User:GillenK
Targeting of proteins to different cellular compartments in E. coli. From MicrobeWiki, the student-edited microbiology resource Jump to: navigation, search Contents [hide]
Introduction
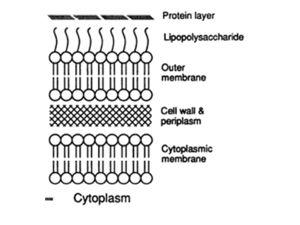
Sorting proteins to their correct cellular location is of critical importance to cells. Mis-localized proteins are non-functional and can lead to cell death. Protein sorting to the various organelles in eukaryotic cells is widely studied and resulted in a Nobel Prize to Blobel for the recognition of the signal sequence that initially targets secreted and membrane proteins to the endoplasmic reticulum. While it has been long assumed that prokaryotes lack the sophistical intracellular architecture of eukaryotic cells, more recent research has shown that prokaryotes do indeed have cytoplasmic organization with various proteins restricted to certain areas. In addition to the cytoplasm, Gram-negative bacteria must selectively localize proteins to the inner membrane (IM), periplasm, outer membrane (OM), and extracellular environment (Figure 1). This page will summarize how proteins are targeted to various compartments in Gram-negative cells, taking most examples from the well-studied Escherichia coli (E. coli). Beyond to contributing to basic biological research, studies of protein sorting in bacteria have biotechnology and medical applications (Cormelius, P., 2000). Purification of recombinant proteins in E. coli is aided by targeting the proteins to particular cell locations and studies of bacterial toxin secretion may lead to novel therapeutic agents.
Inner Membrane
Include some current research, with at least one figure showing data.
Periplasm
Include some current research, with at least one figure showing data.
Outer membrane
Include some current research, with at least one figure showing data. Conclusion
Extracellular
In addition to targeting to various cellular locations, some proteins are destined to leave the cell entirely and enter the extracellular environment. Examples include hemolysins exotoxins that l yse host cells to release nutrients and pili that mediate attachment to substrates. In gram-negative cells secreted proteins must cross both the inner and outer membranes. Secretion across the outer membranes differs from secretion across the inner membrane in a number of respects. In contrast to the IM Sec proteins through which proteins pass in an unfolded state, proteins may partially fold in the periplasm prior to secretion through the OM. (Thanassi and Hultgren 2000).
[[Image:T5SS|thumb|300 × 300 px|right| Figure 2.A schematic diagram of the autotransporter family. From http://en.wikipedia.org/wiki/Secretion courtesy Mike Jones]] Thanassi and Hultgren (2000) reviewed the different pathways proteins may take on their voyage outside the Gram-negative cell. They discuss six different pathways to protein secretion, four of which are Sec dependent and two that are Sec independent. Proteins using the Sec dependent pathways, as the name suggests, cross the inner membrane via the Sec pathway described above. They then rely on different paths to extracellular secretion. The simplest path is taken by autotransporters, proteins that form their own pore in the OM. Autotransporter proteins contain three domains, the N-terminal signal sequence for Sec targeting, a carboxy-terminal beta-barrel that forms a pore in the OM, and a passenger domain. The passenger domain passes through the pore and can be cleaved to be released extracellularly. There is also a single accessory factor pathway that encodes the beta-barrel OM pore as a separate protein. The beta-barrels of the autotransporters and single accessory factors are not homologous and appear to have evolved separately. The chaperone/usher pathway is a second route outside the cell.
References
Cornelis, P. Current opinion in biotechnology 2000;11(5):450-4
Edited by student of Joan Slonczewski for BIOL 238 Microbiology, 2010, Kenyon College. Retrieved from "http://microbewiki.kenyon.edu/index.php/BIOL_238_Paper_2010" Category: Pages edited by students of Joan Slonczewski at Kenyon College
