Serratia marcescens: Difference between revisions
(Marked as uncurated) |
|||
| (66 intermediate revisions by 4 users not shown) | |||
| Line 1: | Line 1: | ||
{{Uncurated}} | |||
=Classification= | =Classification= | ||
===Higher order Taxa=== | ===Higher order Taxa=== | ||
Bacteria(Domain); Proteobacteria(Phylum); Gamma Proteobacteria(Class); Enterobacteriales(Order); Enterobacteriaceae(Family); Serratia(Genus). | Bacteria (Domain); Proteobacteria (Phylum); Gamma Proteobacteria (Class); Enterobacteriales (Order); Enterobacteriaceae (Family); Serratia (Genus). | ||
===Species=== | ===Species=== | ||
''Serratia marcescens'' | ''Serratia marcescens'' | ||
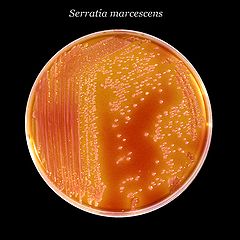
[[Image: Serratia_marcescens_01.jpg|frame|right| ''Serratia marcescens'' on an XLD agar plate. Obtained from the CDC [http://phil.cdc.gov/phil/home.asp Public Health Image Library]]] | |||
=Description and Significance= | =Description and Significance= | ||
Serratia marcescens | ''Serratia marcescens'' is a motile,short rod-shaped, Gram-negative, facultative anaerobe bacterium, classified as an opportunistic pathogen. It was discovered in 1819 by Bartolomeo Bizio in Padua, Italy. Bizio named the genus Serratia in honor of and Italian physicist named Serratia, and chose marcescens for the species name after the Latin word for decay [3]. ''Serratia marcescens'' was first thought to be harmless (non-pathogenic). Due to its ability to produce red pigmentation, it was first used in 1906 as a marker in order to trace bacterial activity or transmission [4]. It was not until later in the 1950’s that the US government experimented with the ''Serratia marcescens'' and the harmful affects that the bacteria causes were revealed. A study using ''Serratia marcescens'' was carried out to determine the possibility of biological weapons being transmitted by wind current. In the famous “Operation Sea Spray” the US Army filled balloons with ''Serratia marcescens'' and exploded them over San Fransisco. ''Serratia marcescens'' was chosen because it was easily tracked due to its pigment production. However,there was an increase in the number of pneumonia and urinary tract infections [1] reported in the region shortly after the experiment was conducted . Although ''Serratia marcescens'' was classified as a human pathogen in the 1960s, scientist still used it as a bacterial tracer well into the 1970s [10]. | ||
Optimally, ''Serratia marcescens'' grows at 37°C, but it can grow in temperatures that range from 5–40°C. They grow in pH levels that range from 5 to 9 [14]. ''Serratia marcescens'' is well known for the red pigmentation it produces called prodigiosin. Prodigiosin is made up of three pyrrole rings [15] and is not produced at 37°C, but at temperatures below 30°C [20]. The red pigment production is not present in all strains but in those that it is present, it can resemble blood [10]. This and the fact that ''Serratia marcescens'' typically grows on bread and communion wafers stored in moist places, has led scientists to suggest Serratia contamination as a possible explanation for transubstantiation miracles (the conversion of bread to the body and blood of Christ). For example, the story of the Miracle of Bolsena states that, in 1263, a priest with doubts of Christ’s presence in the consecrated Host presided over a mass in the Basilica of Bolsena. After speaking the words of consecration, blood began to drip from the consecrated Host onto his hands and the altar [1]. This event was depicted by Raphael on the walls of the Vatican [19]. | |||
=Genome Structure= | =Genome Structure= | ||
The genome of the ''Serratia marcescens'' strain Db11 was sequenced by the Sanger Institute with the collaboaration of Dr.Jonathan Ewbank of the Centre d'Immunologie de Marseille Luminy. The completed genome consists of a single circular chromosome of 5,113,802 bp containing a G+C content of 59.51%. | The genome of the ''Serratia marcescens'' strain Db11 was sequenced by the Sanger Institute with the collaboaration of Dr.Jonathan Ewbank of the Centre d'Immunologie de Marseille Luminy. The completed genome consists of a single circular chromosome of 5,113,802 bp containing a G+C content of 59.51% [21]. | ||
=Cell Structure and Metabolism= | =Cell Structure and Metabolism= | ||
Serratia marcescens is short and rod shaped. It is a facultative anaerobe, meaning that it can grow in either the presence of oxygen (aerobic) or in the absence of oxygen (anaerobic). Primarily | ''Serratia marcescens'' is short and rod shaped. It is a facultative anaerobe, meaning that it can grow in either the presence of oxygen (aerobic) or in the absence of oxygen (anaerobic). Primarily it uses fermentation as the means of gathering energy and has enzymes (superoxide dismutase, catalase or peroxides) that protect it from reactive oxygen species, allowing it to live in oxygenated environments. ''Serratia marcescens'' is a gram negative bacterium. Gram negative bacteria have a thin cell wall made of a single layer of peptidoglycan that is enclosed by an outer membrane. The outer membrane has lipopolysaccharides (LPS), which are a special kind of phospholipid composed of fatty acids that are attached to a glucosamine phosphate dimer. One glucosamine is then attached to a core polysaccharide that extends to the O polysaccharides[18]. The outer membrane also serves as a means to regulate the uptake of nutrients and the exclusion of toxins. The protein pores and transporters found in the envelope layers vary in selectivity [18]. | ||
Serratia marcescens is | ''Serratia marcescens'' is motile and travels by several different means. A single ''Serratia marcescens'' bacterium can swim with the use of its flagellum [17]. As a group they can swarm together on agar of lower concentrations (0 .5-0.8%) [8]. The swarmer cells can range in length from 5-30 µm and they are highly flagellated and nonseptated. ''Serratia marcescens'' have about 100 – 1000 flagella per swimmer cell [16]. ''Serratia marcescens'' can also form a biofilm (complex structure made of secreted mucilaginous matrixto form a protective coating in which they are encased [2]). | ||
Hydrolysis of casein is not a common trait and is | Hydrolysis of casein is not a common trait and is therefore useful in the differentiation of ''Serratia marcescens'' from the 438 strains of Enterobacteriaceae and Pseudomonadaceae families [12]. ''Serratia marcescens'' has the reproducible capability to break down casein producing a clearing on milk agar plates. Casein is a protein precipitated from milk that forms the basis of cheese and certain plastics [5]. ''Serratia marcescens'' uses extracellular enzymes called proteases to break down the peptide bonds (CO-NH) in casein [4]. Similarly, an extracellular enzyme called gelatinase breaks down gelatin, an incomplete protein that lacks tryptophan. Gelatin hydrolysis transforms the protein to individual amino acids and causes it to liquefy in cold conditions (under 25 °C) when it would otherwise be solid [4]. | ||
There are other biochemical tests that help to identify Serratia marcescens in the lab. | There are other biochemical tests that help to identify ''Serratia marcescens'' in the lab. It is negative for the methyl red test due to their production of 2, 3 - butanediol and ethanol, but positive for the Voges-Proskauer test, which shows an organism’s ability to convert pyruvate to acetoin [4]. ''Serratia marcescens'' is negative for acid production on lactose, but positive glucose and sucrose (with gas production) fermentation. Nitrate tests are positive since nitrate is generally used as the final electron acceptor rather than oxygen [4]. Citrate (positive test) is used by ''Serratia marcescens'' to produce pyruvic acid. It is positive for decarboxylase, which is the removal of a carboxyl group from an amino acid, producing an amine and carbon dioxide. | ||
The red pigmentation (prodigiosin) that Serratia marcescens is known for is only present in some | The red pigmentation (prodigiosin) that ''Serratia marcescens'' is known for can be a distinguishing factor but is only present in some strains. It is not exactly known why this is, but it is hypothesized that it is a tightly regulated gene product. Prodigiosin can trigger a body’s immune system (antibodies and T cells), so it is possible that ''Serratia marcescens'' living in a human body will limit prodigiosin synthesis and hence escape detection by the host’s immune system. Many strains appear to have lost the ability to produce it at all. [3] | ||
=Ecology= | =Ecology= | ||
''Serratia marcescens'' is ubiquitous. It is commonly found in soil, water, plants and animals. It is widely present in non potable water in underdeveloped countries due to poor chlorination. This microorganism is a common agent responsible for contamination of Petri plates in laboratories, and is also found to grow on bread [22]. Although ''S. marcescens'' is a pathogenic microorganism, it is only so with immunocompromised individuals such as those found in hospitals where many of the documented infections take place. The mode of transmission of this microorganism is by either direct contact,or by catheters, droplets, saline irrigation solutions, and other solutions that are believed to be sterile [22]. | |||
=Pathology= | =Pathology= | ||
Serratia marcescens cause nosocomial infections. It is resistant to many antibiotics traditionally used to treat bacterial infections, such as penicillin and ampicillin [ | ''Serratia marcescens'' is an opportunistic pathogen that cause nosocomial infections. It is resistant to many antibiotics traditionally used to treat bacterial infections, such as penicillin and ampicillin [9]. This is due to all of ''Serratia marcescens''’ characteristics; unique membrane (LPS) as a Gram-negative bacteria, the ability to survive in aerobic and anaerobic conditions, and its motility [10]. Most strains are resistant to several antibiotics because of the presence of R-factors (genes coding for antibiotic resistance) on plasmids [1]. | ||
There are many diseases that are associated with Serratia marcescens: sepsis, bacteremia, meningitis and cerebral abscesses, urinary tract infections, osteomyelitis, ocular infections, and endocarditis [ | There are many diseases that are associated with ''Serratia marcescens'': sepsis, bacteremia, meningitis and cerebral abscesses, urinary tract infections, osteomyelitis, ocular infections, and endocarditis [10]. Due to the wide range of diseases ''Serratia marcescens'' causes, there is not one determining symptom or source of origin. The biofilms produced are generally pathogenic in the body [2]. | ||
Also, as mentioned in the cell structure, LPS is attached to the outer membrane of the Gram negative bateria. LPS acts as an endotoxin (a cell component that is harmless as long as the pathogen remains intact). The release of LPS would over-stimulate the host defenses and cause them to undergo lethal endotoxic shock [16]. The presence of LPS therefore makes it difficult to kill | Also, as mentioned in the cell structure, the LPS layer is attached to the outer membrane of the Gram negative bateria. The LPS acts as an endotoxin (a cell component that is harmless as long as the pathogen remains intact). The release of LPS would over-stimulate the host defenses and cause them to undergo lethal endotoxic shock [16]. The presence of LPS therefore makes it difficult to kill ''Serratia marcescens'' without causing the death of the host’s cells. | ||
Some antibiotics | Some of the antibiotics have proven to be effective against ''Serratia marcescens'' are the antipseudomonal beta-lactam antibiotics, which kill bacteria by inhibiting cell wall synthesis. Though they have been developed and used to kill pseudomonas, they have also proven effective against ''Serratia marcescens'' [10] and other closely related Gram negative bacteria. Part of the problematic nature of this organism is its ability to colonize any surface. For example, ''Serratia marcescens'' has been identified as the most common organism found in both corneal scrapings and contact lenses [9]. It has been found, however, that polyquaternium-1 (a biocide used commercially in a contact lens disinfecting solution) is active against the cytoplasmic membrane of ''Serratia marcescens'' [6]. | ||
= R-factors= | |||
''S.marcescens'' contain these R-factors which are a specific types of plasmid carries one or more genes that that confers resistance to different types of antimicrobial agents.The contribution of R-factors to the resistance of ''Serratia'' to various drugs has been studied as far back as 1969[23]. This study found that out of 22 multiple resistant strians, 21 were able to transfer some form of their resistance to others. This study also demonstrated that drug resistance was far more prevalent in ''Serratia'' than in any other commonly isolated member of the Enterobacteriaceae. It was also found that not only did the R-factors mediate drug resistance to the strains that were once susceptible to certain drugs, but it further conferred additional resistance to drugs which the strains were already previously resistant to [23]. Since then, other experiments have concluded that the transfer system of R-factors in ''Serratia marcescens'' may be temperature sensitive and more likely to occur between those organisms that are found to be more closely related phylogenetically. | |||
=Efflux Pumps= | |||
Not only does ''S. marcescens'' have R-factors which encode genes for particular drug resistance, but it also contains sophisticated efflux pumps which further remove toxins that may be fatal to the microorganism. Specifically, SdeXY was the first multidrug efflux pump belonging to the RND (Resistance-Nodulation-Cell Division)family found in ''S.marcescens''. The SdeY gene is found to be a member of the RND family, while the SdeX is a member of the membrane fusion proteins. When working properly (unmutated), these proteins reduce the organism's susceptibility to erythromycin, tetracycline, norfloxacin, benzalkonium chloride, ethidium bromide, acriflavine, and rhodamine 6G (reference 4). Other efflux pumps have also been categorized such as the SdeAB RND pump and the SdeCDE RND pump. The former functions with a broad substrate specificity and the latter consists of a membrane fusion proteins (MFP)and two different RND transporters (SdeD and SdeE)[24]. | |||
Another type of multidrug efflux pump found in this microorganism is an ABC-type efflux pump called SmdAB. Both SmdA and SmdB genes must be present and are necessary for resistance[25]. | |||
=Current Research= | =Current Research= | ||
One recent study suggests that ''Serratia marcescens''ost3 produces a novel prodigiosin called MAMPDM ((2,2'-[3-methoxy-1'amyl-5'-methyl-4-(1"-pyrryl)]dipyrrylmethene))that may have an important impact on cancer treatment. This red pigment demonstrated a selective cytotoxic activity in cancer cell lines, but in contrast it revealed a reduced toxicity to non malignant cells. They came to the conclusion that ''Serratia marcescens''ost3 may in one time be used as a source to develop an anti-cancer compound [26]. | |||
Another study suggests that ''Serratia marcescens'' strain NCTC 10211 may serve as a probiotic in inhibiting the growth of ''H.pylori'', the causative agent of stomach ulcers [27]. Upon examining several different species and bacteria strains, ''Serratia marcescens'' strain NCTC 10211 revealed striking inhibition zones when inoculated with different Hp strains.The components and/or mechanism involved in inhibition of ''Hp'' proliferation are still not well understood. Further research is needed to isolate and characterize the inhibitory substances so that they may be used for ''Hp'' therapy. It is largely unclear whether ''Serratia marcescens'' has bactericidial activity or if promotes changes in the physiology and morphology of ''Hp''. | |||
''Serratia marcescens'' has a unique ability to produce extracellular enzymes (28). Several such enzymes have been shown to have the ability to degrade chitin, a substance which mainly comprises fungal cell walls (28). These chitinolytic enzymes could have possible industrial and agricultural uses such as the introduction of these genes for chitin degrading enzymes into crops as well as fermentative bacteria (28). Further research into protein modification of nucleotide sequences would allow for improved expression of chitin producing genes, thus providing a protective mechanism for susceptible plants and fermentative bacteria against fungal infections (28). | |||
=References= | =References= | ||
| Line 86: | Line 104: | ||
20. Yuko Tanaka, Junko Yuasa, Masahiro Baba, Taichiro Tanikawa, Yoji Nakagawa and Tohey Matsuyama. “Temperature-Dependent Bacteriostatic Activity of Serratia marcescens”. Microb. Environ.. Vol. 19: 236-240 (2004) . | 20. Yuko Tanaka, Junko Yuasa, Masahiro Baba, Taichiro Tanikawa, Yoji Nakagawa and Tohey Matsuyama. “Temperature-Dependent Bacteriostatic Activity of Serratia marcescens”. Microb. Environ.. Vol. 19: 236-240 (2004) . | ||
21. http://www.sanger.ac.uk/Projects/S_marcescens/ | |||
22. http://commtechlab.msu.edu/sites/dlc-me/zoo/microbes/serratia.html | |||
23. Cooksey Robert, Grace Thorne, Edmund Farrar. "R Factor Mediated Antibiotic Resistance in ''Serratia marcescens''. Antimicrobia Agents And ChemoTheraphy. Vol 10. No. 1 July 1976 p 123-127. | |||
24. Belgic, Sanela and Elizabeth A. Worobec. "Characterization of the ''Serratia marcescens'' SdeCDE multidrug efflux pump studied via gene knockout mutagenesis". Department of Microbiology, University of Manitoba, Winnipeg, MB R3T 2N2, Canada. | |||
25. Taira Matsuo, Jing Chen, Yusuke Minato, Wakano Ogawa, Tohru Mizushima, Teruo Kuroda, and Tomofusa Tsuchiya. "SmdAB, a Heterodimeric ABC-Type Multidrug Efflux Pump, in ''Serratia marcescens''. Journal of Bacteriology, Jan.2008 p.648-654. | |||
26. <A href="http://search.ebscohost.com/login.aspx?direct=true&db=syh&AN25320763&site=ehost-live">Identification of a red-pigmented bacterium producing a potent anti-tumor N-alkylated prodigiosin as Serratia marcescens.</A> | |||
27. R Krausse, K. Piening, U. Ullman. "Inhibitory effects of various micro-organisms on the growth of ''Heliobacter pylori''. Letter in Applied Mirobiology 2005, 40, 81-86. | |||
28. Hejazi, A.,and Falkiner, F. R. “Serratia marcescens” Journal of Medical Microbiology, 1997. Volume 46. p. 903-912. | |||
Edited by Irina Mashenko and Abrehet Tesfamicael students of M Glogowski at Loyola University 1- http://www.ncbi.nlm.nih.gov/pmc/articles/PMC87738/ | |||
<!-- Do not edit or remove this line-->[[Category:Pages edited by students of Mary Glogowski at Loyola University]] | |||
Latest revision as of 21:37, 1 April 2011
Classification
Higher order Taxa
Bacteria (Domain); Proteobacteria (Phylum); Gamma Proteobacteria (Class); Enterobacteriales (Order); Enterobacteriaceae (Family); Serratia (Genus).
Species
Serratia marcescens
Description and Significance
Serratia marcescens is a motile,short rod-shaped, Gram-negative, facultative anaerobe bacterium, classified as an opportunistic pathogen. It was discovered in 1819 by Bartolomeo Bizio in Padua, Italy. Bizio named the genus Serratia in honor of and Italian physicist named Serratia, and chose marcescens for the species name after the Latin word for decay [3]. Serratia marcescens was first thought to be harmless (non-pathogenic). Due to its ability to produce red pigmentation, it was first used in 1906 as a marker in order to trace bacterial activity or transmission [4]. It was not until later in the 1950’s that the US government experimented with the Serratia marcescens and the harmful affects that the bacteria causes were revealed. A study using Serratia marcescens was carried out to determine the possibility of biological weapons being transmitted by wind current. In the famous “Operation Sea Spray” the US Army filled balloons with Serratia marcescens and exploded them over San Fransisco. Serratia marcescens was chosen because it was easily tracked due to its pigment production. However,there was an increase in the number of pneumonia and urinary tract infections [1] reported in the region shortly after the experiment was conducted . Although Serratia marcescens was classified as a human pathogen in the 1960s, scientist still used it as a bacterial tracer well into the 1970s [10].
Optimally, Serratia marcescens grows at 37°C, but it can grow in temperatures that range from 5–40°C. They grow in pH levels that range from 5 to 9 [14]. Serratia marcescens is well known for the red pigmentation it produces called prodigiosin. Prodigiosin is made up of three pyrrole rings [15] and is not produced at 37°C, but at temperatures below 30°C [20]. The red pigment production is not present in all strains but in those that it is present, it can resemble blood [10]. This and the fact that Serratia marcescens typically grows on bread and communion wafers stored in moist places, has led scientists to suggest Serratia contamination as a possible explanation for transubstantiation miracles (the conversion of bread to the body and blood of Christ). For example, the story of the Miracle of Bolsena states that, in 1263, a priest with doubts of Christ’s presence in the consecrated Host presided over a mass in the Basilica of Bolsena. After speaking the words of consecration, blood began to drip from the consecrated Host onto his hands and the altar [1]. This event was depicted by Raphael on the walls of the Vatican [19].
Genome Structure
The genome of the Serratia marcescens strain Db11 was sequenced by the Sanger Institute with the collaboaration of Dr.Jonathan Ewbank of the Centre d'Immunologie de Marseille Luminy. The completed genome consists of a single circular chromosome of 5,113,802 bp containing a G+C content of 59.51% [21].
Cell Structure and Metabolism
Serratia marcescens is short and rod shaped. It is a facultative anaerobe, meaning that it can grow in either the presence of oxygen (aerobic) or in the absence of oxygen (anaerobic). Primarily it uses fermentation as the means of gathering energy and has enzymes (superoxide dismutase, catalase or peroxides) that protect it from reactive oxygen species, allowing it to live in oxygenated environments. Serratia marcescens is a gram negative bacterium. Gram negative bacteria have a thin cell wall made of a single layer of peptidoglycan that is enclosed by an outer membrane. The outer membrane has lipopolysaccharides (LPS), which are a special kind of phospholipid composed of fatty acids that are attached to a glucosamine phosphate dimer. One glucosamine is then attached to a core polysaccharide that extends to the O polysaccharides[18]. The outer membrane also serves as a means to regulate the uptake of nutrients and the exclusion of toxins. The protein pores and transporters found in the envelope layers vary in selectivity [18].
Serratia marcescens is motile and travels by several different means. A single Serratia marcescens bacterium can swim with the use of its flagellum [17]. As a group they can swarm together on agar of lower concentrations (0 .5-0.8%) [8]. The swarmer cells can range in length from 5-30 µm and they are highly flagellated and nonseptated. Serratia marcescens have about 100 – 1000 flagella per swimmer cell [16]. Serratia marcescens can also form a biofilm (complex structure made of secreted mucilaginous matrixto form a protective coating in which they are encased [2]).
Hydrolysis of casein is not a common trait and is therefore useful in the differentiation of Serratia marcescens from the 438 strains of Enterobacteriaceae and Pseudomonadaceae families [12]. Serratia marcescens has the reproducible capability to break down casein producing a clearing on milk agar plates. Casein is a protein precipitated from milk that forms the basis of cheese and certain plastics [5]. Serratia marcescens uses extracellular enzymes called proteases to break down the peptide bonds (CO-NH) in casein [4]. Similarly, an extracellular enzyme called gelatinase breaks down gelatin, an incomplete protein that lacks tryptophan. Gelatin hydrolysis transforms the protein to individual amino acids and causes it to liquefy in cold conditions (under 25 °C) when it would otherwise be solid [4].
There are other biochemical tests that help to identify Serratia marcescens in the lab. It is negative for the methyl red test due to their production of 2, 3 - butanediol and ethanol, but positive for the Voges-Proskauer test, which shows an organism’s ability to convert pyruvate to acetoin [4]. Serratia marcescens is negative for acid production on lactose, but positive glucose and sucrose (with gas production) fermentation. Nitrate tests are positive since nitrate is generally used as the final electron acceptor rather than oxygen [4]. Citrate (positive test) is used by Serratia marcescens to produce pyruvic acid. It is positive for decarboxylase, which is the removal of a carboxyl group from an amino acid, producing an amine and carbon dioxide. The red pigmentation (prodigiosin) that Serratia marcescens is known for can be a distinguishing factor but is only present in some strains. It is not exactly known why this is, but it is hypothesized that it is a tightly regulated gene product. Prodigiosin can trigger a body’s immune system (antibodies and T cells), so it is possible that Serratia marcescens living in a human body will limit prodigiosin synthesis and hence escape detection by the host’s immune system. Many strains appear to have lost the ability to produce it at all. [3]
Ecology
Serratia marcescens is ubiquitous. It is commonly found in soil, water, plants and animals. It is widely present in non potable water in underdeveloped countries due to poor chlorination. This microorganism is a common agent responsible for contamination of Petri plates in laboratories, and is also found to grow on bread [22]. Although S. marcescens is a pathogenic microorganism, it is only so with immunocompromised individuals such as those found in hospitals where many of the documented infections take place. The mode of transmission of this microorganism is by either direct contact,or by catheters, droplets, saline irrigation solutions, and other solutions that are believed to be sterile [22].
Pathology
Serratia marcescens is an opportunistic pathogen that cause nosocomial infections. It is resistant to many antibiotics traditionally used to treat bacterial infections, such as penicillin and ampicillin [9]. This is due to all of Serratia marcescens’ characteristics; unique membrane (LPS) as a Gram-negative bacteria, the ability to survive in aerobic and anaerobic conditions, and its motility [10]. Most strains are resistant to several antibiotics because of the presence of R-factors (genes coding for antibiotic resistance) on plasmids [1]. There are many diseases that are associated with Serratia marcescens: sepsis, bacteremia, meningitis and cerebral abscesses, urinary tract infections, osteomyelitis, ocular infections, and endocarditis [10]. Due to the wide range of diseases Serratia marcescens causes, there is not one determining symptom or source of origin. The biofilms produced are generally pathogenic in the body [2].
Also, as mentioned in the cell structure, the LPS layer is attached to the outer membrane of the Gram negative bateria. The LPS acts as an endotoxin (a cell component that is harmless as long as the pathogen remains intact). The release of LPS would over-stimulate the host defenses and cause them to undergo lethal endotoxic shock [16]. The presence of LPS therefore makes it difficult to kill Serratia marcescens without causing the death of the host’s cells.
Some of the antibiotics have proven to be effective against Serratia marcescens are the antipseudomonal beta-lactam antibiotics, which kill bacteria by inhibiting cell wall synthesis. Though they have been developed and used to kill pseudomonas, they have also proven effective against Serratia marcescens [10] and other closely related Gram negative bacteria. Part of the problematic nature of this organism is its ability to colonize any surface. For example, Serratia marcescens has been identified as the most common organism found in both corneal scrapings and contact lenses [9]. It has been found, however, that polyquaternium-1 (a biocide used commercially in a contact lens disinfecting solution) is active against the cytoplasmic membrane of Serratia marcescens [6].
R-factors
S.marcescens contain these R-factors which are a specific types of plasmid carries one or more genes that that confers resistance to different types of antimicrobial agents.The contribution of R-factors to the resistance of Serratia to various drugs has been studied as far back as 1969[23]. This study found that out of 22 multiple resistant strians, 21 were able to transfer some form of their resistance to others. This study also demonstrated that drug resistance was far more prevalent in Serratia than in any other commonly isolated member of the Enterobacteriaceae. It was also found that not only did the R-factors mediate drug resistance to the strains that were once susceptible to certain drugs, but it further conferred additional resistance to drugs which the strains were already previously resistant to [23]. Since then, other experiments have concluded that the transfer system of R-factors in Serratia marcescens may be temperature sensitive and more likely to occur between those organisms that are found to be more closely related phylogenetically.
Efflux Pumps
Not only does S. marcescens have R-factors which encode genes for particular drug resistance, but it also contains sophisticated efflux pumps which further remove toxins that may be fatal to the microorganism. Specifically, SdeXY was the first multidrug efflux pump belonging to the RND (Resistance-Nodulation-Cell Division)family found in S.marcescens. The SdeY gene is found to be a member of the RND family, while the SdeX is a member of the membrane fusion proteins. When working properly (unmutated), these proteins reduce the organism's susceptibility to erythromycin, tetracycline, norfloxacin, benzalkonium chloride, ethidium bromide, acriflavine, and rhodamine 6G (reference 4). Other efflux pumps have also been categorized such as the SdeAB RND pump and the SdeCDE RND pump. The former functions with a broad substrate specificity and the latter consists of a membrane fusion proteins (MFP)and two different RND transporters (SdeD and SdeE)[24].
Another type of multidrug efflux pump found in this microorganism is an ABC-type efflux pump called SmdAB. Both SmdA and SmdB genes must be present and are necessary for resistance[25].
Current Research
One recent study suggests that Serratia marcescensost3 produces a novel prodigiosin called MAMPDM ((2,2'-[3-methoxy-1'amyl-5'-methyl-4-(1"-pyrryl)]dipyrrylmethene))that may have an important impact on cancer treatment. This red pigment demonstrated a selective cytotoxic activity in cancer cell lines, but in contrast it revealed a reduced toxicity to non malignant cells. They came to the conclusion that Serratia marcescensost3 may in one time be used as a source to develop an anti-cancer compound [26].
Another study suggests that Serratia marcescens strain NCTC 10211 may serve as a probiotic in inhibiting the growth of H.pylori, the causative agent of stomach ulcers [27]. Upon examining several different species and bacteria strains, Serratia marcescens strain NCTC 10211 revealed striking inhibition zones when inoculated with different Hp strains.The components and/or mechanism involved in inhibition of Hp proliferation are still not well understood. Further research is needed to isolate and characterize the inhibitory substances so that they may be used for Hp therapy. It is largely unclear whether Serratia marcescens has bactericidial activity or if promotes changes in the physiology and morphology of Hp.
Serratia marcescens has a unique ability to produce extracellular enzymes (28). Several such enzymes have been shown to have the ability to degrade chitin, a substance which mainly comprises fungal cell walls (28). These chitinolytic enzymes could have possible industrial and agricultural uses such as the introduction of these genes for chitin degrading enzymes into crops as well as fermentative bacteria (28). Further research into protein modification of nucleotide sequences would allow for improved expression of chitin producing genes, thus providing a protective mechanism for susceptible plants and fermentative bacteria against fungal infections (28).
References
1. amh10. “Serratia Marcescens.” MicrobLog.com. 4 August 2006. 7 Nov. 2008. © 2008 <http://microblog.me.uk/89>
2. "biofilm." The American Heritage® Science Dictionary. Houghton Mifflin Company. 02 Dec. 2008. <Dictionary.com http://dictionary.reference.com/browse/biofilm>.
3. Bry, Lynn. “Re: How and why did Serratia marcescens produce prodigiosin?” 5 June 2005. 2 Dec. 2008. <http://www.madsci.org/posts/archives/2005-06/1117999360.Mi.r.html>
4. Cappuccino, James and Natalie Sherman. Microbiology: a Laboratory Manuel. 7th edition. ©2005 Pearson Education, Inc. pp.146,151,165
5. "casein." Dictionary.com Unabridged (v 1.1). Random House, Inc. 03 Dec. 2008. <Dictionary.com http://dictionary.reference.com/browse/casein>.
6. Codling Caroline, Brian V. Jones, Eshwar Mahenthiralingam, A. Denver Russell and Jean-Yves Maillard. “Identification of genes involved in the susceptibility of Serratia marcescens to polyquaternium-1.” Journal of Antimicrobial Chemotherapy (2004) 54, 370–375
7. “Excerpt from Serratia.” Emedicine.com. 19 Nov. 2008. © 1996-2006 by WebMD <http://www.emedicine.com/med/byname/serratia.htm>
8. Harshey Rasika M. “Bees aren't the only ones : swarming in Gram-negative bacteria.” Molecular Microbiology (1994) 13(3), 389-394
9. JAMA and Archives Journals. "Organisms Found On Contact Lenses Can Provide Clues To Cause Of Corneal Eye Infection." ScienceDaily 13 September 2007. 10 December 2008 <http://www.sciencedaily.com¬ /releases/2007/09/070910160823.htm>.
10. “Serratia Marcescens Bacteria.” serratia-marcescens.org. 9 Nov. 2008. <http://www.serratia-marcescens.org/>
11. "r factor." Dictionary.com Unabridged (v 1.1). Random House, Inc. 07 Dec. 2008. <Dictionary.com http://dictionary.reference.com/browse/r factor>.
12. Salisbury, Willaiam and Joseph Likos. Hydrolysis of casein: a differential aid for the identification of Serratia marcescens. J. clin. Path., 1972, 25, 1083-1085
13. "Serratia marcescens." WordNet® 3.0. Princeton University. 6 Nov. 2008. <Dictionary.com http://dictionary.reference.com/browse/serratia marcescens>.
14. “Serratia marcescens.” Wikipedia.org. 24 Oct. 2008. 19 Nov. 2008. <http://en.wikipedia.org/wiki/Serratia_marcescens>
15. Schlegel, Hans. General Microbiology. © Georg Thieme Verlag, Stuttgart 1992. pp. 88.
16. Shapiro, James and Martin Dworkin. Bacteria as Multicellular Organisms. © 1997 by Oxford University Press, Inc. pp. 210
17. Simurda, Maryanne. “Department of Biology Faculty and Research.” 9 Nov. 2008. <http://biology.wlu.edu/simurda.htm>
18. Slonczewski, Joan and John Foster. Microbiology: An Evolving Science. © 2009 W.W. Norton & Company, Inc. pp. 91, 488
19. “The Miracle Microbe: Serratia marcescens.” 18 Nov. 2008. © 1999 Comm Tech Lab, Michigan State Univeristy. < http://commtechlab.msu.edu/sites/dlc-me/zoo/microbes/serratia.html>
20. Yuko Tanaka, Junko Yuasa, Masahiro Baba, Taichiro Tanikawa, Yoji Nakagawa and Tohey Matsuyama. “Temperature-Dependent Bacteriostatic Activity of Serratia marcescens”. Microb. Environ.. Vol. 19: 236-240 (2004) .
21. http://www.sanger.ac.uk/Projects/S_marcescens/
22. http://commtechlab.msu.edu/sites/dlc-me/zoo/microbes/serratia.html
23. Cooksey Robert, Grace Thorne, Edmund Farrar. "R Factor Mediated Antibiotic Resistance in Serratia marcescens. Antimicrobia Agents And ChemoTheraphy. Vol 10. No. 1 July 1976 p 123-127.
24. Belgic, Sanela and Elizabeth A. Worobec. "Characterization of the Serratia marcescens SdeCDE multidrug efflux pump studied via gene knockout mutagenesis". Department of Microbiology, University of Manitoba, Winnipeg, MB R3T 2N2, Canada.
25. Taira Matsuo, Jing Chen, Yusuke Minato, Wakano Ogawa, Tohru Mizushima, Teruo Kuroda, and Tomofusa Tsuchiya. "SmdAB, a Heterodimeric ABC-Type Multidrug Efflux Pump, in Serratia marcescens. Journal of Bacteriology, Jan.2008 p.648-654.
26. <A href="http://search.ebscohost.com/login.aspx?direct=true&db=syh&AN25320763&site=ehost-live">Identification of a red-pigmented bacterium producing a potent anti-tumor N-alkylated prodigiosin as Serratia marcescens.</A>
27. R Krausse, K. Piening, U. Ullman. "Inhibitory effects of various micro-organisms on the growth of Heliobacter pylori. Letter in Applied Mirobiology 2005, 40, 81-86.
28. Hejazi, A.,and Falkiner, F. R. “Serratia marcescens” Journal of Medical Microbiology, 1997. Volume 46. p. 903-912.
Edited by Irina Mashenko and Abrehet Tesfamicael students of M Glogowski at Loyola University 1- http://www.ncbi.nlm.nih.gov/pmc/articles/PMC87738/
