Enterococcus faecalis: Difference between revisions
No edit summary |
|||
| Line 23: | Line 23: | ||
==Genome structure== | ==Genome structure== | ||
[[Image:new.png|frame|Genome of ''E. faecalis'' strian v583]] | [[Image:new.png|frame|Genome of ''E. faecalis'' strian v583 | ||
Stothard P, Van Domselaar G, Shrivastava S, Guo A, O'Neill B, Cruz J, Ellison M, Wishart DS (2005) BacMap: an interactive picture atlas of annotated bacterial genomes. Nucleic Acids Res 33:D317-D320]] | |||
Due to many public health dangers, the genome sequence data from a strain of Enterococcus was necessary and would signal an important advance in the study of bacterial pathogens. The strain chosen for genome DNA sequencing was E. faecalis V583, the first vancomycin-resistant isolate in the United States. The genome of strain V583 was sequenced by The Institute for Genome Research (TIGR). | Due to many public health dangers, the genome sequence data from a strain of Enterococcus was necessary and would signal an important advance in the study of bacterial pathogens. The strain chosen for genome DNA sequencing was E. faecalis V583, the first vancomycin-resistant isolate in the United States. The genome of strain V583 was sequenced by The Institute for Genome Research (TIGR). | ||
Revision as of 04:49, 2 May 2007
Classification
Higher order taxa
Bacteria; Firmicutes; Bacilli; Lactobacillales; Enterococcaceae; Enterococcus
Genus
Enterococcus faecalis
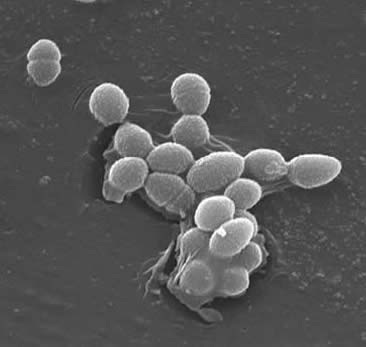
Description and significance
Enterococcus are gram-positive cocci that can survive harsh conditions in nature. They can be found in soil, water, and plants. Some strains are used for the manufacture of foods whereas others are the cause of serious human and animal infections (i.e. they are known to colonize the gastrointestinal and gential tracts of humans). They are associated with both community and hospitial acquired infections. Enterocci can grow at a temperature range of 10 to 42°C and in environments with broad pH values. Some are known to be motile.While there are over 15 species of the Enterococcus genus, 80-90% of clinical isolates are E. faecalis. Enterocci typically form short chains or are arranged in pairs. However, under certain growth conditions, they elongate and appear coocobacillary. In general, enterococci are alpha-hemlytic. Some possess the group D Lancefield antigen and can be detected using monoclonal antibody-based agglutination tests. Enterococci are typically catalase negative, and are anaerobic. They are able to grow in 6.5% NaCl, can hydrolyze esculin in the presence of 40% bile salts and are pyrrolidonyl arylamidase and leucine arylamidase positive. Enterococci have proven to present a therapeutic challenge because of their resistance to many antimicrobial drugs, including cell-wall active agents, aminoglycosides, penicillin and ampicillin, and vancomycin. In the last decade, vancomycin-resistnat enerococci have become a major challenge. The enterococci have the capacity to acquire a wide variety if antimicrobial resistance factors which present serious problems in the management of patients with enteroccoal infections. In general, entercoccal isolates with lowered susceptibility to vancomycin can be categorized as vanA, vanB, and vanC. vanA and vanB pose the greatest threat because they are the most resistant and the resistance genes are carried on a plasmid. Since the resistance genes are carried on a plasmid they readily transferable. E. faecalis strains are categorized as vanA isolates. E. faecalis are also resistant to teicoplanin. Enterococcal strains that are vancomycin dependent have been reported, but are rare and are less common that vancomycin-resistant strains.
Genome structure
Due to many public health dangers, the genome sequence data from a strain of Enterococcus was necessary and would signal an important advance in the study of bacterial pathogens. The strain chosen for genome DNA sequencing was E. faecalis V583, the first vancomycin-resistant isolate in the United States. The genome of strain V583 was sequenced by The Institute for Genome Research (TIGR). Strain V583 contains four DNA molecules: the main 3,218,030 base pair bacterial chromosome and three circular plasmids. The chromosome contains about 3,500 open reading frames (ORFs), about 1/3 of these ORFs have no assignable function. The analysis of the genes harbored in the enterococcal genome shows, as was predicted by its ability to survive a wide range of environmental conditions, that this organism is metabolically diverse and contains a wide range of regulatory systems. The three plasmids are circular DNA molecules identified as Plasmid-1, Plasmid-2, and Plasmid-3. Plasmid-1 contains 66,320bp, Plasmid-2 contains 17,963bp, and Plasmid-3 contains 57,660bp. The plasmids encode a number of genes, including, transposases, multidrug resistance proteins, and a ppGpp-regulated growth inhibitor. The average G+C composition of the E.faecalis chromosome is 37.38%. Since the DNA molecule is so large, regional deviations from the average occur. One of these locations is the large segment associated with the vancomycin resistance gene cluster positioned near 2.22 Mb, showing a large increase in percent G+C content. These differences associated with antibiotic resistance or virulence suggested the acquisition of genetic material from a frogein species through horizontal transfer. Whether the transfers are responsible for the variations in DNA composition is still unknown. The information contained in the genome of E. faecalis V583 will greatly aid the understanding of how the organism has adapted to be a versatile human pathogen. Using comparative genomics, the role of the different regulatory elements can be better understood in how they respond to various environmental stresses and in the expression of potential virulence factors. More studies like these will suggest new or alternate vaccines to bacterial infections caused by the Enterococci.
The TIGR contains a complete list of genes for the E. faecalis chromosomes.
Cell structure and metabolism
The Enterococci inhabit harsh environments, like the intestinal tracts of humans and animals. Growth under these hostile conditions requires that “E. faecalis have a metabolism that is flexible.” E. faecalis are capable of not only fermentation to produce latic acid but also can “catabolize a spectrum of energy sources” from carbohydrates, glycerol, lactate, malate, cirtrate, diamino acids and many alpha-keto acids.
It has been shown that under selected growth conditions E. faecalis can enhance growth through oxidative phosphorylation using a “proton motive force established by electron transport.” A consequence of “nascent respiration is production of potent oxidants” like superoxide and hydrogen peroxide, oxidative stress the E. faecalis can tolerate. The tolerance of this stress, combined with other severe growth conditions, allows the E. faecalis to grow at 10 to 45C, in bile salts, and at extremely low and high pHs. In addition, “E. faecalis resist azide, detergents, heavy metals, and ethanol.”
The capacity to utilize varied sugar sources likely allows enterococcal to “live in diverse environments, especially in the intestine where nutrients are limited.
In the intestine, enterococci derive most of their energy from the fermentation of nonabsorbed sugars. E. faecalis can also get energy by degrading “mucins, a carbohydrate that is heavily glycosylated and produced by intestinal goblet cells.”
Describe any interesting features and/or cell structures; how it gains energy; what important molecules it produces.
Ecology
Describe any interactions with other organisms (included eukaryotes), contributions to the environment, effect on environment, etc.
Pathology
Enterococci have emerged in the last part of the 20th century as a major cause of nosocomial infections, and within this group Enterococcus faecalis causes the majority of human enterococcal infections. These infections may be local or systematic and include uninary tract and abdominal infections, wound infections, bacteremia, and endocarditis. Enterococci are part of the many taxa that make up the normal human intestinal flora and are capable of surviving numerous environmental challenges such as temperature extremes and the presence of bile salts. Their acquisition of resistance to multiple antibiotics has made these bacteria a major health problem. The National Nosocomial Infection Surveillance (NNIS) system has reported a rapid rise in the incidence of infection and colonization with vancomycin-resistant enterococci (VRE) since 1989. This poses serious health problems, which include the lack of available antimicrobial therapy for VRE infections, because most VRE strains harbor resistance to multiple antibiotics (e.g. aminoglyscoides and ampicillin) and vancomycin is often the antibiotic of last resort.
The transfer of vanocmycin resistant genes from VRE to other gram-positive pathogens is a serious public health concern. Thus, the acquisiton of the genome sequence data from a strain of Enterococcus signals an important advance in the study of bacterial pathogens.
How does this organism cause disease? Human, animal, plant hosts? Virulence factors, as well as patient symptoms.
Application to Biotechnology
Does this organism produce any useful compounds or enzymes? What are they and how are they used?
Current Research
Enter summaries of the most recent research here--at least three required
References
De la Maza, Luis M., Marie T. Pezzlo, and Janet T. Shigei. Color Atlas of Medical Bacteriology. Washington, DC: American Society for Microbiology, 2004.
Edited by Richard A. Martinez of UC San Diego, student of Rachel Larsen.