Granulosis Virus
Classification
|
NCBI: Taxonomy |
Domain: Viruses
Group: dsDNA
Order: Unassigned
Family Baculoviridae
Genus: Betabaculovirus
Type Species
Cydia pomonella granulovirus
Description and Significance
Granuloviruses are in the family of insect viruses called the baculoviridae. These viruses target insects and are popular as insecticides for farmers targeting lepidopteran larva. The viruses are present in many historical records for causing silkworm 'jaundice' and have been used as biopesticides since WWII. There are currently 17 species in the genus betabaculovirus. These viruses are popular due to their selectivity of only attacking insects from the order lepidotera. Granuloviruses are well known for their unique ability to completely liquify their hosts in order to spread to more hosts, a trait they share with the closely related nuclear polyhedrosis viruses. The type species of granuloviruses is the species Cydia pomonella granulosis virus (CpGV), which only affects Cydia pomonella larva. C. pomonella larva are pests of various fruits in agriculture, causing great loss of ripened fruits. Granulovirus has been used as a pesticide since World War II (Federici, 1997); however, C. pomonella has been developing resistance to granulovirus since 2005 (Sauer, 2017). Granuloviruses are useful in the pesticide industry due to their ability to efficiently kill lepidopteran pests, such as C. pomonella, to protect crops without damaging them or harming the consumer of the crops. Extensive research is being done on more uses and applications of baculoviridae to operate as pesticides against more specialized or resistant lepidopteran pests.
Genome Structure
CpGV is a double stranded DNA virus with a circular genome. The genome is 123,500 bases with 143 open reading frames. 25 of the genes are unique to CpGV, while 118 are homologous to other Baculovirus species. CpGV encodes six genes required for genome replication, which includes DNA polymerase and helicase. It has genes for granulin/polyhedrin, which is a major protein involved in the formation of viral inclusion bodies. Auxiliary genes include proteases, such as chitinase, which are dedicated to the degradation of host structures and the prevention of host cell death (Luque et al, 2001).
Ecology and Pathogenesis
Granulosis was discovered in the early 1960’s, and was found to only infect the codling moth (C. pomonella) and species closely related to it. The codling moth bores into apples, which makes them unfit for human consumption. Spraying apples with granulosis virus significantly decreased the amount of damaged apples, and killed most codling moth larvae before they could enter the fruit; most died after feeding on the epidermis of the treated fruits, which are still suitable for human consumption (Falcon et al, 1968). Granuloviruses are produced for commercial use on crops by infecting large numbers of lepidopteran larva in a lab setting (D'Amico, Podgwaite, 2015). Upon liquification, the larva and virus products are brought to a processing facility where the liquid is converted into a powder mix (D'Amico, Podgwaite, 2015). This powder is then mixed with water and sprayed on the crops to protect the crops from lepidopteran pests (D'Amico, Podgwaite, 2015). Larva that are killed by the granuloviruses become shiny in appearance and latch on to the leaves of the plant they are on (D'Amico, 2015). The bodies of the larva are generally very flimsy and rupture after proteases break through epithelial layer of the host, causing a release of viral particles into the environment (D'Amico, 2015).
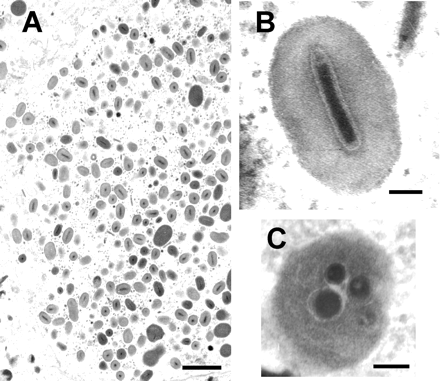
Granuloviruses occlusion bodies contain one or rarely two virions and are about 0.16-0.30 μm by 0.30-0.50 μm in size. The nucleocapsid of the virus contains a double-stranded circular-shaped strand of DNA (Fields Virology, 2013). The outside of the nucleocapsid contains proteins that form ring-shaped subunits through their interactions. The end of the virion contains many unique proteins that are not found anywhere else in the virion, including the protein pp78/83 which aids in assembly of actin and production of offspring viruses (Shuler, Michael L. et al, 1995). Occlusion bodies (Occluded virus, or OV) of granuloviruses contain one or two virions that are wrapped in a protein called granulin (a protein that distinguishes granuloviruses from nuclear polyhedrosis viruses) (Fields Virology, 2013). Occlusion bodies are crystalline gene products that are responsible for the primary viral infection in the gut basal cells. Their function is to release nucleocapsids into gut epithelial cells during infection (Rohrmann, 2013). These occlusion bodies are oval shaped and referred to as 'granules' due to their appearance under a microscope. (Fields Virology, 2013). Granuloviruses also encode homologs of LD130, an envelope fusion protein (Fields Virology, 2013). Granulovirus replication is biphasic cycle, where the budded viruses are formed prior to the occlusion viruses. Budded viruses are produced after primary infection and are simply nucleocapsids surrounded by a membrane (Rohrmann, 2013). Their function is for cell-to-cell transmission of granulovirus.
The granulovirus replication cycle begins when the occluded form of the virus is consumed by hosts off leaves, where it is dissolved in the alkali environment of the C. pomonella larva midgut (Summers, 1971). The cells endocytose the virus released from the occlusion body, which is carried by actin filaments to the nucleus. At this point, the virus uncoats and releases its genome into the nuclear pores. The genome is then incorporated into the host genome and is transcribed by host machinery. Budded viruses are formed using host machinery to infect more cells in the organism, as occluded viruses cannot be transmitted between cells due to presence of granulin, which can not enter cells (Fields Virology, 2013). The assembled budded virus buds off the basal side of gut lumen cells, where it moves to infect new cells. Each budded virus particle buds out from the cell with the LD130 envelope fusion protein, which allows the budded virus to anchor onto the next gut cell. The cell then endocytoses the nucleocapsid of the budded virus and the replication cycle repeats. Granuloviruses encode proteases, such as chitinase, that degrade host proteins and liquify the host, which is the cause of its death (Luque et al, 2001). During these late stages of replication before the larva die, the granulosis viruses begin to express a gene called 'egt' in gut cells (Zimmer, 2014). This gene causes the larva to become agitated and move up the fruit/leaves they are on (Zimmer, 2014). This forced behavior ensures that the viral particles spread further upon liquification of the host (Zimmer, 2014). Scientists are interested in this mechanism of behavior because it is uniquely controlled by a singular gene (Zimmer, 2014). During very late stages of replication, as the larva begin to move to the top of the plant, nucleocapsids remaining in the host cell nuclei are used to form new occluded viruses to be spread upon host liquification (Fields Virology, 2013). The occlusion viruses are produced due to their enhanced survivability in the environment (Fields Virology, 2013). This is because the occlusion body is efficient at protecting viral particles from heat and ultraviolet light protein denaturation while the budded virus membrane does not protect the virus from environmental damage (Fields Virology, 2013).
References
S. Au, et al. Baculovirus Nuclear Import: Open, Nuclear Pore Complex (NPC). Viruses. 23 July 2013.
Federici B.A. (1997) Baculovirus Pathogenesis. In: Miller L.K. (eds) The Baculoviruses. The Viruses.
[www.ncbi.nlm.nih.gov/books/NBK138305/ Rohrmann, George F. “The Baculovirus Replication Cycle: Effects on Cells and Insects.” Baculovirus Molecular Biology [Internet]. 3rd Edition., U.S. National Library of Medicine, 12 Dec. 2013.]
[D'Amico, Vince, and John Podgwaite. “Gypchek - The Gypsy Moth NPV Product - Natural Enemies - Gypsy Moth - Forest Disturbance Processes - Northern Research Station - USDA Forest Service.” USDA Forest Service, USDA, 2015, www.nrs.fs.fed.us/disturbance/invasive_species/gm/control_management/gypchek_production/.]
[D'Amico, Vince. “Baculoviruses.” Biological Control, Cornell University College of Agricultural and Life Sciences, 2015, biocontrol.entomology.cornell.edu/pathogens/baculoviruses.php.]
[Fields, Bernard N., and David Mahan Knipe. Fields Virology. 6th ed., vol. 2th, Raven Press, 2013.]
[Shuler, Michael L., et al. Baculovirus Expression Systems and Biopesticides. Wiley-Liss, 1995.]
[Zimmer, Carl. “Mindsuckers: Meet Nature's Nightmare, Carl Zimmer.” National Geographic, National Geographic, Nov. 2014, carlzimmer.com/mindsuckers-meet-natures-nightmare-128/.]
Author
Page authored by Ben Kelly and Ilise Kundel, student of Prof. Jay Lennon at IndianaUniversity.