Chimpanzee Evolution
Introduction and Early Evolution
Pan troglodytes, better known as chimpanzees, are a species of great ape widely regarded as the closest living relatives to bonobos (Pan paniscus) and one of the closest living relatives to humans (Homo sapiens; Fig. 1).[1][2]

The evolution of chimpanzees is thought to have begun, like that of many other primates, with the divergence of platyrrhines (also known as New World monkeys) and catarrhines (a group containing Old World monkeys and apes) 25-40 million years ago [3] This was followed by a divergence in the catarrhines, forming the Hominoidea (ape) and Cercopithecoidea (Old World monkey) groups 23 million years ago. [4] It is estimated that the genera Homo and Pan diverged 5-6 million years ago, followed by a split in the genus Pan roughly 2.4 million years ago, eventually culminating into the chimpanzees and bonobos present today (Fig. 2). [5][6] [7][8]
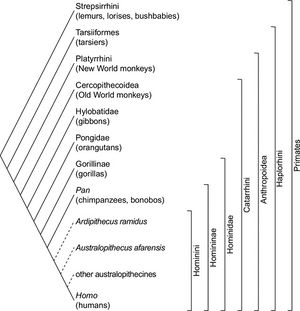
Continuing Evolution and Speciation
Evolution has not stopped with the modern chimpanzee. There are currently four recognized subspecies: the central chimpanzee (P. t. troglodytes), the western chimpanzee (P. t. verus), the eastern chimpanzee (P. t. schweinfurthii), and the Nigeria-Cameroon chimpanzee (P. t. ellioti) (Fig. 3). [9][10][11][12]
The subspecies are geographically isolated by the Sanaga River, which separates the western and Nigeria-Cameroon subspecies from the central and eastern chimpanzees, and the Ubangi River, which is the primary barrier between the latter two subspecies. [13] The Niger River and the Dahomey Gap (a stretch of dry forest across Western Africa) have both been hypothesized as barriers that have separated the western and Nigeria-Cameroon chimpanzees.[14] However, not all of these geographic barriers are complete; researchers have recently recorded migration between eastern and central chimpanzee subspecies.[15][16] Sequencing of mitochondrial DNA has revealed that there are two major chimpanzee lineages; one clade is formed by the central and eastern chimpanzees, while western and Nigeria-Cameroon chimpanzees form the other clade.[17] Subspeciation is further enforced by variable susceptibility to specific viral infections.[18][19]Simian immunodeficiency virus (SIV), the precursor to the human immunodeficiency virus (HIV), is known to infect only P. t. troglodytes and P. t. schweinfurthii.[20][21] Chimpanzee subspecies have also shown specific behavioral differences, most notably that nut-cracking behavior only occurs in the west.[22][23]
Distinct Genetic Adaptations
Include some current research, with a second image.
Behavioral Adaptations
Contributions to Medicine and a Collective Understanding of Homo sapiens
Conclusion
Overall text length should be at least 1,000 words (before counting references), with at least 2 images. Include at least 5 references under Reference section.
References
- ↑ Bjork A, Liu W, Wertheim JO, Hahn BH, Worobey M. Evolutionary history of chimpanzees inferred from complete mitochondrial genomes. Mol Biol Evol. 2011;28(1):615–23.
- ↑ Raaum RL, Sterner KN, Noviello CM, Stewart CB, Disotell TR. Catarrhine primate divergence dates estimated from complete mitochondrial genomes: Concordance with fossil and nuclear DNA evidence. J Hum Evol. 2005;48(3):237–57.
- ↑ Stewart CB, Disotell TR. Primate evolution - In and out of Africa. Curr Biol. 1998;8(16):582–8.
- ↑ Raaum RL, Sterner KN, Noviello CM, Stewart CB, Disotell TR. Catarrhine primate divergence dates estimated from complete mitochondrial genomes: Concordance with fossil and nuclear DNA evidence. J Hum Evol. 2005;48(3):237–57.
- ↑ Bjork A, Liu W, Wertheim JO, Hahn BH, Worobey M. Evolutionary history of chimpanzees inferred from complete mitochondrial genomes. Mol Biol Evol. 2011;28(1):615–23.
- ↑ Raaum RL, Sterner KN, Noviello CM, Stewart CB, Disotell TR. Catarrhine primate divergence dates estimated from complete mitochondrial genomes: Concordance with fossil and nuclear DNA evidence. J Hum Evol. 2005;48(3):237–57.
- ↑ Stewart CB, Disotell TR. Primate evolution - In and out of Africa. Curr Biol. 1998;8(16):582–8.
- ↑ S, Hedges SB. A molecular timescale for vertebrate development. Nature [Internet. 1998;392:917–20.]
- ↑ Bjork A, Liu W, Wertheim JO, Hahn BH, Worobey M. Evolutionary history of chimpanzees inferred from complete mitochondrial genomes. Mol Biol Evol. 2011;28(1):615–23.
- ↑ Palaeontographica VA, Das P, Whiten a, Goodall J, Mcgrew WC, Nishida T, et al. Cultures in chimpanzees troglodytes ) have achieved long-term status across Africa , differ- lated 151 years of chimpanzee observation . This comprehensive analysis reveals patterns of variation that are far more extensive than have previously been docume. Nature. 1999;399(JUNE):15–8.
- ↑ Schmidt JM, De Manuel M, Marques-Bonet T, Castellano S, Andrés AM. The impact of genetic adaptation on chimpanzee subspecies differentiation. Vol. 15, PLoS Genetics. 2019. 1–32 p.
- ↑ Bowden R, MacFie TS, Myers S, Hellenthal G, Nerrienet E, Bontrop RE, et al. Genomic tools for evolution and conservation in the chimpanzee: Pan troglodytes ellioti is a genetically distinct population. PLoS Genet. 2012;8(3):1–10.
- ↑ Bjork A, Liu W, Wertheim JO, Hahn BH, Worobey M. Evolutionary history of chimpanzees inferred from complete mitochondrial genomes. Mol Biol Evol. 2011;28(1):615–23.
- ↑ Bjork A, Liu W, Wertheim JO, Hahn BH, Worobey M. Evolutionary history of chimpanzees inferred from complete mitochondrial genomes. Mol Biol Evol. 2011;28(1):615–23.
- ↑ Bjork A, Liu W, Wertheim JO, Hahn BH, Worobey M. Evolutionary history of chimpanzees inferred from complete mitochondrial genomes. Mol Biol Evol. 2011;28(1):615–23.
- ↑ Schmidt JM, De Manuel M, Marques-Bonet T, Castellano S, Andrés AM. The impact of genetic adaptation on chimpanzee subspecies differentiation. Vol. 15, PLoS Genetics. 2019. 1–32 p.
- ↑ Bjork A, Liu W, Wertheim JO, Hahn BH, Worobey M. Evolutionary history of chimpanzees inferred from complete mitochondrial genomes. Mol Biol Evol. 2011;28(1):615–23.
- ↑ Bjork A, Liu W, Wertheim JO, Hahn BH, Worobey M. Evolutionary history of chimpanzees inferred from complete mitochondrial genomes. Mol Biol Evol. 2011;28(1):615–23.
- ↑ Schmidt JM, De Manuel M, Marques-Bonet T, Castellano S, Andrés AM. The impact of genetic adaptation on chimpanzee subspecies differentiation. Vol. 15, PLoS Genetics. 2019. 1–32 p.
- ↑ Bjork A, Liu W, Wertheim JO, Hahn BH, Worobey M. Evolutionary history of chimpanzees inferred from complete mitochondrial genomes. Mol Biol Evol. 2011;28(1):615–23.
- ↑ Schmidt JM, De Manuel M, Marques-Bonet T, Castellano S, Andrés AM. The impact of genetic adaptation on chimpanzee subspecies differentiation. Vol. 15, PLoS Genetics. 2019. 1–32 p.
- ↑ Palaeontographica VA, Das P, Whiten a, Goodall J, Mcgrew WC, Nishida T, et al. Cultures in chimpanzees troglodytes ) have achieved long-term status across Africa , differ- lated 151 years of chimpanzee observation . This comprehensive analysis reveals patterns of variation that are far more extensive than have previously been docume. Nature. 1999;399(JUNE):15–8.
- ↑ Proffitt T, Haslam M, Mercader JF, Boesch C, Luncz L V. Revisiting Panda 100, the first archaeological chimpanzee nut-cracking site. J Hum Evol. 2018;124:117–39.
Edited by [Author Name], student of Joan Slonczewski for BIOL 116 Information in Living Systems, 2020, Kenyon College.
