Soil Unknown: P. aeruginosa
Classification
| Domain | Bacteria |
|---|---|
| Phylum | Proteobacteria |
| Class | Gammaproteobacteria |
| Order | Pseudomonadales |
| Family | Pseudomonadaceae |
| Genus | Pseudomonas |
| Species Group | Pseudomonas aeruginosa group |
| Species | Pseudomonas aeruginosa |
Species
|
NCBI: Taxonomy |
| Genus species: | Pseudomonas aeruginosa |
|---|
Habitat Information
The organism was isolated from 1 gram of a soil sample gathered from a park in the Barton Creek Landing complex in Travis County. In general, P. aeruginosa can be found in soil, decaying organic matter, and a variety of moist environments, including swimming pools, hot tubs, sponges, washcloths, and contact lens solutions (1).
Description and Significance
Appearance
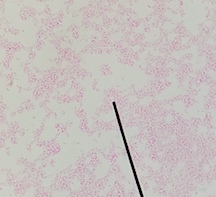
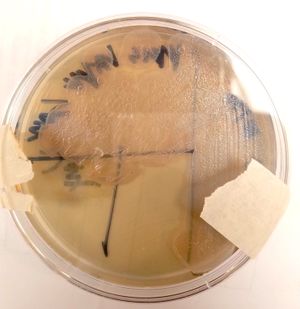
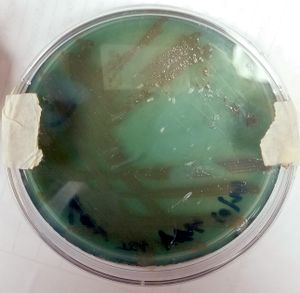
At the cellular level, P. aeruginosa is a Gram Negative bacilli. At the colonial level, P. aeruginosa colonies appear to be of small to moderate, irregular, pulvinate and smooth. When incubated at 30 degrees celsius, the colonies have a small amount of orange pigmentation. When incubated at 37 degrees celsius, the colonies have gray pigmentation, as well as blue-green extracellular pigmentation:
Antimicrobial Activity
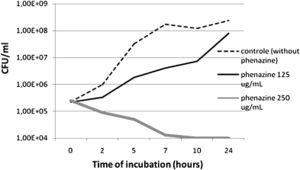
A study published in the Annals of Clinical Microbiology and Antimicrobials concluded that P. aeruginosa-derived extracellular compounds such as phenazines have inhibitory effects against MRSA. A synergic effect was observed when combined with silver nanoparticles produced by Fusarium oxysporum (2).
Strain PR3 of P. aeruginosa has been found to have antifungal properties, inhibiting both Candida albicans and rice blast fungus (3.)
Significance of organism
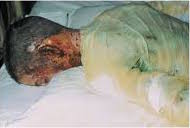
Psuedomonas aeruginosa is a rare opportunist because it cannot penetrate the intact, natural defenses of the skin. Susceptible potential victims include burn victims, cancer patients, and immunocompromised patients. Those with cystic fibrosis have a higher risk of acquiring a P. aeruginosa lung infection.
P. aeruginosa is the most common microorganism seen in burn victims, and the microbe typically grows underneath the scab-like crust that naturally forms over a severe burn. Once established, it kills cells and destroys tissues. Large infections can be diagnosed due to the microbe's blue-green pyocyanin pigment. If P. aeruginosa invades the bloodstream, severe symptoms including fever, chills and shock can result
In general, a penicillin and an aminoglycoside are simultaneously used to treat P. aeruginosa infections. To treat a P. aeruginosa infection in a burn patient, debridement and administration of antimicrobial drugs are necessary. (1)
Genome Structure
Describe the size and content of the genome. How many chromosomes? Circular or linear? Other interesting features? What is known about its sequence? Include S Ribosomal sequence that you obtained from PCR and sequencing here.
At 6.3 million base pairs, this is the largest bacterial genome sequenced.
S Ribosomal sequence: Group_2R-ACC_primer_-_reverse_D07.ab1 NNNNNNANNNNNNNCNNCCTGTNACTCTGTCCCCGAAGGGAAAGCCCTATCTCTAGGGTTGTCAGAGGATGTCAAGACCT GGTAAGGTTCTTCGCGTTGCTTCGAATTAAACCACATGCTCCACCGCTTGTGCGGGCCCCCGTCAATTCCTTTGAGTTTC AGTCTTGCGACCGTACTCCCCAGGCGGAGTGCTTAATGCGTTAGCTGCAGCACTAAGGGGCGGAAACCCCCTAACACTTA GCACTCATCGTTTACGGCGTGGACTACCAGGGTATCTAATCCTGTTCGCTCCCCACGCTTTCGCTCCTCAGCGTCAGTTA CAGACCAGAGAGTCGCCTTCGCCACTGGTGTTCCTCCACATCTCTACGCATTTCACCGCTACACGTGGAATTCCACTCTC CTCTTCTGCACTCAAGTTTCCCAGTTTCCAATGACCCTCCCCGGTTGAGCCGGGGGCTTTCACATCAGACTTAAGAAACC GCCTGCGAGCCCTTTACGCCCAATAATTCCGGACAACGCTTGCCACCTACGTATTACCGCGGCTGCTGGCACGTAGTTAG CCGTGGCTTTCTGGTTAGGTACCGTCAAGGTGCGAGCAGTTACTCTCGCACTTGTTCTTCCCTAACAACAGAGCTTTACG ATCCGAAAACCTTCATCACTCACGCGGCGTTGCTCCGTCAGACTTTCGTCCATTGCGGAAGATTCCCTACTGCTGCCTCC NNNNNNNNACTGNCCNNNNNNTTTNCNA
Cell Structure, Metabolism and Life Cycle
Interesting features of cell structure; how it gains energy; what important molecules it produces.
P. aeruginosa is a Gram negative bacterium that is typically found in water and soil. Considered one of the top 3 opportunistic pathogens... It forms biofilms on rock
Physiology and Pathogenesis
Identifying biochemical characteristics and enzymes made
If relevant, how does this organism cause disease?
| Gram Stain | Negative |
|---|---|
| Motility Test | Positive |
| Gelatinase Test | Positive: produces gelatinase enzyme |
| DNA Hydrolysis Test | Negative: no deoxyribonuclease enzyme |
| Lipid Hydrolysis Test | Positive: Presence of Lipase enzyme |
| Phenol Red Glucose | Orange (shows some glucose fermentation, though P .aeruginosa does not ferment glucose.) |
| Phenol Red Sucrose | Red: no Sucrose fermentation |
| Phenol Red Lactose | Red: no Lactose fermentation. |
| Starch Hydrolysis | Negative: no Amylase enzyme |
| Casein Hydrolysis | Positive: Casease enzyme |
| SIM Medium | Positive for Motility and Sulfur Reduction. Indole production: negative |
| TSI Test | K/NC, H2S: Negative fermentation, Positive H2S reduction |
| Citrate Test | Positive: Citrate as a carbon source |
| Urease Test | Negative: no Urease enzyme |
| MR-VP Tests | Negative MR: does not ferment glucose. Negative VP: does not use butylene glycol pathway |
| Nitrate Reduction Test | Positive: denitrification took place |
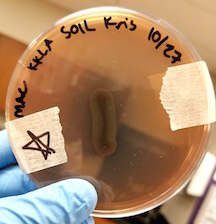
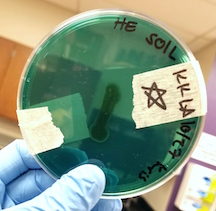
| Hektoen Enteric Agar | Negative: Non-Lactose Fermenting (NLF) |
| MacConkey Agar | Negative: NLF |
| Oxidase Test | Negative: does not produce cytochrome oxidase |
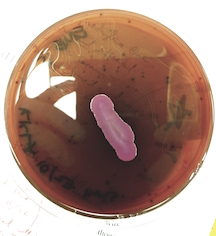
| Eosin Methylene Blue Agar | Weak positive: Indicates weak lactose fermentation, though P .aeruginosa does not ferment lactose. |
| Phenylalanine Deaminase | Negative: no Deaminase enzyme |
| Decarboxylation Test: Arganine | Positive: produces arginine decarboxylase |
| Decarboxylation Test: Lysine | Negative: does not produce lysine decarboxylase |
| Decarboxylation Test: Ornithine | Negative: does not produce ornithine decarboxylase |
| Catalase Test | Positive: Catalase enzyme |
| Blood Agar | Alpha hydrolysis: partial hemolysis |
| Bacitracin & Optichin Susceptibility | Negative: Neither inhibited growth |
| Phenylethyl Alcohol Agar | Very slow growth: not a Gram Positive organism |
| Mannitol Salt Agar | Negative: Inhibited by salt |
| 6.5% Salt Tolerance Test | Negative: Inhibited by salt |
| Bile Esculin Test | Negative: does not hydrolyze esculin |
Hosts
- Humans - those most susceptible include burn victims, cancer patients, and immunocompromised patients. Cystic fibrosis patients have a higher risk of lung infection.(1)
- Animals - similar susceptibility patterns to humans. The most commonly colonized sites include wet or moist anatomic areas like the gut and respiratory tract, along with chronically wet areas of fur. (5)
- Plants - certain strains are capable of infecting plant roots. A study published in Plant Physiology details two P.aeruginosa strains that infect the roots of Arabidopsis and sweet basil, and are capable of causing plant mortality. (6)
Virulence factors
- Fimbriae and adhesions aid in attachment to host, as well as enabling biofilm formation.
- Capsule helps shield the bacteria from phagocytosis, and also aids in bacterial attachment and biofilm formation.
- Neuraminidase enzyme helps bacteria attach to host.
- Elastase enzyme breaks down IgA, IgG, degrades complement components, and breaks down elastic fiber.
- Lipid A component can trigger fever, blood clotting, inflammation, or shock.
- Exotoxin A and exoenzyme S inhibit eukaryotic protein synthesis.
- Pyocyanin pigment triggers the formation of reactive forms of oxygen, which damages host cells (1).
Patient Symptoms
- Blue-green color where the bacterium is growing
- With bacteremia: fever, blood clotting, inflammation and shock are possible
- With lung infections: breathlessness, coughing, wheezing, rapid breathing, and weight loss (1)
References
1. Bauman, R. W. (2015) Microbiology with Diseases by Body System, pp. 567, 570. Glenview, IL: Pearson.
2. Cardozo, V., Oliveira, A.,Nishio, E. Antibacterial activity of extracellular compounds produced by a Pseudomonas strain against methicillin-resistant Staphylococcus aureus (MRSA) strains. Annals of Clinical Microbiology and Antimicrobials. 2013. Volume 12:12.
3. Bajpai, Vivek K.; Shin, Seung Yong; Kim, Hak Ryul; Kang, Sun Chul. 2008. Anti-fungal action of bioconverted eicosapentaenoic acid (bEPA) against plant pathogens. Industrial Crops and Products. 27(1): 136-141.
4. https://www.nature.com/articles/35023079
5. Psuedomonas aeruginosa Technical Sheet. Charles River Laboratories, 2017.
6.Walker, T., Bais, H., Deziel, E., and Schweizer, H., ''Pseudomonas aerutinosa-plant root interactions. Pathogenicity, biofilm formation, and root exudation. Plant Physiology'. 2004. Volume 134. p. 320 - 331.
Author
Page authored by Leila Adell and Katie Kelinske, students of Prof. Kristine Hollingsworth at Austin Community College.