Citrobacter freundii: Difference between revisions
| Line 28: | Line 28: | ||
==Genome structure== | ==Genome structure== | ||
The complete genome of this microbe has not been sequenced yet since it is so large, although some individual strains and plasmids of the microbe have been sequenced. The most prominent one is the plasmid pCTX-M#3. Its sequence was completed on January 6, 2005. It is a circular DNA plasmid and it is 89,468 nucleotide base pairs long. The length of the plasmid is 0.089468 (Mbp). It is composed of 51.0 % GC content, and encodes 105 proteins. | |||
The C. freundii OS60 AmpC β-lactamase gene has also been sequenced and it is composed of 1197 nucleotides. It encodes a 380 amino acid long precursor with a 19-residue signal peptide. The mature protein encoded by this gene has molecular mass of 39 781 Da. 77% of the amino acid positions hold identical to residues in the E. coli K12 chromosomal AmpC β-lactamases. | |||
It also has been disocovered that for the first time in the Enterobacteriaceae family that a gene encoding L_methionine γ_lyase (MGL)is present in the genome of Citrobacter freundii. . The hybrid plasmid pUCmgl obtained from the C. freundii genomic library contains an EcoRI insert which is 3000 bp long. This gene which ensures the appearance of MGL activity. The nucleotide sequence of the C. Freundii EcoRI fragment contains two open reading frames. The first frame (the megL gene) encodes a protein of 398 amino acid residues that has sequence homology with MGLs from different sources. The second frame encodes a protein with sequence homology with proteins belonging to the family of permeases. | |||
==Cell structure and metabolism== | ==Cell structure and metabolism== | ||
Revision as of 02:53, 27 August 2007
A Microbial Biorealm page on the genus Citrobacter freundii
Classification
Higher order taxa
Bacteria; Proteobacteria; Gammaproteobacteria; Enterobacteriales; Enterobacteriaceae; Citrobacter
Species
|
NCBI: Taxonomy |
Citrobacter Freundii
Description and significance
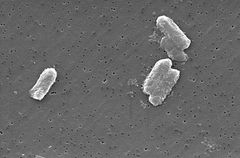
The Citrobacter species, including Citrobacter freundii, are aerobic gram-negative bacilli. Citrobacter freundii is a long rod-shaped bacteria typically 1-5 μm in length. Most have many flagella used to move about, but a few are non-motile. Its habitat includes the enivronment (soil, water, sewage), food, and the intestinal tracts of animals and humans. It belongs to the family of Enterobacteriaceae.
As an opportunistic pathogen, C.freundii is often the cause of significant opportunistic infections. C. Freundii is known to cause a wide variety of nosocomial infections of the respiratory tract, urinary tract, blood and several other normally sterile sites. It represents approximately 29% of all opportunistic infections. Therefore, one of the main reasons the genome of C. Freundii is being sequenced is in order to find antibiotics that can fight these opportunistic infections.
The Citrobacter genus was discovered in 1932 by Werkman and Gillen. Cultures of C. freundii were isolated and identified in the same year from soil extracts.
Genome structure
The complete genome of this microbe has not been sequenced yet since it is so large, although some individual strains and plasmids of the microbe have been sequenced. The most prominent one is the plasmid pCTX-M#3. Its sequence was completed on January 6, 2005. It is a circular DNA plasmid and it is 89,468 nucleotide base pairs long. The length of the plasmid is 0.089468 (Mbp). It is composed of 51.0 % GC content, and encodes 105 proteins.
The C. freundii OS60 AmpC β-lactamase gene has also been sequenced and it is composed of 1197 nucleotides. It encodes a 380 amino acid long precursor with a 19-residue signal peptide. The mature protein encoded by this gene has molecular mass of 39 781 Da. 77% of the amino acid positions hold identical to residues in the E. coli K12 chromosomal AmpC β-lactamases.
It also has been disocovered that for the first time in the Enterobacteriaceae family that a gene encoding L_methionine γ_lyase (MGL)is present in the genome of Citrobacter freundii. . The hybrid plasmid pUCmgl obtained from the C. freundii genomic library contains an EcoRI insert which is 3000 bp long. This gene which ensures the appearance of MGL activity. The nucleotide sequence of the C. Freundii EcoRI fragment contains two open reading frames. The first frame (the megL gene) encodes a protein of 398 amino acid residues that has sequence homology with MGLs from different sources. The second frame encodes a protein with sequence homology with proteins belonging to the family of permeases.
Cell structure and metabolism
Describe any interesting features and/or cell structures; how it gains energy; what important molecules it produces.
Ecology
Describe any interactions with other organisms (included eukaryotes), contributions to the environment, effect on environment, etc.
Pathology
How does this organism cause disease? Human, animal, plant hosts? Virulence factors, as well as patient symptoms.
Application to Biotechnology
Does this organism produce any useful compounds or enzymes? What are they and how are they used?
Current Research
Enter summaries of the most recent research here--at least three required
References
Edited by Sumaira Akbarzada, student of Rachel Larsen