Acidithiobacillus albertensis
Classification
Bacteria; Proteobacteria; Acidithiobacillia; Acidithiobacillales; Acidithiobacillaceae; Acidithiobacillus
Species
|
NCBI: [1] |
Acidithiobacillus albertensis
Phylogenetic Debate
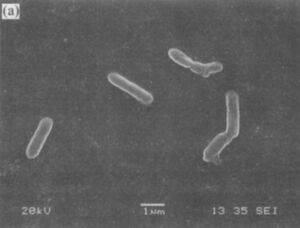
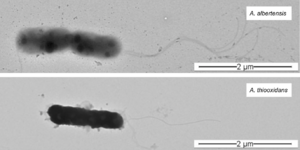
When A. albertensis was initially discovered in Alberta, Canada in 1983, it was reported as a new species within the genus Acidithiobacillus. Multiple studies supported this classification, demonstrating that A. albertensis could be distinguished from A. thiooxidans by its multiple flagella, more narrow pH range (2.5 - 5.5 for A. albertensis vs 0.5 - 5.5 for A. thiooxidans), and higher mol % G + C content (62 vs 52). However, in 2017, Castro et al. published a draft genome sequence of the DSM 14366 strain of A. albertensis based on 16s rRNA that challenged this classification.
They reported that A. albertensis had a 97.4% average nucleotide identity value when compared to A. thiooxidans and the two genomes had an in-silico DNA-DNA hybridization index of 82.9%. Values above 95% and 70%, respectively, for these two tests suggest that the genomes being tested belong to the same species. So, Castro et al. concluded that, “A. albertensis and A. thiooxidans probably comprise a single genospecies.” As of April 2024, NCBI classifies the two as separate species.
Description and Significance
Describe the appearance, habitat, etc. of the organism, and why you think it is important.
Genome Structure
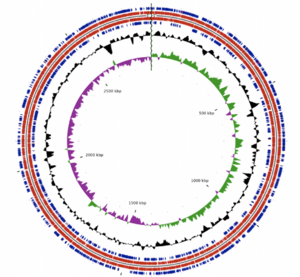
The A. albertensis genome is made of 3.5 Mbp organized into a single, circular chromosome. It contains 3203 genes, 3149 of which are predicted to be protein-coding. Castro et al. report that 63.4% of these genes have known functions, while the other 36.6% were hypothetical or unknown. Those with known functions include suites of genes for sulfur oxidation, carbon dioxide fixation, nitrate/nitrite assimilation and urea hydrolysis in addition to structural and housekeeping genes.
Plasmids and other mobile genetic elements (MGEs) make up 16.2% of the A. albertensis genome and are the main source of genetic difference between A. albertensis and A. thiooxidans. These MGEs encode genes related to sulfur oxidation, nitrate reduction and urea transport, among others. According to Castro et al. (2017): these genes "could confer adaptive advantages to A. albertensis over A. thiooxidans strains under nitrogen and oxygen limitation and/or under extremely low pH."
Flagella
Other genetic differences between A. albertensis and A. thiooxidans are due to non-synonymous single nucleotide polymorphisms in 9 genes related to flagellar structure and patterns. These mutations are likely why A. albertensis has multiple flagella while A. thiooxidans has only one. A. albertensis also has more genes associated with chemotaxis than A. thiooxidans, meaning that it may be able to detect and respond to chemical cues in the environment differently than A. thiooxidans, a particularly important skill for survival in ever-changing mining conditions.
Cell Structure, Metabolism and Life Cycle
Interesting features of cell structure; how it gains energy; what important molecules it produces.
Ecology and Pathogenesis
Habitat; symbiosis; biogeochemical significance; contributions to environment.
If relevant, how does this organism cause disease? Human, animal, plant hosts? Virulence factors, as well as patient symptoms.
References
[1] “Chalcopyrite.” Encyclopædia Britannica, Encyclopædia Britannica, inc., www.britannica.com/science/chalcopyrite.
[2] Castro, M., et al. “Draft genome sequence of the type strain of the sulfur-oxidizing acidophile, Acidithiobacillus albertensis (DSM 14366)”. Standards in genomic sciences. 2017. Volume 12 p. 77-84.
[3] Chen, J., Liu, Y., Diep, P., Mahadevan, R. “Genetic engineering of extremely acidophilic Acidithiobacillus species for biomining: Progress and perspectives”. Journal of Hazardous Materials. 2022. Volume 438 p. 129-247.
[4] Moya-Beltrán, A., Beard, S., Rojas-Villalobos, C. et al. “Genomic evolution of the class Acidithiobacillia: deep-branching Proteobacteria living in extreme acidic conditions”. ISME J. 2021. Volume 15 p. 3221–3238.
[5] Moya-Beltrán, A., Gajdosik, M., Rojas-Villalobos, C. et al. “Influence of mobile genetic elements and insertion sequences in long- and short-term adaptive processes of Acidithiobacillus ferrooxidans strains”. Sci Rep. 2023.Volume 13 p. 10876-10890.
[6] Rzhepishevska, O. “Physiology and Genetics of Acidithiobacillus Species: Applications for Biomining”. Umeå University. 2023.
[7] Schofield, E. “Acid Rock Drainage: More than Just a Mining Project Concern?” Barr Engineering Co., 2023. www.barr.com/Insights/Insights-Article/ArtMID/1344/ArticleID/380/Acid-rock-drainage-More-than-just-a-mining-project-concern.
[8] “Section 3 - Acidithiobacillus: Safety Assessment of Transgenic Organisms”. OECD. 2006. Volume 2.
[9] “Tetrathionate.” Wikipedia, Wikimedia Foundation, 27 Aug. 2023, en.wikipedia.org/wiki/Tetrathionate.
[10] Vardanyan, N.S., and A.K. Vardanyan. “New sulphur oxidizing bacteria isolated from bioleaching pulp of zinc and copper concentrates.” Universal Journal of Microbiology Research. 2014. Volume 2 p. 27–31.
[11] Xia, J., et al. “A new strain acidithiobacillus albertensis by-05 for bioleaching of metal sulfides ores.” Transactions of Nonferrous Metals Society of China. 2007. Volume 17 p. 168–175.
Author
Page authored by Ben Sauer and Rose Schnabel, students of Prof. Jay Lennon at IndianaUniversity.