User talk:Aamirie
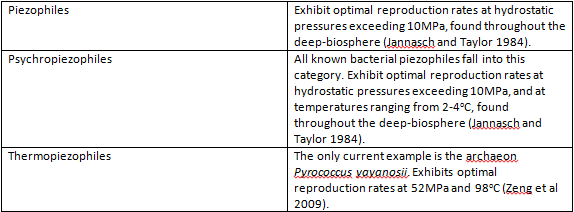
Piezophiles
Introduction
Piezophiles (barophiles) are organisms whose survival and reproduction is optimized to high pressures, such as those in deep sea environments. Piezophiles are found primarily in the depths of the ocean, which has an average pressure of 38 MPa (megapascals) and reaches 110 MPa at its deepest point in the Marianas Trench, in comparison to a pressure of 0.1 MPa at sea level (Abe and Horikoshi 2001). Remarkably, the most pressure resistant organism currently known, Pyrococcus yayanosii, can survive at pressures up to 150MPa (Zeng et al. 2009). In order to inhabit such extreme environments, these organisms have evolved various mechanisms to counteract the effects of the elevated pressures they endure. While relatively little is known about piezophiles as of yet, preliminary research indicates potential industrial and biotechnological applications for high pressure adaptation mechanisms (Abe and Horikoshi 2001).
Origin and distribution
Phylogenetic analysis of the 16S rRNA sequences of piezophilic bacteria shows a high level of similarity to non-piezophilic bacteria from Antarctica, and all but one of the currently known piezophilic species are also psychropiezophiles. Because of this, it is theorized that piezophily arose from psychrophiles, whose pre-existing adaptations to extreme cold overlap significantly with the adaptations necessary to survive in high pressure environments (Lauro et al. 2006). The vast majority of piezophiles are bacteria, and can be grouped into one of five genera – Shewanella, Colwellia, Photobacterium, Moritella, and Psychromonas. The first example of a piezophilic archaeon is the recently discovered thermopiezophile Pyrococcus yayanosii (Zeng et al. 2009).
Physical environment
Piezophilic organisms are found in what is called the deep-biosphere, defined as those environments more than 1000m below sea level and with hydrostatic pressures greater than 10MPa (Jannasch and Taylor 1984) . The deep-biosphere is for the most part oligotrophic, characterized by a lack of sunlight and nutrients – less than 1% of available organic material descends to the levels inhabited by these organisms. The piezophiles that have been isolated as of yet can be divided into thermopiezophiles and psychropiezophiles. The only known thermopiezophile is Pyrococcus yayanosii strain CH1 (Oger and Jebbar 2010), which is found in the vicinity of hydrothermal vents – fissures in the sea floor from which water geothermally heated to temperatures up to 400°C is emitted, resulting in eutrophic, microbially dense communities (Zierenberg et al. 2000). Psychropiezophiles are also found in the depths of the ocean, but in areas not heated by geothermal energy. The average temperature in this environment is approximately 2°C (Kato and Qureshi 1999).
Adaptations to high pressure
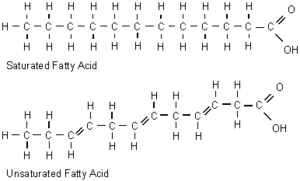
High pressure has a similar effect on cell membranes as low temperature, which reduces fluidity by increasing packing of the fatty acyl chains of phospholipids. Most piezophiles are subject to both intense hydrostatic pressure and low temperatures, and the decrease in membrane fluidity this causes can result in a gel-like membrane, which interferes with nutrient uptake and cell signalling mechanisms. In piezophiles, this is prevented by increasing the proportion of fluidizing monounsaturated and polyunsaturated fatty acids in their membranes, which cannot be packed as tightly as saturated fatty acids (Bartlett 1999).
High hydrostatic pressure is also known to change the conformation of cellular proteins. While pressures in the range relevant to piezophiles are not sufficient to denature protein, the small conformational changes they cause have been found to change or inhibit protein function (van den Berg 2003). Amino acid sequencing of proteins from piezophilic Shewanella bacteria indicates lower concentrations of proline residues (Bartlett 1999), whose cyclic side chains disrupt alpha helices, as well as glycine residues, whose small side chains with high conformational flexibility also destabilize helical protein structure (Richardson 1981). These modifications lead to an overall decrease in protein flexibility, and increased stability in high pressure environments (Bartlett 1999). At the gene expression level, high hydrostatic pressure has been shown to up-regulate the transcription of genes encoding chaperone proteins, which maintain proper folding, and therefore function of the proteins they act on (Oger and Jebbar 2010).
Possible applications of high pressure adaptation mechanisms
Research on piezophiles has yielded promising results in regulating gene expression with hydrostatic pressure. This regulation mechanism was first discovered in Photobacterium profundum, in which hydrostatic pressure was found to act on the ToxR-ToxS complex, which in turn controls expression of two outer membrane proteins, OmpH and OmpL. At the bacterium’s optimal pressure of 28 MPa, OmpH is dominant, but when exposed to decreasing hydrostatic pressure the ratio of OmpH to OmpL decreased accordingly (Simonato et al 2006). A promoter has also been found in Shewanella benthica and Shewanella violacea which increases expression of its downstream genes when grown at a pressure of 50 MPa. It has been theorized that inserting specific genes downstream of this promoter via homologous recombination, or insertion of the promoter sequence upstream of a desired gene would allow pressure regulated control of gene expression (Abe and Horikoshi 2001).
A number of proteins are in fact more stable at high hydrostatic pressures, and increasing pressure has been shown to prevent denaturation of some proteins even past their inactivation temperatures. For example, the half-life of Taq polymerase, an enzyme essential in the polymerase chain reaction process, was found to increase from 5 min at 3 MPa to 12 min at 45 MPa. This effect has also been observed for other DNA polymerases, as well as various hydrogenases (which catalyse oxidation of H2), and α-glucosidases (which are involved in the breakdown of starch and glycogen) (Abe and Horikoshi 2001). The study of these proteins and why they stabilize under these conditions is of use in the design of synthetic proteins to increase efficiency of processes that can be performed at high hydrostatic pressures, such as food processing and antibiotic production (van den Berg 2003).
References
Abe, F., and Horikoshi, K. “The biotechnological potential of piezophiles.” TRENDS in Biotechnology, 2001.
Bartlett D. H. “Microbial adaptations to the psychrosphere/piezosphere.” Journal of Molecular Microbiology and Biotechnology, 1999.
Jannasch, H.W., and Taylor, C.D. “Deep-sea microbiology.” Annual Review of Microbiology, 1984, 38:487-514.
Kato, C., and Qureshi, M. H. “Pressure response in deep-sea piezophilic bacteria.” Journal of Molecular Microbiology and Biotechnology, 1999.
Lauro, F. M., Chastain, R. A., Blankenship, L. E., Yayanos, A. A., and Bartlett, D. H. “The unique 16S rRna genes of piezophiles reflect both phylogeny and adaptation.” Applied Environmental Microbiology, 2006, DOI: 10.1128/AEM.01726-06
Oger, P.M., and Jebbar, M. “The many ways of coping with pressure.” Research in Microbiology, 2010, DOI: 10.1016/j.resmic.2010.09.017
Richardson, J. S. “The anatomy and taxonomy of protein structure.” Advanced Protein Chemistry, 1981.
Simonato, F., Campanaro, S., Lauro, F. M., Vezzi, A., D’Angelo, M., Vitulo, N., Valle, G., and Bartlett, D. H. “Piezophilic adaptation: a genomic point of view.” Journal of Biotechnology, 2006, DOI: 10.1016/j.jbiotec.2006.03.038
van den Berg, B. “Extremophiles as a source for novel enzymes.” Current Opinion in Microbiology, 2003, DOI 10.1016/S1369-5274(03)00060-2\
Zeng, X., Birrien, J.L., Fouquet, Y., Cherkashkov, G., Jebbar, M., Querellou, J., Oger, P., Cambon-Bonavita, M.A., Xiao, X., Preur, D. “Pyrococcus CH1, an obligate piezophilic hyperthermophile: extending the upper pressure-temperature limits for life.” International Society for Microbial Ecology Journal, 2009, Jul;3(7):873-876.
Zierenberg, R. A., Adams, M. W. W., and Arp, A. J. “Life in extreme environments: hydrothermal vents.” National Academy of Sciences, 2000, DOI: 10.1073/pnas.210395997