Enterovirus
Baltimore Classification
Higher order taxa
Virus; Picornaviridae (Family); Enterovirus
Species
Poliovirus
Description and Significance
Poliovirus is the etiologic agent of poliomyelitis, a human disease of the central nervous system. The virus initiates infection by attaching to specific cell surface molecules or receptors. The poliovirus receptor is CD 155 and the normal function of this molecule on the host cell is not known yet.
There are three types of poliovirus and each type has many strains. After entering through the mouth, the virus multiplies in the throat and gastrointestinal tract, then moves into the bloodstream and is carried to the central nervous system. It then replicates and destroys the motor neuron cells, impairing normal functioning of the muscles that control swallowing, circulation, respiration, and the trunk, arms and legs.
When poliovirus encounters the nerve cells, the virus particle attaches itself to the protruding receptors-- a protruding protein structure on the surface of human nerve cells-- and causes infection. Once inside the cell, the virus hijacks the cell's assembly process and makes thousands of copies of itself in hours. The virus kills the cell and then spreads to infect other cells.
Genome Structure of a Poliovirus
The genome of the poliovirus consists of one s/s (+)sense RNA molecule of between 7.2kb (HRB14) to 8.5kb (FMDV). The genomic RNA is infectious. It is less infectious than intact particles but infectivity is increased if and when the RNA is introduced into cells by transfection. The RNA's untranslated region at the 5' end is 600-1200 bases long and is important in translation, virulence and possibly encapsidation while its' untranslated region at the 3' end is shorter, 50-100 bases, and is instrumental in (-) strand synthesis. The 5' UTR contains a 'clover-leaf' secondary structure known as the Internal Ribosome Entry Site (IRES) while the rest of the genome encodes a single 'polyprotein' of between 2100 to 2400 amino acids. Both ends of the genomes are modified: basic protein VPg is covalently attached at the 5' end and polyadentylation modifies the 3' end. While polyadenylic acid sequences are not genetically coded, there is a 'polyadenylation signal upstream of the 3' end as in eukaryotic mRNAs.
The capsid consists of a densely-packed icosahedral arrangement of 60 promoters, each consisting of 4 polypeptides, VP1. 2, and 4, all derived from cleavage of the original promoter VP0, with (pseudo) T=3 packing. The size of the particle depends on the type and degree of dessication and ranges from 27-30nm in diameter while the stretched-out length of the genome is about 2,500 nm: the genome is tightly packed into the caspid, together with sodium or potassium ions to counteract the negative charges on the phosphate groups.
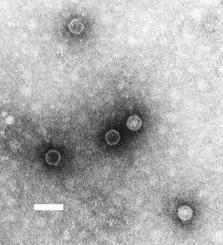
Virion Structure of a Poliovirus
The poliovirus virions are non-enveloped with isometric nucleocapsids that are sometimes penetrated by stain. The virion is of icosahedral symmetry and the nucleocapsids appear to be round. There are 12 capsomers for each nucleocapsid but there are often incomplete virus particles present that are empty capsids.
The single-step growth curve type experiments performed at high multiplicity of infection has lead to a great understanding of the Picornavirus replication process. Replication occurs entirely in the cytoplasm and can even occur in enucleated cells and is not inhibited by actinomycin D.
The primary site of infection by the poliovirus is the lymphoid tissue associated with the oropharynx and gut (GALT). The production of virus at this site leads to a transient viraemia, following which the virus may infect the CNS. This is interesting because of the apparent 'dual tropism' of the virus for two distinct cell types and reflects the distribution of the poliovirus receptor CD 155 on lymphoid cells/epithelial cells in the gut on neurons in the CNS. The virus in the CNS replicates in the 'grey matter', particularly motor neurones in the anterior horns of the spinal cord and the brain stem. Distinctive plaques produced in the grey matter are due to lytic replication of the virus and probably inflammation caused by an over-enthusiastic immune response.
Viral Ecology & Pathology
There is no animal reservoir for polioviruses. The poliovirus enters the human body through the mouth and multiplies in the intestine. The virions of poliovirus are purely protein and can resist the strong detergents present in the liver bile and constantly introduced into the small intestine to help in digestion.
The replication of poliovirus takes place in the intestinal lining but sometimes the virus escapes the gut and infects the nerve cells in the spinal column, causing paralysis. The (+) ssRNA genome of poliovirus, which is a single long RNA molecule, cannot recombine with (+) ssRNA genomes from other related picornaviruses by the process of crossing-over. Crossing-over requires a set of highly specialized enzymes that are not available to the viruses. This means only very few poliovirus strains are closely related. Thus, developing an immunity against one of these strains should work for all all these strains.
Polio has a peak incidence in the summer-early autumn in temperate climates. It mainly affects children under the age of 5. There is no cure yet for polio but it can be prevented. Polio vaccine can be administered multiple times to protect a child for life. As a result of the vaccine and the effort to fight polio, polio cases have decreased by over 99% since 1988. Whereas there were about 350 000 cases of polio then, there were only 1919 reported cases in 2002. The poliovirus has been successfully fought almost all around the world and as of 2003, there were only seven polio-endemic countries: northern India, northern Nigeria, Egypt, Pakistan, Afghanistan, Somalia and Niger.
References. Updated June 6, 2006
The Big Picture Book of Viruses: Picornaviridae
bilbo.bio.purdue.edu: Poliovirus
National Museum of American History: Polio
