Bacillus cereus: Difference between revisions
Slonczewski (talk | contribs) |
Slonczewski (talk | contribs) |
||
| Line 122: | Line 122: | ||
'''8.4 ''B. cereus'' Endophthalmitis''' | '''8.4 ''B. cereus'' Endophthalmitis''' | ||
''B. cereus'' causes endophthalmitis which can lead to blindness. There is currently no universal treatment for this disease. Various research projects are being conducted by the Callegan Lab to understand its pathogenicity and virulence in order to develop better treatment and antibiotics. [http:// | ''B. cereus'' causes endophthalmitis which can lead to blindness. There is currently no universal treatment for this disease. Various research projects are being conducted by the Callegan Lab to understand its pathogenicity and virulence in order to develop better treatment and antibiotics. [http://calleganlab.org Link to Current Research Website] | ||
One current research project is studying the role of ''B. cereus'' proteases in endophthalmitis. It appears that strains without metalloproteases InhA and InhA2, immune inhibitors, were more virulent than the wild-type strain. Further studies are being conducted to determine why [31]. | One current research project is studying the role of ''B. cereus'' proteases in endophthalmitis. It appears that strains without metalloproteases InhA and InhA2, immune inhibitors, were more virulent than the wild-type strain. Further studies are being conducted to determine why [31]. | ||
Latest revision as of 19:53, 28 April 2017
A Microbial Biorealm page on the genus Bacillus cereus
Classification
===Higher order taxa===
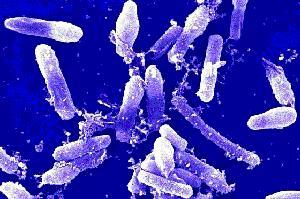
Domain: Bacteria
Phylum: Firmicutes
Class: Bacilli
Order: Bacillales
Family: Bacillaceae
Genus: Bacillus
Species Group: Bacillus cereus group
Species
|
NCBI: Taxonomy |
Bacillus cereus
Description and significance
Bacillus cereus is a large, 1 x 3-4 µm, Gram-positive, rod-shaped, endospore forming, facultative aerobic bacterium [2]. It was first successfully isolated in 1969 from a case of fatal pneumonia in a male patient and was cultured from the blood and pleural fluid [5]. 16s rRNA comparison reveals Bacillus cereus to be most related to Bacillus anthracis, the cause of anthrax, and Bacillus thuringiensis, an insect pathogen used as pesticide [3]. Although they have similar characteristics, they are distinguishable as B. cereus is most motile, B. thuringiensis produces crystal toxins, and B. anthracis is nonhemolytic [4].
B. cereus is mesophilic, growing optimally at temperatures between 20°C and 40°C, and is capable of adapting to a wide range of environmental conditions. It is distributed widely in nature and is commonly found in the soil as a saprophytic organism [2]. B. cereus is also a contributor to the microflora of insects, deriving nutrients from its host, and is found in the rhizosphere of some plants [2].
As a soil bacterium, B. cereus can spread easily to many types of foods such as plants, eggs, meat, and dairy products, and is known for causing 2-5 % of food-borne intoxications due to its secretion of emetic toxins and enterotoxins [4]. Food poisoning occurs when food is left without refrigeration for several hours before it is served. Remaining spores of contaminated food from heat treatment grow well after cooling and are the source of food poisoning.
In addition, Bacillus cereus is an opportunistic human pathogen and is occasionally associated with infections, causing periodontal diseases and other more serious infections [5]. Immunocompromised patients are susceptible to bacteremia, endocarditis, meningitis, pneumonia, and endophthalmitis [6]. Its potential to cause systemic infections are of current public health and biomedical concerns. Thus, the genome sequence of Bacillus cereus is significant in order to establish genetic background information for future investigations. Sequencing its genome is vital to expand understanding of its pathogenicity for treatment and for the development of antimicrobial drugs. Additionally, since Bacillus cereus strains are so genetically closely related to B. anthracis, genomic comparisons between the two species are important to the study of B. anthracis virulence.
Genome structure
3.1 Genome
B. cereus has a circular chromosome measuring 5,411,809 nt in length and was completely sequenced using the shotgun sequencing method [7]. The genome structure of B. cereus consists of 5481 genes, 5234 protein coding, 147 structural RNAs, and 5, 366 RNA operons [1]. An interesting gene cluster found within its genome encodes for arginine deiminase metabolic pathway. This cluster is predicted to have a role in its survival, enabling it to be resistant to acidic conditions in a similar manner as Streptococcus pyogenes [8]. Additionally, B. cereus has a nine gene urease gene cluster that encodes for proteins, blasticidin S deaminase, and an S-layer protein [8]. The urease enzyme increases its vigor in acidic conditions and is similar to the urease found in other bacteria that is required for colonization of the human stomach [8].
Genes present within the chromosome associated with B. cereus virulence include genes encoding for non-hemolytic enterotoxins, channel-forming type III hemolysins, phospholipase C, a perfringolysin O (listeriolysin O), and extracellular proteases [6]. The hbl operon, an RNA transcript of 5.5 kb, transcribes all three proteins of the hemolysin BL enterotoxins associated with food poisoning. These genes along with other genes encoding for metabolic enzymes, proteins involved in motility and chemotaxis, proteins involved in sporulation, and cellular transporters are all regulated by the plcR gene [9]. The plcR gene is also required for full virulence of B. cereus, and is often the target of antimicrobial drugs. Another gene found on its chromosome is the gerA gene which is essential for sporulation when nutrients are depleted, and is responsible for spore germination stimulated by L-alanine and ribosides [9]. It also has 18-23 genes that encode for peptide and amino acid ABC transporter-ATP binding proteins which suggests that amino acids, proteins, and peptids are preferred nutrient sources [9].
3.2 Plasmids
B. cereus has a diverse range of plasmids that vary in size from 5 to 500 kb and is known to have more than one plasmid with only a few that are associated with pathogenesis. B. cereus ATCC has a pXO1 plasmid that is found in B. anthracis [7]. It is avirulent because it does not have the portion that encodes for the toxin and regulatory proteins. B. cereus G9241 has a plasmid that is 99.6% identical to pXO1 plasmid from B. anthracis, but does not have the pXO2 plasmid which is required for full virulence. It also has a second plasmid that encodes for a capsule biosynthesis operon [7]. B. cereus ZK, a pathogenic strain, has five plasmids. Transposase genes are found in the two large plasmids which function in gene exchange between plasmids and chromosome [7]. The three smaller plasmids of the five function in identifying replication and mobilization proteins [7].
Cell structure and metabolism
4.1 Cell Structure
Bacillus cereus is a 1 x 3-4 µm, rod shaped, Gram- positive bacterium. Its cell structure consists of an inner membrane and a thick peptidoglycan which functions to maintain cell shape [10]. The polysaccharide portion makes up 50% percent of the cell wall and consists of a neutral polysaccharide composed of N-acetylglucosamine, N-acetylmannosamine (ManNac), N-acetylgalactosamine and glucose in a molar ratio of 4: 1: 1: 1 [11]. The acidic portion of the cell wall is characteristic in having a repeating tetrasaccharide unit [11]. 5% of the cell wall is made up of techoic acids consisting of N-acetylglucosamine, galactose, glycerol, and phosphorus in a molar ratio of 1: 1.4: 1: 1 [11]. The linkage between the polysaccharide and peptidoglycan is a muramic acid 6-phosphate [11]. The peptidoglycan of some B. cereus strains are unique with only a few oligomers present, the cross-linked muropeptides are dimmers, and many of the muropeptide lack the N-acetly group [12]. These distinguishing features affect cell surface charge which contributes to the attachment of an outer capsule or an S-layer in pathogenic strains.
Clinical isolates of B. cereus have a glycoprotein S-layer over its peptidoglycan which consists of proteinaceous paracrystalline arrays and covers the cell surface. The S-layer is involved in the virulence of B. cereus and functions to promote interactions with human polymorphonuclear leucocytes [13]. It also enables B. cereus to adhere to laminin, type I collagen, fibronectin, and fibrinogen of the epithelium, and thus has a role in increasing interaction between B.cereus and its host [13]. In addition, this proteinaceous layer enhances its resistance to radiation.
B. cereus is motile by means of flagella and exhibits two types of motility including swimming and swarming, depending on the enivronment. Single cells exhibit swimming motility by means of short flagellated rods [14]. On the other hand, swarming is a collective movement of swarm cells with flagellum that is observed to be three to four times longer, and also forty times more flagellated than single swimming cells [14]. 4.2 Spore Structure
B. cereus spore formation occurs when nutrients are scarce within the environment and germinate into vegetative cells once they are available [6]. Therefore, spore structure is important to the survival of this bacterium. B. cereus spores consist of an inner core surrounded by the inner membrane, and outer cortex surrounded by the outer membrane with an additional exterior coat [16]. The spore coat is made of proteins and small amounts of lipids and carbohydrates which contribute to its resistance to oxidizing agents and chemicals by blocking toxic molecules [15]. In addition, the outer spore structure allows them to be heat and γ-radiation resistant [15]. Spore germination is commonly in response to L-alanine which stimulates germination events including hydrations of spores, loss of Ca2+ and dipicolinic acid, and metabolism [15].
4.3 Metabolism
B. cereus is a facultative aerobe so it can utilize oxygen as a terminal electron accepter, but also has methods of anaerobic respiration as a mechanism of energy release. Whole genome sequencing revealed genes encoding for metabolic enzymes such as NADH dehydrogenases, succinate dehydrogenase, complex III, non-proton-pumping cytochrome bd quinol oxidases, and proton-pumping oxidases such as cytochrome c oxidase and cytochrome aa3 quinol oxidase [17].
In aerobic respiration, reducing equivalents produced from glycolysis and the Krebs cycle are reoxidized by the electron transport chain, creating a proton motive force and ATP by ATP synthase [17]. In anaerobic respiration, B. cereus utilizes fermentation to generate energy. Fermentation recycles NAD+ by reducing pyruvate and produces lactate and ethanol [17]. ATP is generated by substrate level phosphorylation.
B. cereus can metabolize a variety of compounds including carbohydrates, proteins, peptides and amino acids for growth and energy. Some of the major products produced from carbon sources such as sucrose or glucose during anaerobic respiration include L-lactate, acetate, formate, succinate, ethanol, and carbon dioxide [18]. During nitrate respiration, nitrate reductase converts nitrate into nitrite which is converted to ammonium by nitrite reductase [18].
Ecology
B. cereus interacts with other microorganisms in the rhizosphere, the region surrounding plant roots. Plants benefit from the presence of B. cereus since it is capable of inhibiting plant disease caused by protest pathogens and also enhances plant growth [23]. It naturally produces the antibiotics zwittermicin A. and Kanosamine which inhibit the growth of plant pathogen, oomycetes, certain fungi, and a few bacterial species [19]. The presence of B. cereus in the rhizosphere also increases nodulation of soybean plants by Bradyrhizobium japonicum [22]. In addition, Cytophaga-Flavobacterium group (CF) benefit from B. cereus by utilizing its peptidoglycan as a carbon and energy source. CF bacteria attain B. cereus peptidoglycan by hydrolyzing the outer layer [21]. The relationship between the two organisms is commensal since the growth of B.cereus is not affected by the presence of CF bacteria.
B. cereus is a also found in the gut microflora of invertebrates, and is an intestinal symbiont of arthropods where it exhibits filamentous growth in sow bugs, roaches, and termites [22]. Arthropods consume feces or soil with spores or cells of B. cereus. This intestinal stage of B. cereus, also known as the Arthromitus stage, involves the attachment of endospore fibers released from a parent cell [22]. After attachment of spores, they begin motile and filament growth stages and attach to the epithelium [22]. B. cereus cells are then defecated back into the soil where they can continue to grow.
As a ubiquitous bacterium, small amounts are consumed by humans from foods. Therefore, it is a contributor to the human intestinal microflora [23]. In addition B cereus is widely known to affect humans by causing food poisoning and infections as an opportunistic pathogen. Refer to pathology section for further elaboration.
Pathology
Bacillus cereus causes two types of food poisoning in humans including diarrhoeal syndrome and emetic syndrome. Food poisoning results from its production of enterotoxins in the gastrointestinal tract. The dosage of ingested B. cereus spores leading to diarrhoeal syndrome is 105–107 g 1 of ingested food, and 105–108 g 1 of ingested food for emetic syndrome [6]. Enterotoxins associated with diarrhoeal syndrome are unresistant to the acidic conditions of the stomach. However, the cereulide peptide toxin associated with emetic syndrome is more resistant to acidic conditions and remains active at 121 °C [6].
Virulence factors associated with diarrhoeal syndrome involve three enterotoxins including hemolysin BL (HBL), non-hemolytic enterotoxin (NHE), and cytotoxin K [24]. The main virulence factor of B. cereus is HBL which is made of the three proteins B, L1, and L2 [24]. Symptoms of diarrhoeal syndrome include watery diarrhea, abdominal cramps, and pain occurring 6-15 hours after ingestion which may last for twenty-four hours [4]. The emetic syndrome is caused by cereulide peptide toxin which is secreted during stationary phase [25]. This toxin has a ring structure, dodecadepsipeptide, which consists of four amino acids, repeating three times, and oxy acids [25]. Symptoms associated with emetic syndrome include nausea and vomiting within half an hour to six hours after ingestion of food and also lasts for about twenty-four hours [6].
Although B. cereus is commonly known to cause food-borne intoxications, it has been reported to cause local and systemic infections, as an opportunistic pathogen, especially among immunocompromised patients, newborns, and patients with surgical wounds [24]. B. cereus can cause ocular infections such as keratitis, endophthalmitis, and panophthalmitis [24]. The main virulence factor in B. cereus endophthalmitis is HBL which can result in the detachment of the retina and blindness. In addition, B. cereus can cause gangrene, bovine mastitis, pyogenic infections, cellulitis, infant death, septic meningitis, periodontal disease, lung abscesses, and endocarditis [6]. However, these infections are less common. Virulence factors associated with non gastrointestinal infections include hemolysins and phospholipase C. Hemolysin III causes erythrocyte lysis [6]. Phospholipase C causes tissue damage by stimulating degranulation of human neutrophil, and breaks down the subepithelial matrix affecting the healing of tissue in infections [24].
Application to Biotechnology
7.1 Biological Control Agent
Biological control agents are alternatives to chemical pesticides that are capable of suppressing plant pests and can also enhance plant growth. Most strains of Bacillus cereus produce toxins known to cause food-borne illnesses. However, there are strains that do not produce the HBL enterotoxin. Thus, antifungal compounds of Bacillus cereus strains have been developed as a useful biological control agent in the suppression of fungi and crop disease.
7.2 Bacillus cereus Strains
Bacillus cereus B4 is used as a pesticide to deter fungi from rotting seedling plants. This strain produces three types of metabolites that enable it to suppress certain plant diseases and promote plant growth. The metabolites include Kanosamine, 3, 4-dihydroxy benzoate, and 2 keto-4 methylthiobutyrate. The metabolite 3,4-DOHB improves crop strength and results in a healthier and larger root system [27].
Bacillus cereus DGA34 was isolated from the environment and is useful in fighting off fungal damping disease in crops. It naturally produces the antibiotic zwittermicin A. by fermentation which is found in the supernantant fluid of its culture medium. This antibiotic is effective in fighting a wide range of fungi and bacteria, and reduces symptoms of damping-off disease and root rot [26].
Bacillus cereus UW85 produces two antibiotic toxins including zwittermicin A. and antibiotic B. Bacillus cereus UW85 is used to protect alfalfa seeds from dampening off caused by Phytophthora medicaginis, cucumber fruits from Pythium aphanidermatum, peanuts from Sclerotinia minor, and tobacco seedlings from Phytophthora nicotianae [28].
These are just a few B. cereus strains with current United States patents as biocontrol agents for crop diseases. There are currently many Bacillus cereus strains that are still under review.
Current Research
8.1 Biofilms of B. cereus
The ability of B. cereus to form biofilms on surfaces can cause potential contamination problems within the food industry. Therefore, biofilm formation of several B. cereus strains are currently being studied to prevent potential food contamination and to ensure safety during production. In a recent study, microtiter assay and assays on stainless steel were completely or partially submerged in liquid in order to observe B. cereus biofilm formation. Since stainless steel is commonly used for pipes and tanks in the food industry, additional tests were conducted to investigate B. cereus biofilm formation from spores on stainless steal coupons. The results from both tests were similar. It appears that B. cereus biofilms preferentially form within the air-liquid interface. This tendency is due to the availability of oxygen in this region, causing bacterial movement toward oxygen. In addition, spore formation was more rapid in the suspension phase of biofilm formation suggesting that biofilms are a cavity for sporulation. The results show that B. cereus biofilms may develop within storage and piping systems when either partially filled or when liquid residues remain during production. In addition, increase in spore formation by B. cereus within biofilms can potentially cause recontamination and equipment failure during food production [29].
8.2 Effects of Porcine Bile on B. cereus
Resistance to bile is important to the survival of B. cereus within the small intestine where it can proliferate and release enterotoxins. On going research projects are being conducted to test B. cereus and factors that may affect its growth and release of enterotoxins once ingested. A recent study tested the effects of porcine bile (PB) on B. cereus and on its HBL enterotoxin in the small intestine by using intestine media with different food types. Different concentrations of porcine bile were added to the gastric media to simulate acid stress. Results show that the growth of B. cereus was affected by the type of food in the small intestine media which may be explained by the protective effect of different types of food against porcine bile. For instance, components of food such as fiber can bind to bile salts reducing their toxic effects on B. cereus. In addition, bile salts may be sequestered by food components that lower cholesterol levels. It was concluded that the tolerance of B. cereus to porcine bile and its ability to grow and produce toxins is dependent on the type of food and on bile concentrations in the small intestine. Additional studies are being conducted to test the effects of bile on B. cereus enterotoxin as well as the effects of different carbon sources on enterotoxin release [32].
8.3 Virulence Factors of B. cereus
B. cereus has been a growing and established opportunistic human pathogen. Therefore, current research is being conducted to understand its pathogenicity and virulence factors in order to find potential targets for antimicrobial drugs. In a recent study conducted, B. cereus fur gene, a transcriptional regulator that is responsible for bacterial iron uptake and metabolism, was shown to reduce virulence in the pathogenic fur mutant. Results in this investigation revealed a decrease in the regulation of iron uptake with three times more intercellular iron in the fur mutant than in the wild-type, resulting in a greater amount of oxidizing free radicals. The virulence of B. cereus fur mutant was measured in an insect infection which revealed the significance of iron metabolism regulation in bacterial pathogens by its reduced virulence. This recent study gives insight to the importance of the fur gene in the regulation of iron concentrations for cell growth, survival, and pathogenesis. The reduced virulence of B. cereus fur mutants in this experiment demonstrates the potential of the fur gene to be a good target for antimicrobial drugs as a conserved protein among pathogenic bacteria [30].
8.4 B. cereus Endophthalmitis
B. cereus causes endophthalmitis which can lead to blindness. There is currently no universal treatment for this disease. Various research projects are being conducted by the Callegan Lab to understand its pathogenicity and virulence in order to develop better treatment and antibiotics. Link to Current Research Website
One current research project is studying the role of B. cereus proteases in endophthalmitis. It appears that strains without metalloproteases InhA and InhA2, immune inhibitors, were more virulent than the wild-type strain. Further studies are being conducted to determine why [31].
Another ongoing research project involves the study of the plcR gene which regulates toxin production and has an important role in B. cereus endophthalmitis pathogenisis. The plcR mutant strain exhibited a reduction in virulence which resulted in 20% retained retinal function. The effects of blocking the peptide PapR as a potential form of treatment is being investigated [31].
Cell wall structures of B. cereus including peptidoglycan, S-layer, capsules, and techoic acids are being studied for their ability to cause intraocular inflammation. Recognition of these cell wall structures by retinal cells are being researched [31].
References
[1] "Bacillus cereus." NCBI website. Accessed on August 18, 2007.
[2] Vilain, S., Luo, Y., Hildreth, M., and Brozel, V. “Analysis of the Life Cycle of the Soil Saprophyte Bacillus cereus in Liquid Soil Extract and in Soil.” Applied Environmental Microbiology. 2006. Volume 72(7). p. 4970–4977.
[3] DelVecchio, V., Connolly, J., Alefantis, T., Walz, A., Quan, M., Patra, G., Ashton, J., Whittington, J., Chafin, R., Liang, X., Grewal, P., Khan, A., and Mujer C. “Proteomic Profiling and Identification of Immunodominant Spore Antigens of Bacillus anthracis, Bacillus cereus, and Bacillus thuringiensis.” Applied Environmental Microbiology. 2006. Volume 72(9). p. 6355–6363.
[4] “Bacillus cereus.” United States Food and Drug Administration, Center for food safety and applied nutrition (FDA). Accessed August 18, 2007.
[5] Hoffmaster, A., Hill, K., Gee, J., Marston, C., De, B., Popovic, T., Sue, D., Wilkins, P., Avashia, S., Drumgoole, R., Helma, C., Ticknor, L., Okinaka, R., and Jackson, J. “Characterization of Bacillus cereus Isolates Associated with Fatal Pneumonias: Strains Are Closely Related to Bacillus anthracis and Harbor B. anthracis Virulence.” Journal of Clinical Microbiology. 2006. Volume 44(9). p. 3352-3360.
[6] Wijnands, L., Dufrenne, J., Zwietering, M. H., and Leusden, F. “Spores from mesophilic Bacillus cereus strains germinate better and grow faster in simulated gastro-intestinal conditions than spores from psychrotrophic strains.” International Journal of Food Microbiology. 2006. Volume 112. Issue 2. p. 120-128.
[7] Rasko, D., Altherr, M., Han, C., and Ravel, J. “Genomics of the Bacillus cereus group of organisms.” FEMS Microbiology Reviews. 2005. Volume 29(2). p.303-329.
[8] Rasko, D., Ravel, J., Okstad, O. A., Helgason, E., Cer, R., Jiang, L., Shores, K. A., Fouts, D., Tourasse, N., Angiuoli, S., Kolonay, J., Nelson, W., Kolsto, A, Fraser, C., and Read, T. D. “The genome sequence of Bacillus cereus ATCC 10987 reveals metabolic adaptations and a large plasmid related to Bacillus anthracis pXO1” Nucleic Acids Research. 2004. Volume 32(3). p. 977–988.
[9] Han, C., Xie, g., Challacombe, J., Altherr, M., Bhotika, S., Bruce, D., Campbell, D., Campbell, M., Chen, J., Chertkov, O., Cleland, C., Dimitrijevic, M., Doggett, N., Fawcett, J., Glavina, T., Goodwin, L., Hill, K., Hitchcock, P., Jackson, P., Keim, P., Kewalramani, A., Longmire, J., Lucas, S., Malfatti, S., McMurry, K., Meincke, L. J., Misra, M., Moseman, B. L., Mundt, M., Munk, C., Okinaka, R. T., Parson-Quintana, B., Reilly, L. P., Richardson, P., Robinson, D. L., Rubin, E., Saunders, E., Tapia, R., Tesmer, J. G., Thayer, N., Thompson, L. S., Tice, H., Ticknor, L., Wills, P., Brettin, T., and Gilna, P. “Pathogenomic Sequence Analysis of Bacillus cereus and Bacillus thuringiensis Isolates Closely Related to Bacillus anthracis.” Journal of Bacteriology. 2006. Volume 188(900). p. 3382–3390.
[10] Ticknor, O., Kolsto, A., Hill, K., Keim. P., Laker, M., Tonks, M., and Jackson, P. “Fluorescent Amplified Fragment Length Polymorphism Analysis of Norwegian Bacillus cereus and Bacillus thuringiensis Soil Isolates.” Applied Environmental Microbiology. 2001. Volume 67(10). p. 4863–4873.
[11] Amano, K., Hazama, S., Akarari, Y., Ito, E. “Isolation and Characterization of Structural Components of Bacillus cereus AHU 1356 Cell Walls.” European Journal of Biochemistry. (1977). Volume 75 (2). p. 513–522.
[12] Severin, A., Tabei, K., Tomasz, A. “The structure of the cell wall peptidoglycan of Bacillus cereus RSVF1, a strain closely related to Bacillus anthracis.” Microbial Drug Resistance. 2004. Volume 10(2). p. 77-82.
[13] Mignot, T., Denis, B., Couture-Tosi, E., Kolsto, A., Mock, M., Fouet, A. “Distribution of S-layers on the surface of Bacillus cereus strains: phylogenetic origin and ecological pressure.” Environmental Microbiology. 2001. Volume 3(8). p. 493–501.
[14] Senesi, S., Celandroni, F., Salvetti, S., Beecher, D., Wong, A., and Ghelardi, A. “Swarming motility in Bacillus cereus and characterization of a fliY mutant impaired in swarm cell differentiation.” Microbiology. 2002. Volume 148. p. 1785-1794.
[15] Pol, I., van Arendonk, W., Mastwijk, H., Krommer, J., Smid, E., and Moezelaar R. “Sensitivities of Germinating Spores and Carvacrol-Adapted Vegetative Cells and Spores of Bacillus cereus to Nisin and Pulsed-Electric-Field Treatment.” Applied Environmental Microbiology. 2001. Volume 67(4). p. 1693–1699.
[16] Kutima, P., and Foegeding, P. “Involvement of the spore coat in germination of Bacillus cereus T spores.” Applied Environmental Microbiology. 1987. Volume 53(1). p.47–52.
[17] Duport, C., Zigha, A., Rosenfeld, E., and Schmitt, P. “Control of Enterotoxin Gene Expression in Bacillus cereus F4430/73 Involves the Redox-Sensitive ResDE Signal Transduction System.” Journal of Bacteriology. 2006. Volume 188. p. 6640–6651.
[18] Mols, M., de Been, M., Zwietering, M., Moezelaar, R., Abee, T. “Metabolic capacity of Bacillus cereus strains ATCC 14579 and ATCC 10987 interlinked with comparative genomics.” Environmental Microbiology. 2007. (Online Early Articles).
[19] Silo-Suh, L., Lethbridge, B., Raffel, S J, He, H., Clardy, J., and Handelsman, J. “Biological activities of two fungistatic antibiotics produced by Bacillus cereus UW85.” Applied Environmental Microbiology. 1994. Volume 60(6). p. 2023–2030.
[20] Halverson, L J, and Handelsman J. “Enhancement of soybean nodulation by Bacillus cereus UW85 in the field and in a growth chamber.” Applied Environmental Microbiology. 1991. Volume 57(9). p. 2767–2770.
[21] Peterson, S., Dunn, A., Klimowicz, A., and Handelsman, J. “Peptidoglycan from Bacillus cereus Mediates Commensalism with Rhizosphere Bacteria from the Cytophaga-Flavobacterium Group.” Applied Environmental Microbiology. Volume 72(8). p. 5421–5427.
[22] Margulis, L., Jorgensen, J, Dolan, S., Kolchinsky, R., Rainey, F., and Shyh-Ching. “The Arthromitus stage of Bacillus cereus: Intestinal symbionts of animals.” Proceedings of the National Science Academy U S A. 1998. Volume 3; 95(3). p. 1236–1241.
[23] Jensen, G., Hansen, B., Eilenberg, J., Mahillon, J. “The hidden lifestyles of Bacillus cereus and relatives.” Environmental Microbiology. 2003. Volume 5(8). p. 631–640.
[24] Kotiranta, A., Lounatmaa, K., and Haapasalo, M. “Epidemiology and pathogenesis of Bacillus cereus infections.” Microbes and Infections. 2000. Volume 2, Issue 2. p. 189-198
[25] Granum, P., Lund, T. “Bacillus cereus and its food poisoning toxins.” FEMS Microbiology Letters. 1997. Volume 157 (2). p. 223–228.
[26] Handelsman, J., Jacobson, L., Stabb, E. “Bacillus cereus strain DGA34; United States Patent 5736382.” Official gazette of the United States Patent and Trademark Office. 1998. Accessed August 13, 2007.
[27] Sunaina, V. “Bacterial metabolites from Bacillus cereus B4 responsible for potato plant growth.” Journal of the Indian Potato Association. 2005. Volume 32 (3-4). p. 187-188.
[28] Silo-Suh, L., Lethbridge, B., Raffel, S., He, H., Clardy, J., and Handelsman, J. “Biological activities of two fungistatic antibiotics produced by Bacillus cereus UW85.” Applied Environmental Microbiology. 1994. Volume 60(6) p. 2023–2030.
[29] Wijman, J., Leeuw, P., Moezelaar, R., Zwietering, M., and Abee, T. “Air-Liquid Interface Biofilms of Bacillus cereus: Formation, Sporulation, and Dispersion.” Applied Environmental Microbiology. 2007. Volume 73(5). p. 1481–1488.
[30] Harvie, D., Vilchez, D., Steggles, J., Ellar, D. “Bacillus cereus Fur regulates iron metabolism and is required for full virulence.” Microbiology. 2005. Volume 151(Pt 2). p.569-77.
[31] Callegan, M., Novsad, Hunt. Callegan Lab; Current research projects. Accessed August 20, 2007.
[32] Clavel, T., Carlin, F., Dargaignaratz, C., Lairon, D., Nguyen-The, C., Schmitt, P. "Effects of porcine bile on survival of Bacillus cereus vegetative cells and Haemolysin BL enterotoxin production in reconstituted human small intestine media." Journal of Applied Microbiology. 2007. Online Early Articles.
Edited by Jacqueline Nguyen student of Rachel Larsen
Edited by KLB
