Natronomonas
A Microbial Biorealm page on the Natronomonas
Classification
Higher order taxa:
Archaea; Euryarchaeota; Halobacteria; Halobacteriales; Halobacteriaceae
Species:
Natronomonas pharaonis
Description and Significance
Natronomonas pharaonis is an aerobic, extremely haloalkaliphilic archaeon that grows optimally in 3.5M NaCl and at pH 8.5, but is sensitive to high magnesium concentrations.
Genome Structure
The genome of Natronomonas pharaonis consists of three circular replicons, the chromosome which is 2,595,221 bp in length, a typical haloarchaeal 131-kb plasmid, and a unique multicopy 23-kb plasmid. Its choromosome has a high G + C content (63.4%). Also, a high proportion of acidic amino acids (average 19.3%) is found in the proteins of N. pharaonis which results in low isoelectric points (average pI 4.6). This is considered to be one of the adaptive features of haloarchaea, which are known to apply the salt-in strategy (high internal salt concentrations) in order to survive in their hypersaline environment (Falb et al). Further, it is noteworthy that because the archaeon lacks the genetic encoding for key enzymes for glycolytic pathways, it is not capable of sugar utilization.
Cell Structure and Metabolism
Natronomonas, like the other members of Halobacteriaceae, has a distinct physiological characteristics because it not only requires high NaCl concentrations but also high pH and low Mg2+ concentrations for growth. It usually utilizes amino acids as the carbon source, but the series of studies discovered that the archaeon has a high degree of nutritional self-sufficiency. Also, in contrast to other alkaliphiles, which use sodium Na+ instead of protons H+ as coupling ion between repiratory chain and ATP synthase, Natronomonas uses protons as coupling ion.
The archaeon grows under highly alkaline conditions of pH around 11, which causes reduced levels of ammonia in addition to low availability of metal ions. The genome analysis shows that, in its nitrogen metabolism process, the archaeon has three mechanisms that supply ammonia, which is then assimilated into glutamate: direct uptake of ammonia, uptake of nitrate and subsequent reduction to ammonia, and uptake of urea which is split by urease to release ammonia (see the figure below).
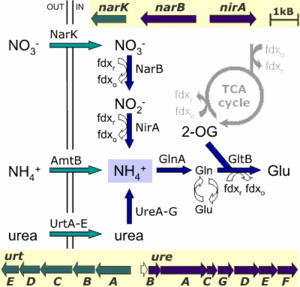
The green arrows in the figure represent the transporters for exogenous nitrogen source ammonia (AmtB), nitrate (NarK), and urea (UrtA-E), and the blue arrows represent the enzymes for reduction of nitrate (NarB + Nir A) and hydrolysis of urea (UreA-G). Other abbreviations: GlnA + GltB = glutamate; 2-OG = oxoglutarate; fdx = ferredoxin.
It is probable that Natronomonas uses ferredoxin and not NADH as the electron donor for all three reductive conversions. This is evident from the occurrence of conserved ferredoxin-biding residues within the N. pharaonis NirA protein and ferredoxin dependence of nitrate and nitrite reductases in the halophile Haloferax mediterranei.
Ecology
Strains of N. pharaonis were fist isolated from highly saline soda lakes in Egypt and Kenya, which show pH values around 11.
References
Kamekura, M. "Diversity of extremely halophilic bacteria." Extremophiles (1998) 2. p. 289-295.
