Comamonas badia
Classification
Higher Order Taxa
Bacteria (Kingdom), Proteobacteria (Phylum), Betaproteobacteria (Class), Burkholderiales (Order), Comamonadaceae (Family), Comamonas(Genus)
Species
Comamonas badia
Description
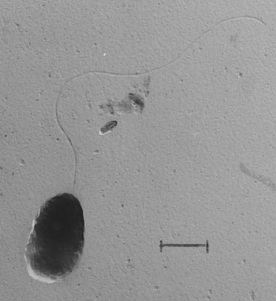
Comamonas badia is a gram negative, rod-shaped, strictly aerobic, and highly motile bacteria with a polar flagellum. It was isolated from phenol-adapted activated sludge from a factory sludge from Japan [1]. It was originally classified under the genus Pseudomonas [3][4]. Based on 16S rRNA gene sequence and G+C content, C. badia was determined to represent its own genus, Comamonas. This novel one was first represented by the species C. badia, type-strain IAM 14839T [1][9].
Comamonas badi forms pink colonies when cultured in Luria slant media and forms visible flocs in Luria broth media, unlike other Comamonas species which do not form flocs [2]. Comamonas badia grows at a very slow rate in a flocculated state with a generation time of about 12 hours at optimal temperatures (27-30°C) in LB media and produces few free-floating cells. When cultured in broths, the supernatant is usually clear [4]. This bacterium grows the slowest at 20°C (19 hours for generation time) and does not grow at 37°C (average human body temperature), unlike other Comamonas species which do undergo growth at 37°C [1]. For floc formation, C. badia grows embedded in a film mesh structure made out of mucopolysaccharides which consists of glucosamine, glucose, mannose, galactose and rhamnose. Only about 10% of the total polysaccharide content found with C. badia is involved with floc formation [3]. The ability to form flocs gives this species a protection advantage from predators compared to free roaming cells, allows other bacteria to form on the flocs, and prevents the organism from being washed out of the system, i.e. activated sludge [10].
Significance
It has been found that C. badia has moderate phenol tolerance and moderate phenol degradation capabilities. C. badia existing in a floc-formation is also the most predominating bacteria species in activated sludge and is involved in degrading highly toxic and carcinogenic phenol compounds in coke wastewater [2]. Being able to tolerate and degrade such toxic compounds, the ability to form flocs is an integral part this microbe in activated sludge to treat wastewaters [10]. C. badia has a unique ability in which it is responsible for its own deflocculation by producing an exoenzyme. Various commercial enzymes are unable to induce deflocculation in C. badia and therefore the bacteria’s life activities are determined by their own accord via polysaccharide-synthesizing enzymes and deflocculating enzymes [5].
Phylogenetics
It was only recently in 2004 that C. badia was introduced and recognized as a novel species, using comparative analysis of almost complete 16S rDNA gene sequence and G+C content of DNA on the strain IAM 14839T. The complete genome of C. badia has yet to be sequenced [1][9].
Cell Structure
C. badia are gram negative, highly motile, 0.8-0.9 × 1.0-2.5 μm, and rod-shaped bacterium that appears in singles or pairs in young cultures. They grow very slowly in a flocculated state (12+ hours for doubling time) and have a single polar flagellum for motility. C. badia are catalase and oxidase-positive, heterotrophic organisms that do not use glucose, mannose, arabinose, or sucrose for growth, but do use glutamic acid or aspartic acid and is able to reduce nitrate [1][6]. They are capable of growing in cultures without growth factors and do not hydrolyze gelatin. C. badia contains the ubiquinone Q-8 as the predominant component of the quinone system for electron transport. Comamonas badia is largely composed of exopolysaccharides (EPS), containing large amounts of straight-chain unsaturated fatty acids, cis-7-hexadecenoic acid (16:1ω7c) and saturated fatty acid, hexadecanoic acid (16:0) [1]. Large amounts of these EPS contribute to the high EPS to protein ratio, which are commonly found in bacteria within activated sludge [7]. This ratio also gives the flocs formed by this microbe a high hydrophobic and negative charge characteristic. A high hydrophobic property helps microbes form flocs easier and is normally affected by growth rate as well [7][8].
Ecology
C. badia was isolated and found in phenol-adapted activated sludge in a factory plant in Japan used for decomposing industrial wastes. This species is able to thrive in such environments not only due to its ability to degrade phenols but also its ability to grow and tolerate an environment with up to 500 mg/L of phenols. It may have a prominent role in structure formation of the activated sludge ecology as C. badia is the most predominate species in activated sludge, roughly 32-66% of total peak area of the community [2]. C. badia can also form a productive interaction with phenol-decomposing bacteria Strain No. 3 from the activated sludge to greatly enhance phenol degradation and floc formation abilities. High phenol-decomposing bacteria such as Strain No. 3 requires floc formation or floc incorporation in order to degrade phenols efficiently. Unable to form flocs on its own, Strain No. 3 is usually incorporated with C. badia and is the only strain that is able to from a floc formation with C. badia. This combination of Strain No. 3 and C. badia in a ratio of 1:6, respectively, in a floc is able to decompose phenol concentrations as high as 700 ppm rapidly without deflocculating for over 50 hours. However, Strain No. 3 bacteria must be incorporated into the C. badia floc rather than interacting with the floc as free roaming cells. The latter method will result with displays of slow and non-effective phenol decomposition and deflocculation after the 48 hour incubation point. Strain No. 3 and C. badia may have a nutritional interaction and C. badia may provide a structural advantage for Strain No.3 as a floc allowing such combination to decompose high levels of phenol with the presence of peptone [4]. Such positive interaction between the bacteria may provide useful environmental interaction in decomposing harmful toxic substances such as phenols in industrial wastewater such as coke wastewater [2].
Unique Deflocculation of Comamonas badia
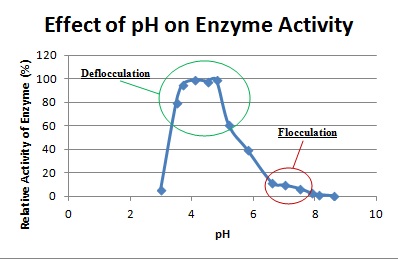
Not only is C. badia capable of producing its own exoenzyme to induce deflocculation, it also has the ability to resist deflocculation by various commercial cellulases, proteases, lysozymes and EDTA. However, the species lose this resisting ability if the cell cultures were dialyzed before treatment with commercial enzymes. The self produced exoenzyme, which is an exopolymer-degrading enzyme or other known as a polysaccharide hydrolase, has been isolated and purified to test its functions and characteristics. When the activity of hydrolysis of polymers is higher than the synthesis of polymers, deflocculation occurs. The optimal pH for deflocculation is around the range of 3.8 – 5.0, which is far from the normal pH of flocculation (7.0). At normal flocculation pH, only less than 10% of enzymatic activity was observed from the hydrolase. The exoenzyme maintains stable conformation in normal temperatures until 35°C and becomes completely inactive if left at a temperature of 45°C for 5 minutes. The exoenzyme’s activity is also inhibited at buffer ionic strengths higher than 0.01, and in mixtures of basic or amino acids at concentrations higher than 10 mM. Amino sugars and uronic acids also inhibits the enzyme at 10 mM concentrations. Glucosamine however, effectively inhibits enzymatic activity at 1 mM concentrations. Although this powerful enzyme was able to induce C. badia deflocculation by degrading and releasing sugars, it cannot hydrolyze cellulose, pectin, or ethylene glycol chitin [5].