Dehalobacter
A Microbial Biorealm page on the genus Dehalobacter
Classification
Higher order taxa:
Bacteria; Firmicutes; Clostridia; Clostridiales; Peptococcaceae; Dehalobacter
Species:
Dehalobacter restrictus PER K23, Dehalobacter sp. E1, Dehalobacter sp. TCA1, Dehalobacter sp. MS Dehalobacter sp. WL
|
NCBI: Taxonomy Genome |
Description and Significance
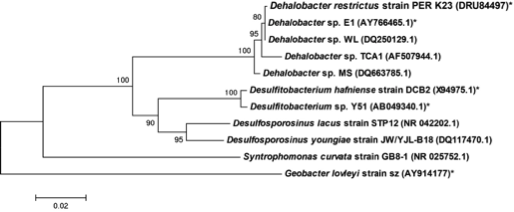
Dehalobacter restrictus is a rod shaped gram-negative obligate anaerobe found in deep soil sediments and groundwater aquifers and is phylogenetically related to several halorespiring genera like Desulfitobacterium and Desulfosporlosinus. It is a typical representative of Dehalobacter spp., which are capable of reductive dechlorination of chlorinated ethenes and chlorinated ethanes (1, 2) and fermentative dehalogenation of Dichloromethane (DCM) (3). These metabolic pathways make them relevant to bioremediation of ground water aquifers contaminated with chlorinated solvents (4). To date, several strains of Dehalobacter have been isolated (1, 3, 5-7)."
Genome Structure
The whole genome of Dehalobacter restrictus PER- K23 was recently sequenced revealing critical insights into phylogeny and metabolic capabilities of the genus. The 2,943 kbp long genome contains 2,826 protein coding and 82 RNA genes, including 5 16S rRNA genes. Interestingly, the genome contains 25 predicted rdase genes, the majority of which appear to be full length. The rdase genes are mainly located in two clusters, suggesting a much larger potential for organohalide respiration than previously anticipated (11).
Cell Structure
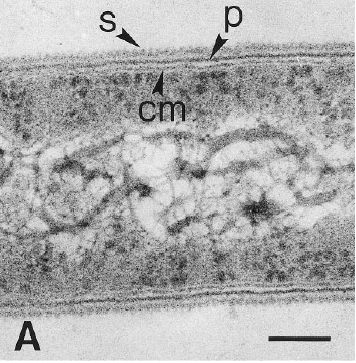
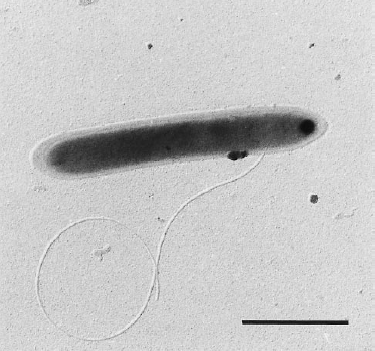
Dehalobacter restrictus cells are rod shaped, 2-3 µm in length and 0.3-0.5 µm in diameter with tapering ends. There is a single lateral flagellum at one end indicating that the cells are motile. The cells do not form spores under substrate and temperature induced stress. Cell wall stains gram negative and is composed of peptidoglycan type A3ϒ enveloped by a layer of proteinaceous surface layer (S – layer) (5)
Metabolism
Dehalobacter restrictus has unique metabolic pathways that make them vital for bioremediation of several key recalcitrant environmental contaminants (4).
Reductive dehalogenation
Isolation and Cultivation
D. restrictus is isolated from a laboratory system that was being fed with tetrachloroethene.