Chlamydophila psittaci
A Microbial Biorealm page on the genus Chlamydophila psittaci
Classification
Higher order taxa
Superkingdom: Bacteria, Superphylum: Chlamydiae/Verrucomicrobia group, Phylum: Chlamydiae, Class: Chlamydiae, Order: Chlamydiales, Family: Chlamydiaceae, Genus: Chlamydophila; Species: Chlamydophila psittaci (4)
Species
Chlamydophila psittaci
previously known as Chlamydia psittaci
Description and significance
Chlamydophila psittaci is an obligate, intracellular, gram negative bacteria that occur as a parasite in eukaryotic cells. These cells are coccoid and non-motile, with sizes ranging from 0.2-1.5 m. The cylamydial cell envelope lacks peptidoglycan, but instead has an outer membrane containing lipopolysaccharide and a cytoplasmic membrane bilayer. (1,6,7)
Chlamydophila psittaci causes a systemic infectious disease, psittacosis, in the parrot family and other avian species. Chlamydophila psittaci is present in feces, nasal secretions, and feathers of infected birds. This bacteria may be transmitted to humans through inhalation of dust from the contaminated bird. In 1930, the largest epidemic of psittacosis affected 750-800 individuals, which lead to the isolation of Chlamydophila psittaci in Europe and the United States. A total of 923 human cases of psittacosis have been reported to the US Centers for Disease Control and Prevention from 1988 through 2003. The term psittacosis is derived from the Greek word for parrot. (2,6,7)
For many years, Chlamydophila psittaci was detected through the isolation of the organism through cell culture, which required scraping of bacterial cells from the site of infection of patients. New techniques of polymerase chain reaction and ligase chain reaction has improved detection of these specimens. These new diagnostic techniques involves fluorescence microscopy and enzyme-linked immunoassays. (6)
Chlamydophila psittaci infection may be treated through antimicrobial therapy such as tetracycline, doxycycline, erythromycin, and sulfonamides. Mortality rate prior to antimicrobial treatment was approximately 15-20% and has decreased to less than 1% with appropriate antibiotic therapy. (3,6)
Genome structure
The complete genomic DNA sequences of Chlamydophila psittaci is still in progress of sequencing. The genomic DNA sequences of Chlamydiaceae have been published for C. trachomatis, C. muridarum, C. pneumoniae, C. caviae, and C. abortus. It is known that the family of Chlamydiaceae genomes are fairly conserved. The chromosome of Chlamydophila psittaci is therefore likely to be circular. The size and content of the genome has yet to be determined. Strains 6BC and E/B have been identified. Chlamydophila psittaci contains a circular plasmid strain, pCpA1, that has been sequenced to be 7,553 nucleotide long. This plasmid was sequenced in 11/15/1991 at the University of Manchester. The plasmid contains genes to encode for the major outer membrane proteins. (5,9,12)
It is known that Omp genes encode for the outer membrane proteins, or polymorphic membrane proteins, and is important for the characteristic of Chlamydophila. The Omp1 outer membrane protein is involved in attachment and entry into host cells. The Omp2 cysteine-rich protein is important for reticular body to elementary body conversion. (6,14)
Cell structure and metabolism
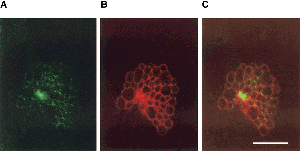
Chlamydophila psittaci have major outer membrane proteins (MOMP), consisting of predominately Beta-sheet content, similar in function to the biochemical properties of porin protein. These channels are permeable to ATP and may be the route in which the bacterium takes advantage of nucleoside triphosphates. Although Chlamydophila psittaci have no known mechanism to metabolize glucose as a source of energy, it is known that Chlamydophila psittaci can obtain ATP and essential amino acids from the host cell. (6,7,8)
Chlamydophila psittaci cannot synthesize pyrimidines or purines de novo, so it must salvage these from its host. Though it can take up ATP, GTP, and TTP directly from host cells. It may not, however, take up UTP, CTP, dATP, dGTP, or dCTP from the host cell, and may not salvage exogenously supplied uridine, cytidine, or deoxycytidine. The primary source of pyrimidine nucleotides is via the salvage of uracil by a uracil phosphoribosyltransferase. The ATP-GTP transport system and intracelullar enzymes allows for metabolism of adenine, adenosine, guanosine, and guanine. The TTP transport system and intraceullar enzymes allows for the metabolism of uracil, deoxyuridine, thymine, and thymidine. In addition, the parasite could incorporate exogenous thymidine and thymine into DNA. There is no interconversion of nucleotides within the cell. (13,14)
Ecology
Chlamydophila psittaci is a pathogen which interacts with parrots, parakeets, canaries, and other avian species. The bacteria may be transmitted to humans through handling of sick birds. Human to human transfer is uncommon, although possible. Certain breeds of Chlamydophila psittaci may infect sheep, goats, and cows. This species does not have any direct contribution to the environment because it cannot live outside its host. (3)
Pathology
Chlamydophila psittaci causes an infection through the respiratory system by using chlamydial elementary bodies to attach to the respiratory epithelial cells of the host and is engulfed through phagocytosis. Elementary bodies spread via the blood steam to the reticuloendothelial system and become reticulate bodies, which depend on host cell ATP to grow. Within the inclusion of the host cell, reticular bodies undergo binary fission for 8-10 hours after infection, and continue to divide for 20 hours. Reticular bodies give rise to elementary bodies after 20 hours of infection. After 48-72 hours, the development cycle is completed and the infected host cell’s inclusion becomes filled with 10-1000 elementary bodies. Elementary bodies are released through lysis of the cell and may then infect fresh host cells. (3,6,7)
Chlamydophila psittaci affects the parrot family and other avian species. Once infected, these species will be susceptible to symptoms such as appetite and weight loss, diarrhea, sinusitis, and respiratory distress. Humans contract the disease through handling sick birds. Symptoms in humans include fever, cough, dyspnea, mild phryngitis, epistaxis, severe headache, and pneumonia. Certain strains of Chlamydophila psittaci may also infect sheep, goats, and cows. (3,6)
Application to Biotechnology
Chlamydophila psittaci is not known to produce any useful compounds or enzymes. Due to its harmful pathogenic nature, studies have been developed to find proper vaccination against this bacteria. Current research have confirmed that the major outer membrane protein gene of Chlamydophila psittaci may serve as a target for vaccination. (10)
Current Research
July 2007 - Construction and immunogenicity of recombinant adenovirus expressing the major outer membrane protein (MOMP) of Chlamydophila psittaci in chicks.
The first study explores the use of the major outer membrane protein encoded by the outer membrane protein 1 (omp1) gene, as a candidate for a vaccine against avian chlamydiosis. Pathogen free chicks were inoculated. Detection through an indirect hemagglutination test showed that these chicks generated antibodies against the major outer membrane protein of Chlamydophila psittaci. 9 out of 10 vaccinated chicks were protected from the disease when challenged, whereas the control groups showed clinical signs of the disease when challenged. This study shows that the major outer membrane protein gene of Chlamydophila psittaci may serve as a candidate vaccine against avian chlamydiosis. (10)
March 2007 - Chlamydiae and Atherosclerosis: Can Psittacine Cases Support the Link?
The second study examines whether Atherosclerosis, a common disease in birds, is caused by the pathogen Chlamydophila psittaci. Out of 103 cases of advanced stages of atherosclerosis, only 4 (3.9%) were positive of chlamydiae in atherosclerotic tissue. Sequential analysis revealed high correlation (94%-100%) with Chlamydophila psittaci in three of the four cases. Because of the low occurrence of chlamydiae in atherosclerotic tissue (3.9%) during advanced stages of Atherosclerosis, a relationship between chlamydiae and atherosclerosis in pet birds is improbable. (11)
August 2007 - Chlamydophila psittaci genotype E/B transmission from African grey parrots to humans.
The third study explores Chlamydophila psittaci genotype E/B transmission from parrots to humans. The presence of Chlamydophila psittaci was examined in a parrot relief and breeding center. 5 of 20 African grey parrots showed depression, ruffled feathers, loss of weight and mild dyspnoea. The manager also showed signs of shortness of breath. Chlamydophila psittaci genotype E/B was identified as the transmitted strain. It is believed that this is the first genotype E/B strain transmitted from parrot to human. Studies reveal that inheritance of the genotype E/B strain, in both parrots and humans, cause no severe clinical symptoms.(12)
References
3. Lessnau, K., Arjomand F., “Psittacosis” 2006 May
5. Chlamydophila psittaci plasmid pCpA1, complete sequence
6. Lederberg, J., “Encyclopedia of Microbiology Second Edition A-C.” 2000. Volume 1. p. 781-787.
7. Singleton, P., Saisbury, D. “Dictionary of Microbiology and Molecular Biology 3rd Edition.” 2001. p 154.
11. Schenker, OA, Hoop, RK. “Chlamydiae and atherosclerosis: can psittacine cases support the link? 2007
Edited by Coleman Ho, student of Rachel Larsen
Edited by KLB
