Yersinia pestis, the History of the Plague and Adaptation to Animal Host
Classification
• Kingdom: Eubacteria
• Phylum: Proteobacteria
• Class: Gammaproteobacteria
• Order: Enterobacteriales
• Family: Enterobacteriaceae
• Genus: Yersinia
Structure and Significance
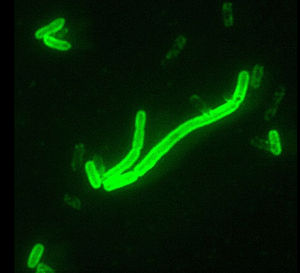
By: Blake Calcei
Yersinia pestis, is most notable for its devastating role in plagues of the past. These bacteria are Gamaproteobacteria that are Gram negative with a coccobacillus shape and are facultative anaerobes. The shape of these bacteria are demonstrated in figure 1, which displays the rod looking shape and is seen through green fluorescence. This bacterium causes infection in humans and other non-human animals. These bacteria can be found in animal or insect blood, and this blood is required for the bacteria to survive. The most common hosts of these bacteria include, insects (fleas), rodents and humans. The most notable carrier of this bacteria is the Oriental Rat flea, which contains bacteria in its blood but is unaffected by the bacteria. These fleas then infect rats, which eventually pass the bacterium to humans and other non-human animals. The three main infections caused by this bacterium include: pneumonic, septicemic and bubonic plagues. These diseases are also commonly found in highly populated areas that have poor sanitation habits therefore increasing the rat population and making the location prime for Y. pestis infections. The progression from fleas to rats and the widespread populations of both of these organisms has allowed these bacteria to be exposed to a wide variety of environmental conditions allowing them to adapt to multiple strains that are very resilient. The regulatory systems involved in the virulence of these bacteria have been found to be quite complex posing a large problem for antibiotics.[1]
Discovery
Alexandre Yersin first discovered Yersinia pestis in 1894 at the Pasteur Institute, while a plague occurred in Hong Kong, cause by this Y. pestis bacterium. Yersin was the first to link the plague to Y. pestis and therefore the original name of the bacterium, Pasteurella pestis, was changed to the modern name, Yersinia pestis. This outbreak in China, which began in 1855, was said to have killed over 12 million people in both India and China, and was not considered over until 1959. There have been three major outbreaks of plague caused by these bacteria, the plague of Justinian, the Black Death, and the third plague pandemic.[2]
History
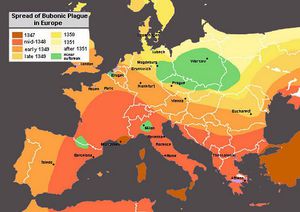
The Plague of Justinian
The Plague of Justinian occurred during the 6th century and primarily between 541-542 AD in the Byzantine Empire, where it wiped out nearly one third of the population. The spread of this plague was Constantinople, to southern Europe and to the Persian Empire in the east. This pandemic is similar to the bubonic plague of the 14th century but has distinct differences, including some victims undergoing hallucinations and delirium, which was not reported during the 14th century bubonic plague.[3] This plague is also one of the first to have been recorded in human history.
This plague is believed to have been the first rise of the bubonic plague and DNA research by Wagner et. al, has been conducted on bodies from the era and the bacteria Yersinia pestis was found.[4] In order to determine if Y. pestis was the cause of this plague the researchers sequenced and analyzed genomes of Y. pestis, which were taken from two specific individuals known to have died in this first plague pandemic. The two individuals used were carbon dated to the period of the first plague pandemic. To determine this, teeth were taken from each individual and DNA was extracted. This DNA was then compared to the known genetic sequence of 131 strains that were sequenced from the second and third plague pandemics. Finally the researchers created a maximum likelihood phylogenetic tree. The branch created for the two Justinian strains have no modern agents, and therefore the Y. pestis that caused the Justinian Plague is either extinct or the strains have yet to be documented in wild rodent populations. This specific strain of Y. pestis, which is blamed for the Justinian Plague, is distant from the strains that caused the Black Death and the Third pandemic.[4]
The Black Death
The Black Death is one of the best-known pandemics in human history. This plague is speculated to have begun in Asia and was brought to Europe through trade routes, most notably the famous Silk Road, approximately in 1343. The locations and timing of the Bubonic Plague can be seen in figure 2, which depicts the location and time of the plague across Europe. It has been demonstrated through research of DNA from bodies that date to the 14th century, that this plague was brought about by, once again, Yersinia pestis. The Black Death had a death toll of approximately 75 to 200 million people. With the plague killing such a large portion of the world’s population there were large shifts in religious and economic stages, in European and World history in general.[5]
Yersinia pestis Adaptation to Animal Host
The bubonic plague, even though it has had devastating role in past plagues, is still a relevant international health concern. The concern is also growing because there are new and multiple drug resistant strains of Y. pestis now in existence. The general method of infection begins with a bite from an infected flea. Figure 3 depicts an infected flea and the dark colored mass in the body of the flea is the bacterial infection. The bacterium is then taken up by macrophages in the animal host where it then replicates. Once the macrophage lyses it releases bacteria that attack the lymph nodes of the host where they once again replicate and lyse cells called polymorphonuclear neutrophils (PMN). When this happens the lymph nodes begin to get swollen and painful. This swelling gives the well known appearance of the bubo, which are the large boil looking lumps that are central to the bubonic plague. Once this bubo forms, the lymph node begins to get overwhelmed by the presence of bacteria, which then begin to enter the blood stream. Once bacteria enter the blood stream septicemia begins to set in, which generally leads to death unless intervention occurs. This general mechanism is well accepted and known but the exact genes used for this process are not well known.[10]
There has been a large amount of research into the specifics of the infection mechanism of Y. pestis due to the lethality and highly contagious characteristics of this bacterium. Elizabeth Pradel and her colleagues decided to attempt to uncover these genes that code for the virulence of Y. pestis. In order to begin this research the team built a store of mutants, where each mutant is missing at least one of the genes that have been previously identified as being those that are up regulated when the genes are in vivo. These mutant strains were created by allelic exchange. This process of allelic exchange requires a recombinase enzyme that allows PCR primers to be placed onto targeted genes.[12] These plasmids and the strains used fell under the broad function categories of metabolism, attachment, stress response and uncharacterized genes. Each of the mutant genes' genotypes was verified through PCR assays.[10]

In all of the strains that were tested, approximately 525 genes were discovered to be part of the virulence of the bacterium. In order to determine that these genes are part of virulence, the amount of activity of these genes is measured, and these genes were up-regulated by at least two times in the rat boils or bubos. Of these approximately 525 genes that are involved in the virulence of the plague, 79 of them were determined to be essential for pathogenesis. These 79 genes either encoded for the bacterium’s ability to create the bubo in the rat and/or are important to the bacterium’s ability to adapt to stress caused by the host rodent’s immune system. The stress that the rodent immune system puts on the bacteria is oxidative and iron stress. There are also important genes that help regulate and express specific genes. This was seen with the yebA gene when it was deleted it reduced the virulence of another mutant, ΔznuABC yupo2062. This interaction between these genes displays the interconnectedness of the genes that are required for the virulence of Y. pestis.[10]
After these specific 79 genes were tested there were still approximately 450 genes that displayed an up-regulation for the plague. For these remaining genes they were placed in specific mutant pools and tested together. In order to determine between these multiple mutants within a pool, each mutant was given a specific antibiotic resistance. These mutants were then counted in the corresponding lymph nodes and spleens of the rodents. This was done in order to identify specific genes that are important for both the initial and later stages of the infection. Most of these pools acted similarly to the wild type strains, which therefore display that at least one of the mutants in the pool are significant for virulence. [10]
Some of the important genes for infection are genes that encode proteins that are important for the bacterium's metabolism. The main carbon source used by the bacteria is carbohydrates, which are essential for the colonization of the host. The most crucial form of carbohydrates utilized by these bacteria are glucose and gluconate. In order to metabolize the glucose, glycolysis is used but when the initial genes for the beginning steps of glycolysis are knocked out glucose is unable to be broken down. The most important genes for glycolysis are those that encode for the latter portion of glycolysis. This was determined when the gene gpmA, the gene that encodes for the breakdown of 3-phosphoglycerate to 2-phosphoglycerate, one of the last steps in glycolysis.[10] The final product of glycolysis, pyruvate, moves into the tricarboxylic acid cycle (TCA cycle), but this cycle has been shown not to be essential for the virulence of the bacteria.
At the initial sequence of infection the bacteria comes into close contact with the polymorphonuclear neutrophiles, which release nitric oxide. This nitric oxide works against the bacteria by inducing oxidative stress. In order to combat this oxidative stress these bacteria use flavoglobin Hmp in order to detoxify these nitrogen species. One of the most important genes for this reaction to the oxidative stress is aceE. This gene is also important for the optimal cell growth within the host, which makes sense because there is a large amount of reproduction and growth within the lymph nodes near the polymorphonuclear neutrophiles. Another important gene that encoded for protection against reactive nitrogen species was nrdHIEF.[10] This was discovered when the gene was knocked out and the disease took considerably more time to set in.
Unlike the nitric stress that the bacteria encounter the researchers have found that the bacteria do not encounter much oxidative stress. Even though these bacteria do not encounter much oxidative stress they still contain some genes that encode to protect against the stress.[6]These genes encode for proteins that function in detoxifying the environment of the host. These genes were up regulated while the bacteria were replicating and growing in the macrophage cells.[7] This infection of the macrophage cells is considered to be one of the most important steps for the overall plague.[8] There are four essential genes that encode for protection against oxidative stress; these genes are mntH, fumC, yfiD, and ibpA. The mntH gene encodes for the ability to introduce manganese allowing for protection against oxidative stress. The fumC gene encodes for enzymes that repair certain genes that are destroyed by oxidative stress. When the ibpA gene is deleted in a mutant the virulence of the bacterium is reduced greatly.[10] This gene is also important for protecting against other stresses; one of these stresses is high temperature.[9] When the first three genes were deleted from the mutants the virulence of the bacterium was not reduced by much. Therefore these genes that are protecting against oxidative stress are not essential because there is not much oxidative stress produced by the host in guard.
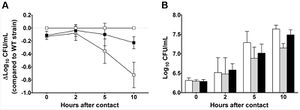
One of the most important genes found was gene ympt1.66c. This gene was found to be necessary for encoding for a putative helicase. When this specific gene was deleted, the ability of the bacteria to spread to the lymph node near the injection site is greatly reduced. Figure 4 depicts the effects of gene ympt1.66c, and what happens when this gene is mutated. The mutant of this gene is not able to initiate infection, which can be due to the mutants inability to take advantage of nutrients available. Another possibility is that the mutant is not able to survive when contact is made with PMN's. The researchers have seen evidence that suggests that horizontal gene transfer transferred this gene to the bacterium. Due to the function of this gene one can infer that horizontal gene transfer is important for these bacteria to initiate the infection in the early stages. This is most likely due to a large diminution of the bacterium’s ability to survive within the macrophage. Some of the important functions of the genes that were found encode for the bacterium’s ability to survive within the macrophages in the early parts of the infection. The methodology that Pradel and her colleagues used can further be applied to more and different pathogens, and also further study into Y. pestis will allow for a more general sense of bacterial pathogenicity mechanisms in other bacteria. Through this study it has been shown that there are only a few genes that are absolutely necessary in order to successfully infect their human and non-human animal hosts. One reason for the small amount of essential genes is the presence of alternative genes that sit in a type of reserve should any of the essential genes be knocked out by host’s immune defense. This is similar to the fact that there are also genes that have the same function but are useful under differing conditions, allowing the infection to express different genes under varying environmental conditions.[10]
Methods for Prevention and Vaccines
According to the Center for Disease control there a preventative measures that can be taken in order to limit the risk of contracting the plague caused by Y. pestis. One of the best ways is to exterminate any rodent populations that are living either in or around one’s home. Another suggestion made is to rid pets of fleas, which are known to be major carriers of the plague. There are vaccines that can be used to combat the plague and these vaccines have been around for quite some time, some dating back to around 1890. These vaccines are known to reduce the susceptibility of contracting the plague and by reducing the severity of the plague. The vaccine that is approved in the United States has been created from Y. pestis, which was incapacitated by formaldehyde. These vaccines are required for people who are involved in both field and laboratory work with the plague, and essential to field physicians who are treating those with the plague in an infected area.[11]
Conclusion
The bubonic plague caused by Y. pestis is known to have been one of the most devastating diseases in human history. It has been identified as the cause of the plague of Justinian, the Black Death, and the third plague pandemic. One of the most notable is the Black Death, which was an outbreak of the bubonic plague in 1300’s Europe and wiped out approximately a third of the population. In modern times this disease does not have as devastating an effect as it has had in history. There are still outbreaks in rural areas in Asia and South America but is much less disastrous when compared to the historical records. There have even been outbreaks in the Western United States. The preventative measures to be taken are to eradicate any rodent populations that exist near one’s home. There are also vaccines available but these are only required in locations of high plague incidence.
There has been a significant amount of research into Y. pestis due to its ability to cause bubonic and pneumonic plague. Some of the most recent research suggests that there are only a few genes that are absolutely essential for infection. This is because there are multiple genes for one function, these multiple copies allow for certain genes to be up regulated under certain environmental conditions. Due to this bacterium’s ability to adapt to its animal host environment it still poses a significant threat to the human population.
Fleas most often carry these bacteria, the most common being the Oriental rat flea; rodents then transport the flea over long distances. The fleas are able to infect both rodents and humans with Y. pestis. This is one of the main reasons why there is such a reoccurrence of the plague in rural areas that have high rodent infestations and have poor sanitation. These fleas can also be carried by pets which poses a substantial threat for urban communities. Overall the bacteria Yesinia pestis still poses a threat to the human population due to its ability to adapt to their host, and to be transmitted rapidly.
References
1. Perry RD; Fetherston JD (1997). “Yersinia pestis—etiological agent” Clin Microbiol Rev. 10(1): 35-36.http://cmr.asm.org/content/10/1/35.short
2. Bockemühl J (1994). "100 years after the discovery of the plague-causing agent--importance and veneration of Alexandre Yersin in Vietnam today". Immun Infekt 22 (2): 72–5.http://europepmc.org/abstract/med/7959865
3. http://historymedren.about.com/od/plagueanddisease/p/The-Sixth-century-Plague.htm
4. Wagner M, David. “Yesinia pestis and the Plague of Justinian 541-543 AD: a genomic analysis”. The Lancet Infectious Diseases. Apr. 2014: 319-326.http://www.sciencedirect.com/science/article/pii/S1473309913703232
5. Haensch, Stephanie; Bianucci, Raffaella; Signoli, Michel; Rajerison, Minoarisoa; Schultz, Michael; Kacki, Sacha; Vermunt, Marco; Weston, Darlene A.; Hurst, Derek; Achtman, Mark; Carniel, Elisabeth; Bramanti, Barbara (2010). "Distinct Clones of Yersinia pestis Caused the Black Death". PLoS Pathogens 6 (10): e1001134.http://journals.plos.org/plospathogens/article?id=10.1371/journal.ppat.1001134#ppat-1001134-g003
6. Sebbane F, Lemaitre N, Sturdevant DE, Rebeil R, Virtaneva K, et al. (2006) Adaptive response of Yersinia pestis to extracellular effectors of innate immunity during bubonic plague. Proc Natl Acad Sci U S A 103: 11766–11771. doi: 10.1073/pnas.0601182103.http://www.pnas.org/content/103/31/11766.short
7. Fukuto HS, Svetlanov A, Palmer LE, Karzai AW, Bliska JB (2010) Global gene expression profiling of Yersinia pestis replicating inside macrophages reveals the roles of a putative stress-induced operon in regulating type III secretion and intracellular cell division. Infect Immun 78: 3700–3715. doi: 10.1128/iai.00062-10.http://iai.asm.org/content/78/9/3700.short
8. ujol C, Bliska JB (2005) Turning Yersinia pathogenesis outside in: subversion of macrophage function by intracellular yersiniae. Clin Immunol 114: 216–226. doi: 10.1016/j.clim.2004.07.013.http://www.sciencedirect.com/science/article/pii/S1521661604002335
9. Kitagawa M, Miyakawa M, Matsumura Y, Tsuchido T (2002) Escherichia coli small heat shock proteins, IbpA and IbpB, protect enzymes from inactivation by heat and oxidants. Eur J Biochem 269: 2907–2917. doi: 10.1046/j.1432-1033.2002.02958.x.http://onlinelibrary.wiley.com/doi/10.1046/j.1432-1033.2002.02958.x/full
10. Pradel E, Lemaître N, Merchez M, Ricard I, Reboul A, et al. (2014) New Insights into How Yersinia pestis Adapts to Its Mammalian Host during Bubonic Plague. PLoS Pathog 10(3): e1004029. doi:10.1371/journal.ppat.1004029.http://journals.plos.org/plospathogens/article?id=10.1371/journal.ppat.1004029#ppat-1004029-g006
11. http://www.cdc.gov/mmwr/preview/mmwrhtml/00041848.htm (accessed on May 19th, 2015)
12. Datsenko KA, Wanner BL (2000) One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci U S A 97: 6640–6645. doi: 10.1073/pnas.120163297 http://www.pnas.org/content/97/12/6640
