Beyond the Basics: Genetic, Intestinal, and Viral Contributors to Autism
Introduction
Autism spectrum disorder (ASD) is a comprehensive label for a wide range of diagnoses and accompanying manifestations. A diagnosis of ASD typically comes after a comprehensive psychological examination and possibly assessment of other clinical symptoms, such as gastrointestinal distress. Common traits associated with autism include social communication challenges and restrictive and repetitive behaviors. [1] It is important to note that these traits vary widely between individuals. Additionally, most of the current research has been conducted on males; females may exhibit very different manifestations. [2]Currently approximately one in 54 children are diagnosed with autism, with males contributing a substantially larger proportion of this figure. [3]This statistic is also subject to the clinical and diagnostic bias surrounding autism and gender, and the subjectivity of what is classified as autism spectrum disorder.[2]
Genetics
Polygenetic Inheritance
Autism is a heterogeneous disorder, meaning that there are several factors - which may be dramatically different across ASD individuals - that can lead to a diagnosis.[4] Associated with a heterogeneous disorder is polygenic inheritance, the combined effect of multiple polymorphisms to have a large effect on an individual.[5] Possible variation as to the type and number of polymorphisms that an autistic individual receives from their parents during recombination partially explains the wide spectrum in the presentation of autistic traits. “De novo” mutations occur randomly in germ line cells; resultantly, they produce a phenotype in the offspring that is not present in the parental generation. These de novo mutations are frequently copy number variants, (CPVs) - duplications or deletions - single nucleotide variations, (SNVs) or likely-gene-disrupting (LGD) mutations - causing premature stop codons, frameshifting, or altered splicing sites.[6]Once established in the genome, mutations can be inherited by progeny from sexual reproduction. An accumulation of these mutations over multiple generations may eventually result in a child presenting with ASD disorder. If this is the case, their family members, who have some of the same heritable mutations, will likely have some ASD traits but be on the ‘higher functioning’ end of the spectrum.[6]
Individual Genes of Interest
Infrequently, one mutation alone is enough to greatly increase the risk of an individual presenting with autistic traits.[4] Two of the most well-known of these mutations are CPVs of the 16p11.2 locus and 22q11.2 loci.[4] Relative to individuals with heterogeneous autism (here referring to those diagnosed with ASD but not possessing either of the aforementioned CPVs, thus the individual's presentation is considered to be due in much greater part to polygenic inheritance), the 16p11.2 deletion and duplication individuals and the 22p11.2 deletion individuals have a differing mean presentation; the 22p11.2 individuals have statistically insignificant differences from the heterogeneous group.[4] Within the two CPV groups themselves, there is a drastic variability in the function of individuals across social, emotional, and intellectual domains, up to 97% within a group with the same single-locus abnormality.[4]
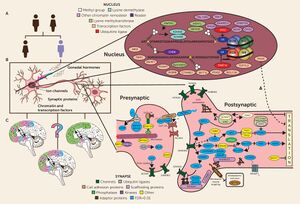
16p11.2
A critical role of the 16p11.2 loci is the regulation of the kynurenine pathway (KP). The kynurenine pathway is a subsect of the metabolic degradation of the amino acid tryptophan.[7] The metabolites produced in this pathway have been linked to multiple neurological and developmental disorders including autism.[8] Quinolinate phosphoribosyltransferase (QPRT) is an essential protein in this metabolic pathway because it catabolizes quinolinic acid (QA), a compound that is cytotoxic at high levels.[9] Increased concentrations of QA are found in ASD patients when compared to control groups.[10] Because QPRT is located on the 16p11.2 gene, a deletion at this loci reduces the QPRT concentration in neurons and glial cells. Consequently QA levels rise, possibly causing or exacerbating ASD symptoms.
22p11.2
22p11.2 is associated with several other conditions including schizophrenia and anxiety, immunodeficiency disorders, and learning disabilities.[11] Coexisting diagnoses and overlapping traits may make it difficult to discern what elements of a patient’s phenotype are due to autism and its relative severity and what parts are a consequence of alternative disorders.
Treatment
Progress in genetic sequencing may make it possible for this clinical testing for autism to be supplemented with an evaluation of the genome of the patient as well. It is critical to note that this does not mean that genetic testing could confirm or deny an ASD diagnosis. For individuals with ASD that is contributed to through haploinsufficiency, it is not inconceivable that there might be some benefit to targeted gene therapy that would supplement the missing gene product, but right now this idea is merely speculative.[5]
Gut Microbiome
Although autism has most commonly been associated with strictly neurological abnormalities, recent research points to the intestine as a significant contributor to the disorder. 23-70% of patients with ASD also present with gastrointestinal (GI) dysfunction, which often manifests as increased intestinal permeability and digestive issues.[13][14] [15] Additionally, relating to GI dysfunction is the gut-brain axis, another likely influential factor in autism. The axis is a relatively new area of research, but already many studies are demonstrating bidirectional communication between the brain and the intestine.[16] Increases in autistic speech, social, and behavioral challenges, as well as aggression and irritability, have been observed to trend with the severity of GI dysfunction, demonstrating the link between GI issues and the gut-brain axis.[17]
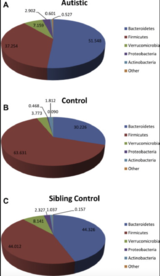
Answers to how and why the gut plays a role in ASD may be found within the gut microflora. The intestinal microbiome hosts a signature pattern to the types and amounts of bacteria present; maintaining proper homeostasis of these colonies is crucial for both disease prevention and normal neurological development.[18][19] ASD patients frequently present with dysbiosis in their microflora: a comparison of ASD presenting children and their siblings found a significant increase in Bacteroidetes and decrease in Firmicutes.[12][20] These findings are supported by rodents with abnormal populations of certain gut microbiota having altered emotional and physical development.[16][21] Of note are bacterial orders Clostridia and Bacteroidia, members of the two aforementioned phylums, which show the greatest diversity between ASD and presenting mice and control mice.[15] Microbes in greater quantities in ASD mice were suspected of contributing to pathogenesis, whereas those that were deficient might promote normal activity.[15]
Re-regulation of irregular microbe levels may be an avenue of future treatments for ASD. Mice bred to exhibit ASD traits with associated irregular gut microbe composition were injected with Bacteroides fragilis. The treated rodents experienced alleviated ASD symptoms along with improved gut barrier integrity.[15] Further research is needed to determine whether similar microbe supplementation for ASD individuals could establish normal intestinal flora and subsequently relieve ASD symptoms involved in the gut-brain axis and GI issues.
Vaccines
Even today, a sizable population - one study claims approximately 20% of adults in the 38 participating countries - falsely believe that vaccines cause autism.[22] While there are many contributing factors to this belief, three scientific misconceptions form the basis of this myth. The measles-mumps-rubella (MMR) vaccine became antagonized in 1998 when Dr. A. J. Wakefield’s hypothesis about the correlation between the MMR vaccine, gastrointestinal issues, and autism, was prematurely released as fact in media outlets.[23][24] Furthermore, in 1997 the FDA released guidelines to regulate the ingestion of thimerosal, a mercury-containing compound found in many vaccines, and in 1999 mandated its removal from vaccines.[25] Subsequent studies found no negative impact regarding thimerosal in vaccines, but the idea about a possible link between vaccines and autism was already in circulation.[26] A portion of the public only absorbed the news of a potentially dangerous chemical being injected via vaccination. Finally, in 2008 an anecdotal account of a 19-month-old girl who experienced a negative reaction to the MMR vaccine and months later had developmental and neurological abnormalities drew attention when it became a national court case. [24]This incident fueled the belief that an overload of vaccines at one time could cause autism. All of these factors, combined with more modern issues such as political divisions, have led to widespread misconceptions that have continued perpetuating to this day.[27] Despite the plethora of misinformation, over a dozen studies have validated the safety of vaccines and concluded that there is no link between vaccines and autism.[28]
Conclusion
Understanding the pathology of autism is fundamental in establishing the way that it is perceived in society. The rhetoric of negligent parenting raising robotic individuals is an antediluvian manner of thinking that doesn’t take into account the multitude of factors that contribute to autism, including genetics, the gut microbiome, and maternal viral infections. [29] Although it is highly unlikely that a ‘cure’ for autism is in the future, the proliferation of new research means that there is an increasing number of treatment options available, ranging from traditional DBT therapies to more experimental supplements to reestablish proper gut microflora. However, above all else, it is essential to view autism as it is - a complex spectrum encompassing a wide range of individuals, with - instead of a problem that needs to be fixed. All of the positive qualities that ASD individuals have, not just their challenges, deserve acknowledgment. Indeed, some of the most notable names today and in history belong to people who would today likely be classified as autistic: Susan Boyle, Emily Dickenson, Albert Einstein, Bill Gates, Steve Jobs, Thomas Jefferson, Michelangelo, Wolfgang Amadeus Motzart, ad Nikola Tesla.[30] Future research will hopefully unlock more of the intricacies of autism and help autistic individuals be better understood both by society and by themselves.
References
Edited by Juliette Lowe, student of Joan Slonczewski for BIOL 238 Microbiology, 2021, Kenyon College.
- ↑ [Mayo Clinic Staff. (2018, January 6). Autism spectrum disorder - Symptoms and causes. Mayo Clinic. Retrieved November 16, 2021, from https://www.mayoclinic.org/diseases-conditions/autism-spectrum-disorder/symptoms-causes/syc-20352928]
- ↑ 2.0 2.1 [Ratto, A. B., Kenworthy, L., Yerys, B. E., Bascom, J., Wieckowski, A. T., White, S. W., Wallace, G. L., Pugliese, C., Schultz, R. T., Ollendick, T. H., Scarpa, A., Seese, S., Register-Brown, K., Martin, A., & Anthony, L. G. (2018). What About the Girls? Sex-Based Differences in Autistic Traits and Adaptive Skills. Journal of autism and developmental disorders, 48(5), 1698–1711. https://doi.org/10.1007/s10803-017-3413-9]
- ↑ [Maenner, M. J. (2020, March 27). Prevalence of Autism Spectrum Disorder Among Children Aged 8 . . . Centers for Disease Control and Prevention. Retrieved November 16, 2021, from https://www.cdc.gov/mmwr/volumes/69/ss/ss6904a1.htm?s_cid=ss6904a1_w]
- ↑ 4.0 4.1 4.2 4.3 4.4 [Chawner, S. J., Doherty, J. L., Anney, R. J., Antshel, K. M., Bearden, C. E., Bernier, R., ... & van den Bree, M. B. (2021). A genetics-first approach to dissecting the heterogeneity of autism: phenotypic comparison of autism risk copy number variants. American Journal of Psychiatry, 178(1), 77-86. https://ajp.psychiatryonline.org/doi/abs/10.1176/appi.ajp.2020.20010015]
- ↑ 5.0 5.1 5.2 [Manoli, D. S., & State, M. W. (2021). Autism spectrum disorder genetics and the search for pathological mechanisms. American Journal of Psychiatry, 178(1), 30-38. https://ajp.psychiatryonline.org/doi/10.1176/appi.ajp.2020.20111608]
- ↑ 6.0 6.1 [Yoon, S., Munoz, A., Yamrom, B. et al. Rates of contributory de novo mutation in high and low-risk autism families. Commun Biol 4, 1026 (2021). https://doi.org/10.1038/s42003-021-02533-z]
- ↑ [Badawy AA-B. Kynurenine Pathway of Tryptophan Metabolism: Regulatory and Functional Aspects. International Journal of Tryptophan Research. January 2017. https://doi.org/10.1177%2F1178646917691938]
- ↑ [Bryn, V., Verkerk, R., Skjeldal, O., Saugstad, O., Ormstad, H., Gillberg Neuropsychiatry Centre, Sahlgrenska akademin, Göteborgs universitet, Gothenburg University, Gillbergcentrum, & Sahlgrenska Academy. (2018;2017;). Kynurenine pathway in autism spectrum disorders in children. Neuropsychobiology, 76(2), 82-88. https://doi.org/10.1159/000488157]
- ↑ [Braidy, N., Grant, R., Adams, S. et al. Mechanism for Quinolinic Acid Cytotoxicity in Human Astrocytes and Neurons. Neurotox Res 16, 77–86 (2009). https://doi-org.libproxy.kenyon.edu/10.1007/s12640-009-9051-z]
- ↑ [Lim, C. K., Essa, M. M., de Paula Martins, R., Lovejoy, D. B., Bilgin, A. A., Waly, M. I., Al-Farsi, Y. M., Al-Sharbati, M., Al-Shaffae, M. A., & Guillemin, G. J. (2016). Altered kynurenine pathway metabolism in autism: Implication for immune-induced glutamatergic activity. Autism Research, 9(6), 621-631. https://doi.org/10.1002/aur.1565]
- ↑ [Du, Q., de la Morena, M. T., & van Oers, N. S. (2020). The genetics and epigenetics of 22q11. 2 deletion syndrome. Frontiers in genetics, 10, 1365. https://www.frontiersin.org/articles/10.3389/fgene.2019.01365/full]
- ↑ 12.0 12.1 [Finegold, S. M., Dowd, S. E., Gontcharova, V., Liu, C., Henley, K. E., Wolcott, R. D., ... & Green III, J. A. (2010). Pyrosequencing study of fecal microflora of autistic and control children. Anaerobe, 16(4), 444-453. https://www.sciencedirect.com/science/article/pii/S1075996410001010]
- ↑ [Chaidez, V., Hansen, R.L. & Hertz-Picciotto, I. Gastrointestinal Problems in Children with Autism, Developmental Delays or Typical Development. J Autism Dev Disord 44, 1117–1127 (2014). https://doi.org/10.1007/s10803-013-1973-x]
- ↑ [de Magistris, Laura*; Familiari, Valeria*; Pascotto, Antonio†,#; Sapone, Anna*; Frolli, Alessandro†; Iardino, Patrizia§; Carteni, Maria‡; De Rosa, Mario‡; Francavilla, Ruggiero¶; Riegler, Gabriele*; Militerni, Roberto†; Bravaccio, Carmela|| Alterations of the Intestinal Barrier in Patients With Autism Spectrum Disorders and in Their First-degree Relatives, Journal of Pediatric Gastroenterology and Nutrition: October 2010 - Volume 51 - Issue 4 - p 418-424 https://journals.lww.com/jpgn/Fulltext/2010/10000/Alterations_of_the_Intestinal_Barrier_in_Patients.6.aspx]
- ↑ 15.0 15.1 15.2 15.3 [Hsiao, E. Y., McBride, S. W., Hsien, S., Sharon, G., Hyde, E. R., McCue, T., Codelli, J. A., Chow, J., Reisman, S. E., Petrosino, J. F., Patterson, P. H., & Mazmanian, S. K. (2013). Microbiota modulate behavioral and physiological abnormalities associated with neurodevelopmental disorders. Cell, 155(7), 1451–1463. https://www.sciencedirect.com/science/article/pii/S0092867413014736]
- ↑ 16.0 16.1 [Carabotti, M., Scirocco, A., Maselli, M. A., & Severi, C. (2015). The gut-brain axis: interactions between enteric microbiota, central and enteric nervous systems. Annals of gastroenterology, 28(2), 203–209. https://pdfs.semanticscholar.org/8758/c70237699371670eb2ebe88f3bf6c3f26496.pdf]
- ↑ [Adams, J.B., Johansen, L.J., Powell, L.D. et al. Gastrointestinal flora and gastrointestinal status in children with autism – comparisons to typical children and correlation with autism severity. BMC Gastroenterol 11, 22 (2011). https://doi.org/10.1186/1471-230X-11-22]
- ↑ [Lin, L., Zhang, J. Role of intestinal microbiota and metabolites on gut homeostasis and human diseases. BMC Immunol 18, 2 (2017). https://doi.org/10.1186/s12865-016-0187-3]
- ↑ [Lu, J., & Claud, E. C. (2019). Connection between gut microbiome and brain development in preterm infants. Developmental psychobiology, 61(5), 739-751. https://onlinelibrary.wiley.com/doi/full/10.1002/dev.21806]
- ↑ [Parracho, H. , Bingham, M. , Gibson, G. & McCartney, A. (2005). Differences between the gut microflora of children with autistic spectrum disorders and that of healthy children. Journal of Medical Microbiology, 54 (10), 987-991. https://www.microbiologyresearch.org/content/journal/jmm/10.1099/jmm.0.46101-0?TRACK=RSS]
- ↑ [Bravo, J. A., Forsythe, P., Chew, M. V., Escaravage, E., Savignac, H. M., Dinan, T. G., Bienenstock, J., & Cryan, J. F. (2011). Ingestion of Lactobacillus strain regulates emotional behavior and central GABA receptor expression in a mouse via the vagus nerve. Proceedings of the National Academy of Sciences of the United States of America, 108(38), 16050–16055. https://doi.org/10.1073/pnas.1102999108]
- ↑ [Duffy, B. (2018, August 22). Autism and vaccines: more than half of people in Britain, France, Italy still think there may be a link. The Conversation. Retrieved November 6, 2021, from https://theconversation.com/autism-and-vaccines-more-than-half-of-people-in-britain-france-italy-still-think-there-may-be-a-link-101930]
- ↑ [Wakefield, A. J., Murch, S. H., Anthony, A., Linnell, J., Casson, D. M., Malik, M., ... & Walker-Smith, J. A. (1998). RETRACTED: Ileal-lymphoid-nodular hyperplasia, non-specific colitis, and pervasive developmental disorder in children. https://www.sciencedirect.com/science/article/pii/S0140673697110960]
- ↑ 24.0 24.1 [Offit, P. A., & Coffin, S. E. (2003). Communicating science to the public: MMR vaccine and autism. Vaccine, 22(1), 1-6. https://www.sciencedirect.com/science/article/pii/S0264410X03005322]
- ↑ [Centers for Disease Control Prevention. Thimerosal in vaccines: a joint statement of the American Academy of Pediatrics and the Public Health Service, MMWR Morb Mortal Wkly Rep, 1999, vol. 48 (pg. 563-5) https://publications.aap.org/pediatrics/article/104/3/568/62479/JOINT-STATEMENT-OF-THE-AMERICAN-ACADEMY-OF]
- ↑ [Thompson WW, Price C, Goodson B, et al. Early thimerosal exposure and neuropsychological outcomes at 7 to 10 years, N Engl J Med, 2007, vol. 357 (pg. 1281-92) https://www.nejm.org/doi/full/10.1056/NEJMoa071434]
- ↑ [Larson, H. J. (2018). Politics and public trust shape vaccine risk perceptions. Nature human behaviour, 2(5), 316-316. https://www.nature.com/articles/s41562-018-0331-6]
- ↑ [Dudley M.Z. et al. (2018) Do Vaccines Cause Autism?. In: The Clinician’s Vaccine Safety Resource Guide. Springer, Cham. https://doi.org/10.1007/978-3-319-94694-8_26]
- ↑ [Kanner, L. (1943). Autistic disturbances of affective contact. Nervous child, 2(3), 217-250. http://mail.neurodiversity.com/library_kanner_1943.pdf]
- ↑ [History's 30 most inspiring people on the autism spectrum. Applied Behavior Analysis Programs Guide. (2017, September 22). Retrieved November 21, 2021, from https://www.appliedbehavioranalysisprograms.com/historys-30-most-inspiring-people-on-the-autism-spectrum/]
