Acidithiobacillus thiooxidans: Difference between revisions
No edit summary |
|||
| Line 1: | Line 1: | ||
{{Uncurated}} | |||
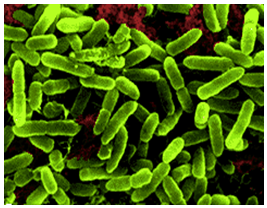
[[Image:acidithiobacillus_thiooxidans_green.png|thumbnail|320px|Figure 1. Electron microscope image of ''Acidithiobacillus thiooxidans''. From Khan et al., 2012.]] | [[Image:acidithiobacillus_thiooxidans_green.png|thumbnail|320px|Figure 1. Electron microscope image of ''Acidithiobacillus thiooxidans''. From Khan et al., 2012.]] | ||
Latest revision as of 12:34, 6 September 2017
Classification
Higher order taxa
Domain Bacteria; Phylum Proteobacteria; Class Gammaproteobacteria; Order Acidithiobacillales; Family Acidithiobacillaceae; Genus Acidithiobacillus
Species
Acidithiobacillus thiooxidans (type strain ATCC 19377)
Description and significance
Acidithiobacillus thiooxidans was once referred to as Thiobacillus thiooxidans before it was reclassified to a newly classified genus, Acidithiobacillus within the Gammaproteobacteria in 2000 (Kelly and Wood, 2000). The organism is a Gram-negative, rod-shaped bacteria that metabolizes sulfur as a method to obtain energy (Figure 1). A. thiooxidan is mesophilic with a temperature range of 10-37°C, the optimum temperatures ranging between 28-30°C. The acidophilic bacteria prefers environments with an optimum pH range of 2.0-3.0 (Kelly and Wood, 2000). The microbe is often found in cave biofilms and can contribute to the formation of cave systems (Jones, 2013).
16S Ribosomal RNA Gene Information
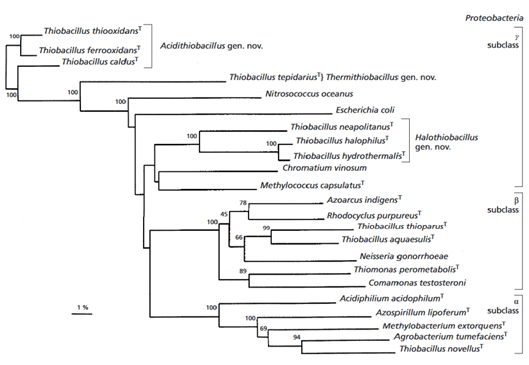
It was found that the species is related A. ferrooxidans at 99% and A. albertensis at 96% (Figure 2). It is also related to A. caldus ( Valdes et al., 2011).
Genome Structure
Whole-genome shotgun strategy was used to determine the complete draft sequence of the A. thiooxidans with the A. thiooxidans strain ATCC 19377 as the representative. The organism has a total length of 3,019,868 base pairs and 3,235 predicted proteins. It also has a GC of 53.1% ( Valdes et al., 2011). Sequence data can be found on the National Center for Biotechnology Information (NCBI) GenBank https://www.ncbi.nlm.nih.gov/genome/?term=thiobacillus+thiooxidans.
Cell structure and metabolism
A. thiooxidans utilizes a polar flagella to move around and does not accumulate sulfur, unlike other sulfur oxidizing microbes. The microbe does not produce spores. The average cell length is 1 µm or less with an average diameter of 0.5 µm (Waksman and Joffe, 1922).
Oxygen, carbon, and nitrogen are the essential nutrients required for metabolism. A. thiooxidans is an obligate aerobe that uses atmospheric oxygen as an electron acceptor and energy from inorganic sulfur, which produces sulfuric acid (H2SO4). The organism fixes atmospheric carbon dioxide to meet its need for carbon (Khan et al., 2012). Presence of carbonates, which turns the medium alkaline, stunts their growth. Small amounts of nitrogen are gathered from ammonium salts, such as sulfate, to aid in growth (Waksman and Joffe, 1922).
Ecology and Pathogenesis
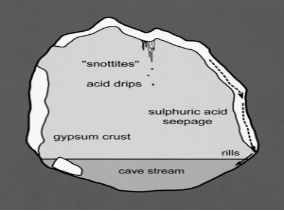
Acidithiobacillus thiooxidans are most commonly located in soil, sewer pipes and cave biofilms also known as “snottites”. Snottites are acidic biofilms that are formed by sulfur oxidizing microbes in caves rich in sulfide. A. thiooxidans in the biofilm produce sulfuric acid as a byproduct, which degrades the cave surfaces (Figure 3). This same process can occur in industrial pipes and cause corrosion. Distribution and morphology of the snottites are dependent on the abundance of nutrients essential for the microbial growth (Jones et al., 2012).
A. thiooxidans are able to inhabit environments where many other microbes cannot due to their high tolerance for Copper and Zinc. The organism is utilized in a mining technique called “bioleaching”, which involves extracting metals from their ores through the action of microbes. The technique uses the catalytic effect produced by the organism’s metabolism to accelerate the degradation of the sulfides. This process is applied to waste treatment and decontamination (Pathak et al., 2009). It has also been used in mining for over 20 years (Hansford, 2013).
A. thiooxidans are non pathogenic (Kelly and Wood, 2000).
Current Research
Not much is known about the abiotic and enzymatic components of reduced inorganic sulfur compound (RISC) oxidation for acidophilic microorganisms. A study was done to combine old RISC models and literature with various experiments in partial oxidation and abiotic reactions. This new model, including the organism’s biomass stoichiometry, would provide assistance in predicting the growth of A. thiooxidans. It could also aid studies in biohydrometallurgical, which is the extraction of metal from ore, and environmental situations (Fazzini et al., 2013).
References
Fazzini, Roberto A.B.; Cortes, Maria P.; Padilla, Leandro; Maturana, Daniel; Budinich, Marko; Maass, Alejandro; Parada, Pilar (2013). "Stoichiometric modeling of oxidation of reduced inorganic sulfur compounds (RISCs) in Acidithiobacillus thiooxidans". Biotechnology and Bioengineering. 110 (8): 2242–2251. doi:10.1002/bit.24875.
Hansford, G. S.; T. Vargas (2001). "Chemical and electrochemical basis of bioleaching processes". Hydrometallurgy. 59 (2–3): 135–145. doi:10.1016/S0304-386X(00)00166-3.
Hose, Louise D; James A. Pisarowicz (1999). "Cueva de Villa Luz, Tabasco, Mexico: Reconnaissance Study of an Active Sulfur Spring Cave and Ecosystem" (PDF). Journal of Cave and Karst Studies. 61 (1): 13–21.
Jones, Daniel S (2005). "Geomicrobiology of highly acidic, pendulous biofilms ("snottites") from the Frasassi Caves, Italy" (PDF). www.carleton.edu. Carleton College.
Jones, Daniel S; Heidi L Albrecht; Katherine S Dawson; Irene Schaperdoth; Katherine H Freeman; Yundan Pi; Ann Pearson; Jennifer L Macalady (2012). "Community genomic analysis of an extremely acidophilic sulfur-oxidizing biofilm". ISME Journal. 6 (1): 158–170. doi:10.1038/ismej.2011.75. PMC 3246232free to read. PMID 21716305.
Khan, Shahroz; Haq, Faizul; Hasan, Fariha; Saeed, Kausar; Ullah, Rahat (2012). “Growth and Biochemical Activities of Acidithiobacillus thiooxidans Collected from Black Shale”. Journal of Microbiology Research 2012, 2(4): 78-83 DOI: 10.5923/j.microbiology.20120204.03.
Kelly, Donovan P.; Wood, Ann P. (2000). "Reclassification of some species of Thiobacillus to the newly designated genera Acidithiobacillus gen. nov., Halothiobacillus gen. nov. and Thermithiobacillus gen. nov". International Journal of Systematic and Evolutionary Microbiology. 50: 511–516. doi:10.1099/00207713-50-2-511.
Pathak, Ashish; Dastidar, M.G.; Sreekrishnan, T.R. (2009). “Bioleaching of heavy metals from sewage sludge”. Journal of Environmental Management. Volume 90 (8): 2343–2353.
Valdes, Jorge; Francisco Ossandon; Raquel Quatrini; Mark Dopson; David S. Holmes (2011). "Draft genome sequence of the extremely acidophilic biomining bacterium Acidithiobacillus thiooxidans ATCC 19377 provides insights into the evolution of the Acidithiobacillusgenus". Journal of Bacteriology. 193 (24): 7003–7004. doi:10.1128/JB.06281-11. PMC 3232857free to read. PMID 22123759. Waksman, Selman A.; Joffe, J. S. (1922). "Microorganisms concerned in the oxidation of sulfur in the soil: II. Thiobacillus thiooxidans, a new sulfur-oxidizing organism isolated from the soil". Journal of Bacteriology. 7 (2): 239–256.
Author
Page created by Catherine List, Molly Norris, Lindsey Paul, and Blake Torchon, students of Dr. Hidetoshi Urakawa at Florida Gulf Coast University.