Logic in Microbial Systems
Introduction

By Jeremy Moore
Synthetic biology is a quickly evolving field that fuses biological and chemical science with engineering principles. One major task for synthetic biologists is harnessing cell’s innate ability to perform tightly concerted metabolic processes in order to produce molecules of industrial and medical relevance. In other words, one goal of synthetic biology is to create a programming language for cellular processes that can be altered in deterministic ways to generate specific products. Applying computer logic to living systems is challenging, as gene regulation is highly sensitive to the environment and requires a tightly controlled balance of regulatory factors[2]. Nonetheless, several methods have been developed to translate logical operators into genetic circuits.
Programmable cells have myriad applications to industry and medicine. As an example, a strain of Yersinia pseudotuberculosis was modified to invade cancerous cells in response to environmental conditions [1]. Applications like this could be used to create synthetic organisms capable of accomplishing highly specific tasks such as targeting specific tissues or compounds.
Logic Gates in Biological Context


tion.
Counting and Memory Function Using Invertases

Construction of Complex Logic


Oscillating Synthetic Gene Networks
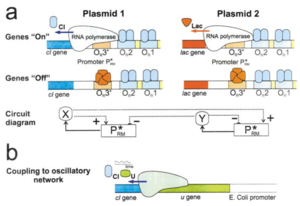
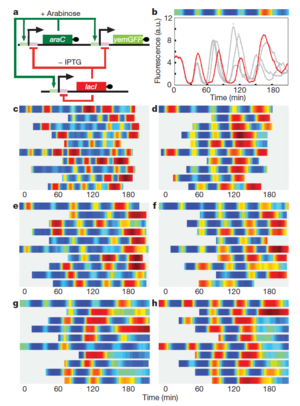
Circuit Interactions with the Host Cell
Conclusion
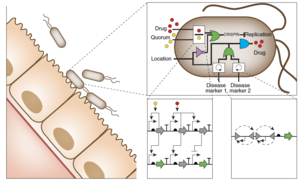
References
- ↑ 1.0 1.1 JC, Clarke EJ, Arkin AP, Voigt CA. Environmentally Controlled Invasion of Cancer Cells by Engineered Bacteria. (2006). JMB 355: 619 – 627. doi:10.1016/j.jmp.2005.10.076.
- ↑ 2.0 2.1 2.2 Brophy JAN, Voigt CA. Principles of Genetic Circuit Design. (2014). Nature Methods 11(5): 508 – 520. DOI:10.1038/NMETH.2926.
- ↑ 3.0 3.1 3.2 http://www.nature.com/nature/journal/v469/n7329/abs/nature09565.html Tamsir A, Tabor JJ, Voigt CA. Robust multicellular computing using genetically encoded NOR gates and chemical ‘wires’. (2011) Nature 469: 212 – 215. doi:10.1038/nature09565.
- ↑ . Friedland AE, Lu TK, Wang X, Shi D, Church G, Collins JJ. Synthetic Gene Networks that Count. (2009). Science 324(5931): 1199 – 1202.
- ↑ https://journals.aps.org/prl/abstract/10.1103/PhysRevLett.88.148101 Hasty J, Dolnik M, Rottschafer V, Collins JJ. Synthetic Gene Network for Entraining and Amplifying Cellular Oscillations. 2002. Physical Review Letters 88(14) 148101.
- ↑ http://www.nature.com/nature/journal/v456/n7221/full/nature07389.html Stricker J, Cookson S, Bennett MR, Mather WH, Tsimring LS, Hasty J. A fast, robust and tunable synthetic gene oscillator. 2008. Nature 456(27) 07389.
Authored for BIOL 238 Microbiology, taught by Joan Slonczewski, 2017, Kenyon College.
