Deinococcus deserti

Description and Significance
Deinos is a Greek derivation which means strange or unusual. Deinococcus is an “unusual coccus”. Initially placed in the genus Micrococcus, Brooks and Murray suggested that they make up their own genus Deinococcus [1] . They have been isolated from a wide range of environments such as soil, water, air, faeces, hot springs and irradiated food [2] . Deinococcus are characterized with the ability to cope with exceptional levels of DNA-damaging agents such as ionizing radiation, UV light and desiccation [2] .
According to NCBI, as of November 23, 2012, there are 46 characterized species of Deinococcus.
Deinococcus deserti was recently isolated from mixed surface sand samples from the Sahara desert [3] . The Sahara desert is an extreme environment, where any surviving organisms would’ve had to adapt to harsh conditions of UV irradiation, cycles of high and low temperature, desiccation and nutrient limitation [3] . These sand samples from which D. deserti were isolated were also “enriched” for radiation tolerance with exposure to 15kGy gamma-radiation [3] . Based on 16S rRNA gene sequence and DNA-DNA hybridization, the novel strain D. deserti was shown to belong to the genus Deinococcus [3] . Like the other members of the genus Deinococcus, D. deserti are non-pathogenic and non-motile [5] .
Deinococcus deserti grow as whitish, circular, uniform-edged colonies of 0.5-1mm when grown on trypticase soy broth (TSB) ten-fold diluted for 72 h at 30˚C [6] . The other Deinococcus type strains usually grow as pink or red colonies on TSB rich media plates, but D. deserti showed no growth on rich media [6] . Growth was also observed at 37˚C, slow growth at 23˚C, and no growth at 45˚C [3] . Growth occurred between pH of 6-9, with optimal pH being 7.5 [3] . A generation time of 3 h is observed [6] . Similar to many Deinococcus spp., D. deserti are strictly aerobic, protease and catalase positive [6] .
Genome
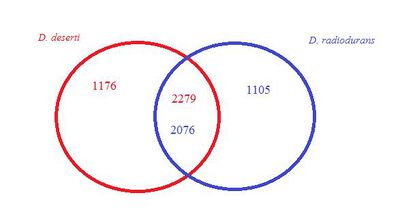
Deinococcus deserti consists of a 2.8 Mb chromosome and three plasmids of 324 kb, 314, and 396 kb [6] . The genome has 3,455 predicted genes and 1348 proteins have been uncovered [6] . The DNA G+C contents of D. deserti VCD115 is 59.8 mol% [4]
Nuclear Organization
The nucleoids of Deinococcus species show a higher degree of compaction compared to radiation sensitive bacteria [10]. A condense nucleoid structure may contribute to radiation resistance by providing a scaffold for efficient DNA repair and limiting dispersion of radiation-induced DNA fragments [10]. Comparative genomics revealed a low diversity of Deinococcus homologs of known histone-like nucleoid associated proteins (NAPs) in E. coli [10]. Only 4 of the 12 NAPs characterized in E. coli (HU, DnaA, Dps, and Lrp) have been identified in D. radiodurans and D. deserti [10]. HU and DNA gyrase being the most abundant proteins involved in the organization of the Deinococcus nucleoids [10]. It is proposed that the condensed nucleoid structure may be consequential absence HU and DNA-gyrase modulators, normally found in radio-sensitive bacteria [10].
Cell Structure
The unusual structure of Deinococcus spp. is their cell envelope [1] . It is unusual in both structure and composition [1] . The cell envelope resembles the cell wall of Gram-negative bacteria, but they have a thick peptidoglycan layer as Gram-positive bacteria [1] . Many members of Deinococcus, including D. radiodurans , stain Gram-positive [1] . Unlike all Deinococcus spp., except D. grandis and D. indicus, D. deserti strain Gram-negative.
Metabolism
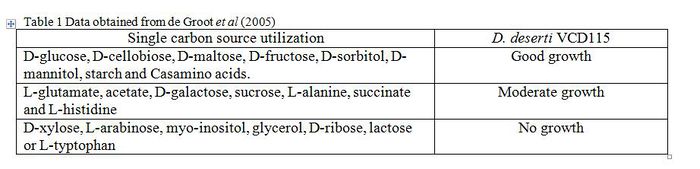
Deinococcus deserti is an obligatory chemoorganoheterotroph. Metabolic capabilities of Deinococcus are similar to those of E. coli, but less diverse [5] . Deinococcus spp. genome encodes the expected functional pathways for carbohydrate metabolism, namely glycolysis, gluconeogenesis, pentose phosphate pathway, and tricarboxylic acid cycle [5] . Amino acids are used as a nitrogen source, especially sulfur-containing amino acids [5] . They possess the standard bacterial lipid metabolism enzymes, but they do lack phosphatidylglycerophosphate synthase. This is enzyme typically involved in the biosynthesis of acidic phospholipids [5] .
Incubation of 9 days in API 20 NE strips, showed that D. deserti was weakly positive for β-glucosidase and β-galactosidase [6] .
Radioactive Tolerance
The adaptation for tolerance to a high quantity of ionizing radiation is much more puzzling, because there is an absence of high radioactive environments on Earth over geologic times [5] . For the last 4 billion years, the radiation levels on Earth’s surface have generally given about 0.05-20 rads/year [5] .
A variety of agents, such as UV light or oxidizing agents, and dynamic environments, such as cycles of desiccation/hydration and extreme temperatures, can also cause damages to DNA which mirrors damage caused by ionizing radiation [5] .
The key technique of this tolerance is the capacity to rebuild a functional genome from radiation-induced DNA fragments, whereas the genomes of majority of other organisms would stay irreversibly fragmented under those same conditions [10]. A combination of different active and passive mechanisms play a role in survival when subjected to DNA-damaging stresses which include efficient DNA repair processes, prevent protein from oxidation, and nucleoid organization [10]
IrrE, a novel protein initially found in D. radiodurans , was shown to be a positive regulator of some of the genes which were up-regulated as a response to desiccation and ionizing radiation [11]. Deinococcus deserti IrrE is able to restore radioresistance in D. radiodurans irrE mutants, indicating that irrE is conserved in Deinococci [11]. The IrrE gene is constitutively expressed [11]. Its inactivation results in a dramatic decrease in resistance to ionizing radiation and UV light [11] This suggest that IrrE is a regulator of genome protection, repair pathways and a signal transduction pathway relative to genome damage [11].
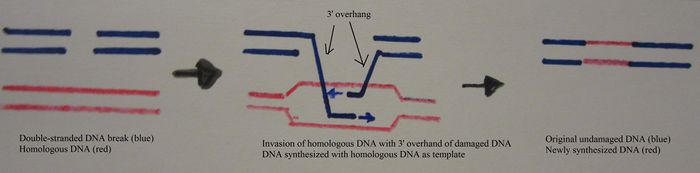
DNA repair processes
The main pathway involved in DNA double-strand break repair is homologous recombination [2] . This technique uses intact homologous DNA molecules to correct the damage sites of DNA. A single-stranded 3’ DNA overhang is produced and loads the RecA protein [2] . The overhang invades a homologous DNA molecule, and new stretches of DNA are synthesized with DNA polymerase I using the homologous DNA as a template [2] . DNA ligase seals the remaining nicks [2] .
Prevent Protein Oxidation
Compared to other radiation sensitive bacteria, D. radiodurans accumulates very high intracellular manganese and low iron levels [8] . Accumulating manganese is shown to prevent protein oxidation and increase radio-resistance [8] . Namely, the proteins involved in DNA repair aren’t damaged thus can function more efficiently in radioresistant bacteria [2, 8]. The Mn/Fe ratio was found to be even higher in D. deserti than D. radiodurans [6] .
Biotechnical Applications
The comparative genomic and proteomics of D. deserti and other members of Deinococcus has revealed several gene sequences and proteins important in surviving in such harsh conditions. In addition, has revealed mechanisms which lead to extreme capacity of radioactive tolerance in Deinococcus. An understanding of DNA damage and repair can be exploited for practical purposes, such as clean up and stabilization of radioactive waste sites [5] .
References
1. Brooks B. and Murray G. “Nomenclature for “Micrococcus radiodurans” and Other Radiation-Resistant Cocci: Deinococcaceae fam. nov. and Deinococcus gen. nov., Including Five Species.” International Journal of Systematic Bacteriology, 1981,0020-7713/81/030353-08$02.00/0
2. Blasius M., Hubscher, U., Sommer, S. “What belongs in the survival kit?” Critical Reviews in Biochemistry and Molecular Biology, 2008, DOI: 10.1080/10409230802122274
3. de Groot A., Chapon, V., Servant, P., Christen, R., Fischer-Le Saux, M., Sommer, S. and Heulin, T. “Deinococcus deserti sp. nov., a gamma-radiation-tolerant bacterium isolated from the Sahara Desert.” International Journal of Systematic and Evolutionary Microbiology, 2005, DOI 10.1099/ijs.0.63717-0
4. Deinococcus deserti genome http://www.ncbi.nlm.nih.gov/genome/?term=Deinococcus+deserti
5. Makarova K.., Aravind, L., Wolf, Y. I., Tatusov, R., Minton, K.., Koonin, E., Daly, M. “Genome of the extremely radiation-resistant bacterium Deinococcus radiodurans viewed from the perspective of comparative genomics.” Microbiology Molecular Biological Review, 2001, DOI: 10.1128/MMBR.65.1.44-79.2001
6. de Groot A., Dulermo R., Ortet P., Blanchard L., Guerin P., Fernandez B., Vacherie B., Dossat C., Jolivet E., Siguier P., Chandler M., Barakat M., Dedieu A., Barbe V., Heulin T., Sommer S., Achouak W. and Armengaud J. “Alliance of proteomics and genomics to unravel the specificities of Sahara bacterium Deinococcus deserti.” PLoS Genet, 2009, DOI: 10.1371/journal.pgen.1000434
7. Tanaka M., Earl, A., Howell, H., Park, M., Eisen, J., Peterson, S., Battista, J. “Analysis of Deinococcus radiodurans’s transcriptional response to ionizing radiation and desiccation reveals novel proteins that contribute to extreme radioresistance.” Genetics, 2004, DOI: 10.1534/genetics.104.029249
8. Daly, M.J., Gaidamakova, E. K., Matrosova, V.Y., Vasilenko, A., Zhai, M., Venkateswaran, A., Hess, M., Omelchenko, M.V., Kostandarithes, H.M., Makarova, K.S., Wackett, L.P., Fredrickson, J.K., Ghosal, D. “Accumulation of Mn(II) in Deinococcus radiodurans Facilitates Gamma-Radiation Resistance” Science, 2004, DOI: 10.1126/science.1103185
9. Daly, M.J., Gaidamakova, E.K., Matrosova, V.Y., Vasilenko, A., Zhai, M., Leapman, R., Lai, B., Ravel, B., Li, S.W., Kemners, K.M., Fredrickson, J.K. “Protein oxidation implicated as the primary determinant of bacterial radioresistance” PLoS Biol, 2007, DOI:10.1371/journal.pbio.0050092
10. Toueille, M., Mirabella, B., Guérin, P., de la Tour, C.B, Boisnard, S., Nguyen, H.H., Blachard, L., Servant, P., de Groot, A., Sommer, S., Armegaud, J. “A comparative proteomic approach to better define Deinococcus nucleoid specificities.” Journal of Proteomics, 2012, DOI:10.1016/j.jprot.2012.03.002
11. Vujičić-Žagar A., Dulermo, R., Gorrec, M., Vannier, F., Servant, P., Sommer, S., de Groot, A., Serre, L. “Crystal Structure of the IrrE Protein, a Central Regulator of DNA Damage Repair in Deinococcaceae” Journal of Molecular Biology, 2009, DOI:10.1016/j.jmb.2008.12.062
12. Images from http://www.genoscope.cns.fr/spip/
Page authored by Michelle Tran, B.Sc. '14 for MICB 301 Microbial Ecophysiology, University of British Columbia, December, 2012.
