Rhodospirillum
A Microbial Biorealm page on the Rhodospirillum
|
NCBI: |
Classification
Higher order taxa:
Bacteria; Proteobacteria; Alphaproteobacteria; Rhodospirillales; Rhodospirillaceae
Species:
Rhodospirillum centenum, R. photometricum, R. rubrum, R. sp.
Description and Significance
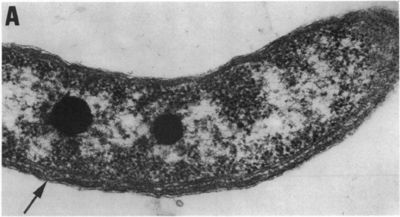
Rhodospirillum rubrum is a purple nonsulfur bacterium that can grow aerobically or anaerobically. It has the ability to live through cellular respiration, fermentation, photosynthesis, or photoautotrophic growth.
Genome Structure
None of the genomes for bacteria in the Rhodospirillum genera have been sequenced. However, the regulation of nitrogen fixation occurs at the transcriptional level with the nif expression and the posttranslational level with dinitrogenase reductase by "reversible ADP-ribosylation catalyzed by the DRAT-DRAG (dinitrogenase reductase ADP-ribosyltransferase-dinitrogenase reductase-activating glycohydrolase) system" (Zhang et al. 1999). In addition, a genetic system of the bacterial photosynthesis for Rhodospirillum centenum has been developed through studying mutants (Yildiz et al. 1991). Visit the draft of the Rhodospirillum rubrum analysis files made for the Joint Genome Institute Microbial Sequencing program for great information on the genome of Rhodospirillum rubrum.
Cell Structure and Metabolism
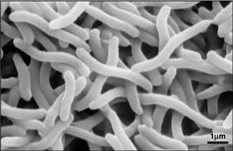
Rhodospirillum bacteria are Gram-negative, motile, spiral-shaped bacteria. They can grown under many different types of conditions including aerobic or anaerobic environments. Anaerobically, the bacterium uses fermentation or photosynthesis in order to produce energy as well as photoautotrophic growth (DOE). The nitrogen fixation system of R. rubrum uses a MoFe and an Fe-only nitrogenase. This system, which works with both translational and post-translational regulation of the nitrogenase activity and responds to both nitrogen and energy status signals, has been called the "best-understood example of reversible ADP-ridosylation as a regulatory system in any organism" (DOE). Rhodospirillum rubrum was found to be most efficient and produce the maximum levels of internal photosynthetic membranes when it was grown with both succinate and frutose as carbon sources under microaerophilic conditions (Grammel et al. 2003). However, it can grow with CO as its sole energy source (DOE). The structure o fthe CODH that is the center of R. rubrum's CO oxidation system has become a model for more complex CODHs for other organisms. In addition, the CO-oxidation regulon has a unique CO-sensing protein, CooA, that is "becoming a paradigm for gas-sensors and transcriptional regulators" (DOE).
In the absence of fructose, the bacterium only produced 20% of its maximum level of photosynthetic membranes. In aerobic conditions, R. rubrum consumed the succinate and fructose simultaneously; however, during oxygen-limiting conditions, the bacterium preferentially consumed fructose. The cell processed the frutose through the Embden-Meyer-Parnas pathway. It was also found that under oxygen-limiting conditions, NADPH was produced mostly by the pyridine-nucleotide transhydrogenase. (Grammel et al. 2003)
Ecology
Rhodospirillum bacteria can generally be found in marine environments or in some types of mud and soil where light is available for photosynthesis. Rhodospirillum centenum can form swarm colonies that "rapidly migrate toward or away from light, depending on the wavelength of excitation" by using surface-induced lateral flagella, chemotaxis, and a photosynthetic apparatus (Jiang et al. 1997).
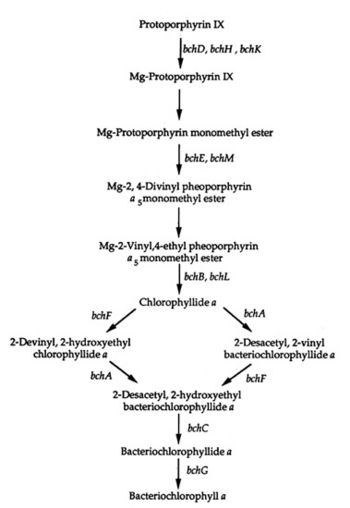
Bchl a Pathway
A genetic system of the bacterial photosynthesis in Rhodospirillum centenum was developed by studying mutants. The mutants blocked bacteriochlorophyll a biosynthesis at certain steps of the biosynthetic pathway leading from protoporphyrin IX to bacteriochlorophyll a. Some of the mutants blocked carotenoid biosynthesis early in the pathway and "exhibited pleiotropic effects on stability or assembly of the photosynthetic apparatus" (Yildiz et al. 1991). Other mutants lacked the ability to make a functional reaction center complex; others still had defective cytochromes which resulted in defective electron transport. The last type of mutant in this study had an "enhanced repression" of bacteriochlorophyll due to the presence of oxygen. The genetic system and biosynthetic pathway can be viewed in the diagram to the right.
CODH
Rhodospirillum rubrum's Ni-Fe-S carbon monoxide dehydrogenase (CODH) "catalyzes the biological oxidation of CO at an unusual Ni-Fe-S cluster called the C-cluster" that contains a mononuclear site and a four-metal cubane (Drennan et al. 2001). The CODH does this oxidation in a two-electron process:
It is thought that the CO binds to a unique CO-binding site on the C-cluster adjacent to the hydroxide; this allows a metal-bound hydroxide to "attack" the CO carbon. The metal-COOH intermediate is then deprotonated (loses the H) and the CO2 is "lost to yield a two-electron-reduced C-cluster" (Drennan et al. 2001).
References
General:
- DOE Joint Genome Institute:Rhodospirillum rubrum
- Jiang, Ze-Yu, Brenda G. Rushing, Yong Bai, Howard Gest, and Carl E. Bauer. 1998. "Isolation of Rhodospirillum centenum mutants defective in phototactic colony motility by transposon mutagenesis." Journal of Bacteriology, vol. 180, no. 5. American Society for Microbiology. (1248-1255)
- Yildiz, Fitnat H., Howard Gest, and Carl E. Bauer. 1991. "Genetic analysis of photosynthesis in Rhodospirillum centenum." Journal of Bacteriology, vol. 173, no. 13. American Society for Microbiology. (4163-4170)
Cell Structure and Metabolism:
- Grammel, Hartmut, Ernst-Deiter Gilles, and Robin Ghosh. 2003. "Microaerophilic cooperation of reductive and oxidative pathways allows maximal photosynthetic membrane biosynthesis in Rhodospirillum rubrum." Applied and Environmental Microbiology, vol. 69, no. 11. American Society for Microbiology. (6577-6586)
CODH
- Drennan, Catherine L., Jongyun Heo, Michael D. Sintchak, Eric Schreiter, and Paul W. Ludden. 2001. "Life on carbon monoxide: X-ray structure of Rhodospirillum rubrum Ni-Fe-S carbon monoxide dehydrogenase." Proceedings of the National Academy of Sciences of the USA, vol. 98, no. 21. (11973-11978)