Desulfovibrio: Difference between revisions
No edit summary |
|||
| Line 1: | Line 1: | ||
{{Biorealm}} | |||
{| | {| | ||
| width="152" height="69" bgcolor="#FFDF95" | | | width="152" height="69" bgcolor="#FFDF95" | | ||
Revision as of 14:03, 14 August 2006
A Microbial Biorealm page on the Desulfovibrio
|
NCBI: |
Classification
Higher order taxa:
Bacteria; Proteobacteria; delta/epsilon subdivisions; Deltaproteobacteria; Desulfovibrionales; Desulfovibrionaceae
Species:
Desulfomonas oviles; D. acrylicus; D. aerotolerans; D. aespoeensis; D. africanus; D. alaskensis; D. alcoholovorans; D. alkalitolerans; D. aminophilus; D. arcticus; D. baarsii; D. bastinii; D. brasiliensis; D. burkinensis; D. caledoniensis; D. capillatus; D. carbinolicus; D. carbinoliphilus; D. cavernae; D. cuneatus; D. dechloracetivorans; D. desulfuricans; D. fairfieldensis; D. ferrireducens; D. ferrophilus; D. frigidus; D. fructosovorans; D. gabonensis; D. giganteus; D. gigas; D. gracilis; D. halophilus; D. hydrothermalis; D. indonesiensis; D. inopinatus; D. intestinalis; D. longreachensis; D. longus; D. magneticus; D. mexicanus; D. multispirans; D. oryzae; D. oxyclinae; D. piger; D. profundus; D. putealis; D. salexigens; D. senezii; D. simplex; D. sulfodismutans; D. termitidis; D. vietnamensis; D. vulgaris; D. zosterae; D. sp.
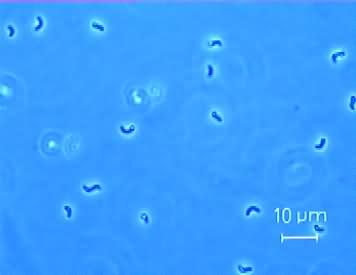
Description and Significance
This rod-shaped bacterium is identified as anaerobically growing, motile organism. Desulfovibrio is known as a sulfate reducing bacterium, which has put it to the forefront of biological research. Because of its metal corroding ability, which has consequently led to numerous health and safety concerns in industry, a means to neutralize and better understand Desulfovibrio species is the goal of research on Desulfovibrio (ITQB). The organism however also shows potential for bioremediation, in that it may undergo anaerobic conversion of pollutants in the soil.
Genome Structure
Because of Desulfovibrio's historical importance, two strains have already been genomically sequenced and one is currenlty in progress. These strains include Desulfovibrio desulfuricans G20 (completed), Desulfovibrio vulgaris subsp. vulgaris str. Hildenborough (completed), and Desulfovibrio magneticus (in progess), which were sequenced by DOE Joint Genome Institute, TIGR, and NITE, respectively. Both of the completely sequenced genomes showed Desulfovibrio to have one chromosome and measure over 3 Mbp in length. Both sequencings also found the number of proteins to be above 3000.
Cell Structure and Metabolism
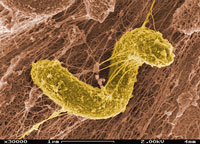
Desulfovibrio cells typically grow anaerobically, but certain strains have been identified growing in the presence of oxygen. Species of Desulfovibrio are traditionally Gram-negative, non-spore forming, and about 0.7 um in cell diameter. Desulfovibrio is known for its flexbility in response to the extended amount of electron acceptors it utilizes including sulfate, sulfur, nitrate, and nitrite among others (NCBI).
Species of Desulfovibrio have long been the subject of interest since they were first indentified as bioremediators. With the ability to reduce several toxic metals such as uranium (VI), chromium (VI) and iron (III), Desulfovibrio has the potential to be both quite useful and harmful (Cabrera).
Ecology
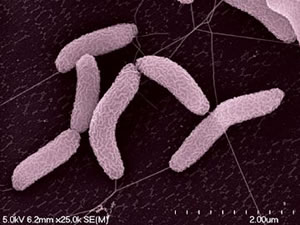
Desulfovibrio strains have been found in a variety of habitats, including soil, the intestines and feces of animals, and both salinic and fresh water. Desulfovibrio desulfuricans strain G20, which was found at an oil well corrosion site (2can). Desulfovibrio cells have mesophillic growth patterns with an ideal growth rate between the temperatures of twenty-five to fourty degrees Celcius (NCBI).
References
2can. 2can Genomes. Bacteria Genomes: Desulfovibrio vulgaris.
NCBI. Genome Project > Desulfovibrio desulfuricans G20 project at DOE Joint Genome Institute.
