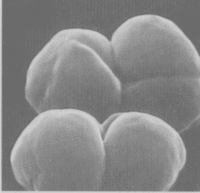
Methanosarcina barkeri
A Microbial Biorealm page on the genus Methanosarcina barkeri
Classification
Higher order taxa
|
NCBI: Archaea; Euryarchaeota; Methanomicrobia; Methanosarcinales; Methanosarcinaceae; Methanosarcina |
Species
|
NCBI: Methanosarcina barkeri |
Strains
Methanosarcina barkeri str. MS
Methanosarcina barkeri str. W
Methanosarcina barkeri str. 227
Description and significance
Describe the appearance, habitat, etc. of the organism, and why it is important enough to have its genome sequenced. Describe how and where it was isolated. Include a picture or two (with sources) if you can find them.
Genome structure
Describe the size and content of the genome. How many chromosomes? Circular or linear? Other interesting features? What is known about its sequence? Does it have any plasmids? Are they important to the organism's lifestyle?
Cell structure and metabolism
Methanosarcina barkeri is a methanogen and produces methane from a variety of carbon sources, including CO2, acetate, and methanol. It is also an anaerobe and is very oxygen sensitive, so all of its energy production pathways involve some sort of fermentation.
Ecology
Methanosarcina barkeri is an anaerobe has been isolated from mud samples in lakes and bogs. M. barkeri also lives in the rumen of cows where it helps digest organic matter for the cow. A USA Today article has reported that up to 17% of the world's atmospheric methane comes from cows, a large majority of which would come from M. barkeri. Because methane is a green house gas and can interfere with the ozone layer, this small organism may be partially responsible for two of the major environmental crises that we have face: thinning of the ozone layer and global warming.
Pathology
Methanosarcina barkeri is an archaea and therefore causes no known diseases.
Application to Biotechnology
As a methanogen, Methanosarcina barkeri has been looked at as a source of methane (natural gas) for use as an energy source.
Current Research
Longstaff, DG and Blight, SK and Zhang, L and Green-Church, KB and Krzycki, JA. "In vivo contextual requirements for UAG translation as pyrrolysine". Molecular Microbiology. 2007. 63-1, p. 229-241.
"Pyrrolysine and selenocysteine have infiltrated natural genetic codes via the translation of canonical stop codons. UGA translation as selenocysteine is absolutely dependent on message context. Here we describe the first experimental examination of contextual requirements for UAG translation as pyrrolysine. A hexahistidine-tagged Methanosarcina barkeri mtmB1 gene, encoding monomethylamine methyltransferase MtmB1, was introduced into Methanosarcina acetivorans. Host mtmB expression was minimized by growth on methanol and recombinant mtmB1 products monitored by anti-MtmB and anti-hexahistidine immunoblotting. UAG translation was not compromised, as recombinant MtmB1 was 1% of cellular protein with only trace UAG-terminated mtmB1 product detectable. Untranslated regions flanking mtmB1 were not required for UAG translation, but loss of a downstream pyrrolysine insertion sequence (PYLIS) significantly increased the UAG-termination product of mtmB1 and decreased the UAG-translation product, which nonetheless contained pyrrolysine. An in-frame UAG within a bacterial uidA transcript was translated in the methanogen as pyrrolysine with 20% efficiency, suggesting UAG translation in the absence of evolved context. However, predominant UAG-directed termination with enhancement of UAG translation by the PYLIS appears analogous to cis-acting elements for UGA translation as selenocysteine, although different mechanisms may underlie these recoding events."
Sato, T, Atomi, H, and Imanaka, T. "Archaeal Type III RuBisCOs Function in a Pathway for AMP Metabolism". Science. 2007. 315-5817, p. 1003.
"The type III ribulose-1,5-bisphosphate carboxylase-oxygenase (RuBisCO) present in the archaeon Thermococcus kodakaraensis' was found to participate in adenosine 5'-monophosphate (AMP) metabolism, a role that is distinct from that of classical RuBisCOs of the Calvin-Benson-Bassham cycle. Genes annotated as thymidine phosphorylase (deoA) and eucaryal translation initiation factor 2B (e2b2) were found to encode AMP phosphorylase and ribose-1,5-bisphosphate isomerase, respectively. These enzymes supplied the RuBisCO substrate, ribulose-1,5-bisphosphate, from AMP and phosphate. Archaea with type III RuBisCOs all harbor both DeoA and the corresponding E2b2 homologs. In this pathway, adenine was released from AMP and the phosphoribose moiety entered central-carbon metabolism."
Chu, HM and Andrew, HJW. "Enzyme-Substrate Interactions Revealed by the Crystal Structures of the Archaeal Sulfolobus PTP-Fold Phosphatase and its Phosphopeptide Complexes". PROTEINS: Structure, Function, and Bioinformatics. 2007. 66, p. 996-1003.
"The P-loop-containing protein phos-phatases are important regulators in signal transduction. These enzymes have structural and functional similarity with a conserved sequence of Dx(25-41)HCxxGxxR(T/S) essential for catalysis. The singular protein tyrosine phosphatase (PTP) from archaeal Sulfolobus solfataricus is one of the smallest known PTPs with extreme thermostability. Here, we report the crystal structure of this phosphatase and its complexes with two tyrosyl phosphopeptides A-(p)Y-R and N-K-(p)Y-G-N. The structure suggests the minimal structural motif of the PTP family, having two variable sequences inserted between the 2-3 and 3-4 strands, respectively. The phosphate of both phosphopeptide substrates is bound to the P-loop through several hydrogen bonds. Comparison of several phosphatase-substrate complexes revealed that Gln135 on the Q-loop has different modes of recognition toward phosphopeptides. The substrate specificity of SsoPTP is primarily localized at the phosphotyrosine, suggesting that this phosphatase may be a prototypical PTP."
Ambrogelly, A, Gundllapalli, S, Herring, S, Polycarpo, C, Frauer, C, and Soll, D. "Pyrrolysine is not hardwired for cotranslational insertion at UAG codons". Proceeds of the National Academy of Sciences. 2007. 104-9, p. 3141-3146.
"Pyrrolysine (Pyl), the 22nd naturally encoded amino acid, gets acylated to its distinctive UAG suppressor tRNA(Pyl) by the cognate pyrrolysyl-tRNA synthetase (PylRS). Here we determine the RNA elements required for recognition and aminoacylation of tRNA(Pyl) in vivo by using the Pyl analog N-epsilon-cyclopentyloxycarbonyl-l-lysine. Forty-two Methanosarcina barkeri tRNA(Pyl) variants were tested in Escherichia coli for suppression of the lac amber A24 mutation; then relevant tRNA(Pyl) mutants were selected to determine in vivo binding to M. barkeri PylRS in a yeast three-hybrid system and to measure in vitro tRNA(Pyl) aminoacylation. tRNA(Pyl) identity elements include the discriminator base, the first base pair of the acceptor stem, the T-stem base pair G51:C63, and the anticodon flanking nucleotides U33 and A37. Transplantation of the tRNA(Pyl) identity elements into the mitochondrial bovine tRNA(Ser) scaffold yielded chimeric tRNAs active both in vitro and in vivo. Because the anticodon is not important for PylRS recognition, a tRNA(Pyl) variant could be constructed that efficiently suppressed the lac opal U4 mutation in E. coli. These data suggest that tRNA(Pyl) variants may decode numerous codons and that tRNA(Pyl):PylRS is a fine orthogonal tRNA:synthetase pair that facilitated the late addition of Pyl to the genetic code."
Feist, AM, Scholten, JCM, Palsson, BO, Brockman, FJ, and Ideker, T. "Modeling methanogenesis with a genome-scale metabolic reconstruction of Methanosarcina barkeri". Molecular Systems Biology. 2006. vol. 2-1.
"Methanogenesis is a unique way of life for a group of archaea (methanogens) that generate energy by converting simple substrates such as acetate, methanol or H2/CO2 to methane. Because of this, methanogens serve as a key component of the carbon cycle by degrading low carbon molecules in a number of anaerobic environments. The methane they produce contributes to the greenhouse effect and is a potential source of renewable energy. In addition, some methanogens can form syntrophic relationships with other microorganisms, making them an interesting target for the study of interactions between different organisms. Although many pieces of methanogenic metabolism are understood, there are still many questions to be answered about the biochemistry of methanogenesis and how these pieces work together in the context of the whole organism. To address these questions, we reconstructed a genome-scale metabolic network for one of the most versatile methanogens, Methanosarcina barkeri, and analyzed the network to determine biochemical properties of key components and methanogenic metabolism as a whole."
References
Balch, WE, Fox, GE, Magrum, LJ, Woese, CR, and Wolfe, RS. "Methanogens: Reevaluation of a Unique Biological Group". Microbiological Reviews. June 1979. p. 260-296.
Maeder DL, Anderson I, Brettin TS, Bruce DC, Gilna P, Han CS, Lapidus A, Metcalf WW, Saunders E, Tapia R, Sowers KR. "The Methanosarcina barkeri Genome: Comparative Analysis with ...". Journal of Bacteriology. 2006. 188-22, p. 7922-7931.
Stadtman, TC and Barker, HA. "STUDIES ON THE METHANE FERMENTATION IX. The Origin of Methane in the Acetate and Methanol Fermentations by Methanosarcina". Journal of Bacteriology. 1951. 61-1, p. 81-86.
"Cutting cattle's methane emissions". USA Today (Magazine). June 01, 2002.
Atkins, JF and Gesteland, R. "The 22nd Amino Acid". Science. 2002. 296-5572, p. 1409-1410.
TIGR Comprehensive Microbial Resource: Methanosarcina barkeri
US Department of Energy Joint Genome Institute Organism Detail page for Methanosarcina barkeri
Edited by Ian Kerman, a student of Rachel Larsen at UCSD.