Group B Strep and Pregnancy
Introduction
By Shawn Ruiz
Group B Strep (GBS), also known as Streptococcus agalactiae, is a Gram-positive, beta-hemolytic, catalase-negative, facultative anaerobe that is a normal component of the gastrointestinal and genitourinary tracts[1]. In fact, GBS colonizes the gastrointestinal and genitourinary tracts of up to 50% of healthy adults[2]. Most healthy adults who are colonized by GBS will not experience any symptoms or GBS-related infections. While the bacteria is usually harmless in healthy adults, it is a major cause of meningitis, pneumonia, and sepsis in neonates[3]. GBS is the leading infectious cause of neonatal mortality and morbidity in the United States; between four and six percent of babies who develop GBS disease die[4][5]. GBS causes both early onset (<7 days old) and late onset (7-90 days old) infections in neonates[4]. The main risk factor for an early-onset GBS infection in neonates is colonization of a birthing person's genital tract with Group B strep during labor[4]. About one in four pregnant individuals carry GBS in their body[5]. If the bacteria is present in a pregnant person, it can be directly transferred to their neonate(s) in a multitude of ways. For example, GBS can travel from the vagina into the amniotic fluid where the neonate(s) can ingest it. The neonate(s) can also come into contact with the bacteria as they make their way down the birth canal[3]. In the early 1990s, the early GBS infection rate was 1.7 cases per 1,000 births[3]. In an effort to decrease this infection rate, the American Congress of Obstetricians and Gynecologists and the American Academy of Pediatrics recommended screening pregnant individuals for GBS before they go into labor[3]. It is now common practice to screen pregnant individuals for GBS at some point between 35 and 37 weeks of pregnancy[5]. Pregnant people who test positive for GBS are treated with intravenous antibiotics during labor[5]. Penicillin and ampicillin are the recommended antibiotics for intrapartum GBS prophylaxis[6]. If a pregnant person tests positive for GBS and they are treated with antibiotics during labor, the risk of their neonate(s) developing a serious, life-threatening GBS infection drops by 80% [3]. Early GBS infection rates in the United States have significantly dropped (0.25 cases per 1,000 births) since these preventative measures went into effect around 1995[3]. Intrapartum prophylaxis effectively prevents GBS transmission from the birthing individual to their neonate(s) during labor and delivery. This preventative measure, however, does not target in utero infections that occur earlier in pregnancy, and little is known about the mechanisms that result in the infection of the amniotic cavity[7]. In utero GBS infections have devastating effects, including preterm birth and mortality in both the pregnant person and their neonate(s)[7]. That said, it is critical that researchers and public health officials work toward understanding exactly how GBS infects the amniotic cavity.
The Hemolytic Pigment of GBS
GBS has been isolated from the amniotic fluid of birthing people with intact chorioamniotic membranes, suggesting that GBS is capable of invading and breaching amniotic epithelium and chorioamnion[7][8][9][10][11]. Whidbey et al. hypothesized that “intra-amniotic GBS infections in patients with intact placenta or chorioamniotic membranes may be due to elevated virulence factor expression”[7][8][9][10][11]. Previous studies showed that the expression of GBS virulence genes is regulated by a two-component regulatory system: COVR/S[7][12][13][14]. More specifically, COVR/S was described to repress a multitude of GBS virulence genes, including the cyl operon, which contains the cylE gene that is important for the production of the pluripotent toxin, hemolysin [7][12][13][14]. In order to test whether or not increased expression of GBS virulence genes promotes the invasion of amniotic epithelium, Whidbey et al. compared the abilities of wild-type (WT) GBS and hyper-hemolytic GBS (ΔcovR) to adhere to and invade human amniotic epithelial cells (hAECs) that were isolated and cultured from “normal, term placentas obtained immediately after cesarean delivery from” birthing people without labor[7]. Non-hemolytic GBS strains (ΔcovRΔcylE and ΔcylE) were also included in the experiment to investigate hemolysin’s role in GBS strains’ ability to adhere to and invade hAECs[7][15]. The results, shown in Figures 2 and 3, show that all GBS strains adhered to hAECs. WT GBS invaded hAECs (~4% invasion), non-hemolytic GBS showed a significantly decreased invasion when compared to the WT (~0.3% invasion), and the hyper-hemolytic GBS strain (ΔcovR) was significantly more invasive when compared to the WT strain (~80% invasion)[7]. In sum, Whidbey et al. concluded that hemolysin promotes GBS invasion of hAECs.
Next, Whidbey et al. investigated if the increased expression of hemolysin in the hyper-hemolytic GBS strain (ΔcovR) activates an inflammatory response[7]. Whidbey et al. isolated RNA from GBS-infected hAECs four hours after infection, and they used qRT-PCR to examine any changes in the expression of inflammatory genes in hAECs[7][16]. The results, depicted in Figure 4, suggest that infection with ΔcovR caused a significant increase in the transcription of cytokines (e.g., IL-6 and IL-8) in hAECs[7]. In sum, the results suggest that hAECs that were infected with the hyper-hemolytic GBS strain (ΔcovR) did show an inflammatory response.
It is known that microbial toxins use the nuclear transcription factor, NF-𝜅B, which is recruited from the cytoplasm to the nucleus, to activate inflammatory signaling pathways[17]. Given that hAECs that were infected with ΔcovR showed an inflammatory response, Whidbey et al. investigated if the increase in the expression of pro-inflammatory genes was associated with the nuclear recruitment/localization of NF-𝜅B[7]. Whidbey et al. isolated total nuclear and cytoplasmic proteins from infected and uninfected hAECs, and they resolved them on 10% SDS-PAGE and Western blots[7]. As shown in Figure 5, hAECs that were infected with the hyper-hemolytic GBS strain (ΔcovR) showed a 2.5 fold increase in the nuclear recruitment of NF-𝜅B compared with WT GBS-infected or uninfected controls. These results suggest that the increase in the expression of inflammatory genes seen in ΔcovR-infected hAECs "is associated with the nuclear recruitment of NF-𝜅B"[7].
Despite finding that hemolysin is associated with an increase in the expression of pro-inflammatory genes, as well as the nuclear recruitment of NF-𝜅B, Whidbey et al. still did not know if the presence of hemolysin accelerates the failure of the human amniotic epithelial barrier[7]. To examine this, Whidbey et al. monitored the changes in transepithelial electrical resistance across hAEC using electric cell-substrate impedance sensing[7][18]. Figure 6 shows that hAECs that were infected with the hyper-hemolytic strain (ΔcovR) showed an accelerated decrease in their barrier resistance compared to hAECs that were infected with WT GBS[7]. In sum, increased hemolysin expression decreased the barrier function of the amniotic epithelium, enabling GBS to breach the epithelial barrier[7].
Knowing that an increase in hemolysin expression enabled hyper-hemolytic GBS to breach the hAEC epithelium, Whidbey et al. investigated if GBS are capable of penetrating intact chorioamniotic membranes[7]. Whidbey et al. mounted, maintained, and, after 48 hours of stabilization, infected chorioamniotic membranes with various GBS strains (WT, ΔcovR, ΔcovRΔcylE). Figure 7 shows that only ΔcovR penetrated and invaded the chorioamnion, including the amniotic epithelium. This suggests that an increase in hemolysin expression can “facilitate bacterial penetration of chorioamniotic membranes and the amniotic cavity”[7].
Considering that hyper-hemolytic GBS is capable of penetrating the chorioamniotic membranes and the amniotic cavity, which may result in an in-utero infection and/or preterm labor, Whidbey et al. hypothesized that increased hemolytic activity could be observed in GBS isolated from birthing people in preterm labor[7]. To test this hypothesis, they collected clinical isolates from amniotic fluid and chorioamnion from six birthing people in preterm labor[7]. These isolates were examined for potential mutations in the covR/S locus, as well as hemolytic properties. Eight GBS strains were obtained: most of them demonstrated increased hemolytic activity, and six of them had a mutation in the covR/S loci[7]. Notably, two GBS isolates did not have a mutation in the covR/S loci, but they did demonstrate increased hemolytic activity. That said, it is likely that hemolytic activity is regulated by multiple regulatory mechanisms. Site-directed mutations of the COVR/S mutations that were collected from clinical samples confirmed that the increase in hemolytic activity was in part attributable to these mutations[7]. Thus, hyper-hemolytic GBS can be associated with birthing people in preterm labor.
One study suggested that the cylE gene, which is a part of the cyl operon, encoded and produced hemolysin[15]. Whidbey et al. proved, however, that, while CylE is necessary for the production of hemolysin, it is not sufficient for GBS hemolysis. Notably, studies have shown that the hemolytic phenotype of GBS is pigmented and the non-hemolytic phenotype is not; this pigment is described as an ornithine rhamnolipid (i.e., granadaene)[15][19][20]. Like hemolytic activity, pigment biosynthesis in GBS also requires the cyl open. Whidbey et al. conducted a sequence profile analysis of CylE, which they used in combination with predicted homology profiles, to propose a pathway for GBS pigment biosynthesis that is dependent on the majority of genes in the cyl operon, which is depicted in Figure 8[7].
That said, Whidbey et al. propose that the ornithine rhamnolipid pigment, rather than the CylE protein, is the cause of the hemolytic activity in GBS. To test this, they extracted and purified pigment from WT GBS and examined the hemolytic activity of the pigment. Lysis of red blood cells occurred in the presence of the pigment, and brief (8 min) exposure to the pigment dramatically altered the membrane morphology of the disc-shaped red blood cells, as shown in Figure 9. Researchers confirmed that the hemolytic activity observed in the lysis of red blood cells could not be attributed to a protein toxin by demonstrating that the purified pigment was not affected by proteinase K. Other experiments, such as SDS-PAGE analysis of the purified pigment, also helped confirm this finding[7].
The Lytic Mechanisms and Cytolytic Abilities of the GBS Hemolytic Pigment
Whidbey et al. investigated how the GBS pigment induces cell lysis by measuring the kinetics of both K+ and hemoglobin (Hb) release from red blood cells treated with 400nM of the pigment[21]. These results suggest that membrane permeabilization enables K+ and Hb to efflux, respectively. This delay in Hb suggests that the GBS pigment induces a colloidal osmotic mechanism of lysis rather than direct lysis[21]. Protection assays revealed that small osmoprotectants (e.g., PEG200) are incapable of protecting RBC from pigment-mediated hemolysis, whereas larger osmoprotectants (e.g, PEG1500) completely protect RBC from pigment-mediated hemolysis[21]. Interestingly, pigment-mediate membrane permeability is independent of cellular response, and it does not conform to a pore-forming protein toxin nor the induction of instant lysis[21].
To investigate the cytolytic abilities of GBS, researchers treated macrophages with various GBS strains (WT, ΔcovR, ΔcylE, and ΔcovRΔcylE) for four hours; cytotoxicity was measured by the release of lactate dehydrogenase[21]. Macrophage death and IL-1β and IL-18 release in cells that were infected with hyper-hemolytic GBS was significantly higher compared to those infected with non-hemolytic strains[21]. This, combined with further experiments that confirmed these findings and revealed an increase in IL-1β and IL-18 levels in macrophages treated with hyper-hemolytic GBS strains, suggest that the purified hemolytic GBS pigments is pro-inflammatory and cytotoxic[21].
The increase in the secretion of IL-1β and IL-18 in macrophages treated with the GBS pigment suggests that the pigment is capable of triggering activation of the inflammasome, which is a cytosolic complex that is responsible for activating inflammatory responses to injury and/or illness[21]. Whidbey et al. write, "one major inflammasome comprises the NLR (nucleotide binding, leucine-rich repeat containing) protein known as NLRP3, which associates with the adaptor ASC (apoptosis-associated speck-like protein containing the caspase recruitment domain, CARD)"[21][22]. Whideby et al. conducted several experiments to investigate if the GBS pigment activates NLRP3, and they concluded that GBS induced hemolysin-dependent cell death in macrophages that contain the NLRP3 inflammasome[21].
The GBS Hemolytic Pigment is the Critical Component of GBS-Related Fetal Injuries
To determine if hyper-hemolytic GBS strains induce fetal injury or preterm birth, Whidbey et al. used an intrauterine model of inoculation to introduce GBS strains (WT, hyperpigmented (ΔcovR), or non-hemolytic (ΔcovRΔcylE)) into the right horn of the uteruses of pregnant mice[21][24][25][26]. The inoculated mice were monitored for 72 hours for signs of preterm, which Whidbey et al. defined as the birth of at least one pup in a mouse’s cage[21]. One of six mice infected with WT GBS and three of six mice infected with ΔcovR GBS experienced preterm birth compared with the controls groups[21]. Moreover, Figure 10 shows that there were significantly higher fetal deaths in mice that were infected with WT GBS and ΔcovR GBS (hemolytic strains)[21]. Whidbey et al. reported that there was not a statistically significant difference between the frequency of fetal deaths between the two hemolytic strains, and they explained that this may be due to a decrease in the repression of hemolysin/pigment by COVR/S in vivo[21][27][28].
Whidbey et al. also investigated if the activation of NLRP3 inflammasome is important for fetal injuries that are the caused by hyper-hemolytic GBS strains[21]. Notably, Whidbey et al. found that the production of the hemolytic pigment is the critical component of GBS fetal injury, as it contributes to fetal injuries that are associated with GBS infections in ways that are independent and dependent on the NLRP3 inflammasome[21].
GBS Exploits Vaginal Epithelial Exfoliation for Ascending Infection
In order for in utero infections to occur in humans, vaginal microbes must enter the uterus by ascending infection. The mechanisms behind ascending infection are not clear, but Vornhagen et al. propose a convincing mechanism, which is depicted in Figure 11, that GBS may use to cause in utero infections.[23]. Notably, not all pregnant people experience ascending microbial infections. Vornhagen et al. propose that one reason for this may be a breakdown in the host’s defenses that work to prevent ascending infection[23]. Epithelial cells, for example, create a physical barrier that is able to recognize a pathogenic attack and respond to it appropriately[23]. One of these defense mechanisms is called epithelial exfoliation (i.e., shedding), during which epithelial cells detach from their basement membrane, which prevents pathogens from harming the host[23]. Vaginal epithelial cells use exfoliation to protect their host[23]. Interestingly, vaginal GBS colonization increases a pregnant person’s risks of experiencing an ascending infection and, in turn, adverse pregnancy outcomes[23]. Given this, Vornhagen et al. hypothesized that vaginal epithelial exfoliation may regulate GBS colonization[23].
To test if GBS may trigger the loss of barrier function in and cellular detachment of vaginal epithelial cells, which will eventually result in exfoliation, Vornhagen et al. infected immortalized human vaginal epithelial cells (hVECs) with the GBS WT strain COH1[23]. Figure 12 shows that Vornhagen et al. noticed (A) a significant amount of cellular detachment in hVECs, (B) a significant difference in hVEC structure and shape, and (C) a loss of barrier function in hVECs infected with WT GBS[23]. In other words, these results suggest that WT GBS infection of hVECs stimulates vaginal epithelial cell exfoliation[23].
Next, Vornhagen et al. measured “the ability of GBS to migrate across hVEC monolayers in [a] Transwell assay[23]." Vornhagen et al. observed that a significant amount of GBS crossed the epithelial barrier. Notably, these crossing times were associated with vaginal epithelial cell exfoliation time points, suggesting that GBS induces vaginal epithelial cell exfoliation, allowing for an increase in GBS cell migration across vaginal epithelia[23].
Vornhagen et al. vaginally inoculated female mice with WT GBS to see if GBS infection results in epithelial exfoliation[23]. Interestingly, blinded quantification scoring of scanning electron microscopy images revealed that mice that were infected with GBS experienced significantly higher levels of vaginal exfoliation than the control group 72 and 96 hours post-inoculation[23]. Vornhagen et al. probed the vaginal epithelial with fluorescent nanoparticles in vivo to confirm that GBS-induced epithelial exfoliation also resulted in a loss of epithelial barrier function in vivo[23]. Notably, over time, more nanoparticles penetrated the epithelial barrier in GBS-inoculated mice than in the control sample, suggesting that in vivo GBS infections resulted in epithelial exfoliation, as well as a disruption in epithelial barrier function, which is associated with increased bacterial dissemination[23].
It is known that the loss of barrier function and increased mobility of epithelial cells is associated with EMT, which is the process whereby nonmotile epithelial cells become mobile metastatic cells; this process is often characterized by a loss of the epithelial marker E-cadherin and a gain in the expression of the mesenchymal marker N-cadherin[23][29]. Although, there is evidence of EMT occurring in cells when only the expression of E-cadherin was decreased without the increase in the expression of N-cadherin[30]. Vornhagen et al. infected hVECs with WT GBS, and they used flow cytometry to detect whether or not infected cells showed evidence of the EMT mesenchymal change. WT GBS-infected hVECs showed a decrease in the epithelial marker E-cadherin, and immunohistochemistry of in vivo murine samples of GBS-infected hVECS validated this finding[23]. While a decrease of the epithelial marker E-cadherin was observed in murine vaginal samples, Vornhagen et al. did not observe a decrease in the expression of the mesenchymal marker N-cadherin in murine vaginal tissue, which may be the result of the difference in the expression levels of N-cadherin in different types of cells[23]. That said, Vornhagen et al. concluded that GBS are capable of inducing EMT in vaginal epithelial cells, which may be the cause of epithelial exfoliation in vaginal epithelial cells[23].
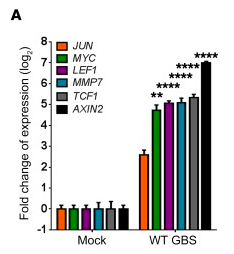
It is known that the β-catenin signaling pathway can regulate EMT[29]. To determine if the GBS can trigger the β-catenin signaling pathway to regulate EMT, Vornhagen et al. examined the gene expression of β-catenin regulated genes in GBS-infected hVECs[23]. Interestingly, Figure 13 shows that β-catenin regulated genes were significantly upregulated in WT GBS-infected hVECs and vaginal tissues from GBS inoculated mice, suggesting that GBS may trigger the β-catenin signaling pathway. This finding, combined with the findings of several other molecular experiments, lead Vornhagen et al. to conclude that the data suggest that β-catenin signaling controls EMT in vaginal epithelial cells and that GBS may use this signaling mechanism to induce epithelial exfoliation[23].
While interesting, these findings did not enable Vornhagen et al. to identify the extracellular signal that GBS may use to prompt β-catenin signaling[23]. Vornhagen et al. eventually turned their attention to the integrin protein family, because they are known for their ability to control various cellular processes [31]. The molecular mechanisms of the integrin signaling pathway are described in detail in the Vornhagen et al. study[23]. Vornhagen et al.’s knowledge of the molecular mechanisms of the integrin signaling pathway encouraged them to conduct several molecular experiments that lead to the suggestion that GBS are able to stimulate integrin signaling, which permits the nuclear translocation of β-catenin[23]. This translocation potential leads to EMT, barrier disruption, and ultimately vaginal epithelial exfoliation[23]. This mechanism is depicted in Figure 11. The data suggest that, while vaginal epithelial cell exfoliation serves as a defense mechanism against certain pathogens, GBS actually use this defense mechanism as a way to ascend further into the vagina[23].
Conclusion
GBS infections are associated with adverse pregnancy outcomes, such as stillbirth. The molecular mechanisms of these outcomes are still unknown, but the two Whidbey et al. studies and the Vornhagen study discussed above significantly advanced the efforts to combat these pregnancy outcomes by expanding our knowledge about the molecular mechanisms of GBS infections and their negative effects on pregnant and birthing individuals and their neonate(s). These studies show that GBS hemolysin is not a protein, as it was previously described, but rather a toxic lipid pigment. Increased expression of this pigment has several consequences, including but not limited to bacterial (GBS) penetration of chorioamniotic membranes and the amniotic cavity, activation of the NLRP3 inflammasome, and higher rates of murine fetal death. The proposed mechanism of hemolysin biosynthesis and the proposed mechanism that explains how GBS exploits vaginal epithelial exfoliation for ascending infection (which were proposed by Whidbey et al. and Vornhagen et al., respectively) are promising starting points for future research that aims to discover and implement preventative measures against in utero GBS infections, which is important because the current intrapartum prophylaxis to prevent GBS transmission from the birthing individual to their neonate(s) does not target in utero GBS infections.
References
Authored for BIOL 238 Microbiology, taught by Joan Slonczewski, 2021, Kenyon College.
- ↑ “Streptococcus Agalactiae.” Wikipedia, Wikimedia Foundation, 24 Mar. 2021, en.wikipedia.org/wiki/Streptococcus_agalactiae..
- ↑ Atul Kumar, et al. “Group B Streptococcus: Global Incidence and Vaccine Development.” Nature Reviews Microbiology, vol. 4, no. 12, 2006, pp. 932–942., doi:10.1038/nrmicro1552..
- ↑ 3.0 3.1 3.2 3.3 3.4 3.5 Dekker, Rebecca. “The Evidence on: Group B Strep.” Evidence Based Birth , Evidence Based Birth , 17 July 2017, evidencebasedbirth.com/groupbstrep/..
- ↑ 4.0 4.1 4.2 John A. “Group B Streptococcus And Pregnancy.” StatPearls, U.S. National Library of Medicine, 29 Jan. 2021, www.ncbi.nlm.nih.gov/books/
- ↑ 5.0 5.1 5.2 5.3 “Group B Strep: Fast Facts and Statistics.” Centers for Disease Control and Prevention, Centers for Disease Control and Prevention, 11 June 2020, www.cdc.gov/groupbstrep/about/fast-facts.html#references.
- ↑ Horsley, Elizabeth. “CDC Updates Guidelines for the Prevention of Perinatal GBS Disease.” American Family Physician, The American Academy of Family Physicians Foundation , 1 May 2011, www.aafp.org
- ↑ 7.00 7.01 7.02 7.03 7.04 7.05 7.06 7.07 7.08 7.09 7.10 7.11 7.12 7.13 7.14 7.15 7.16 7.17 7.18 7.19 7.20 7.21 7.22 7.23 7.24 7.25 7.26 7.27 7.28 7.29 7.30 7.31 7.32 7.33 Whidbey, Christopher, et al. “A Hemolytic Pigment of Group B Streptococcus Allows Bacterial Penetration of Human Placenta.” Journal of Experimental Medicine, vol. 210, no. 6, 2013, pp. 1265–1281., doi:10.1084/jem.20122753.
- ↑ 8.0 8.1 Bobitt, JR, and WJ Ledger. “Unrecognized Amnionitis and Prematurity: a Preliminary Report.” The Journal of Reproductive Medicine., vol. 19, no. 1, 30 June 1977, pp. 8–12., europepmc.org/article/med/874942.
- ↑ 9.0 9.1 Naeye , RL, and EC Peters. “Amniotic Fluid Infections with Intact Membranes Leading to Perinatal Death: a Prospective Study.” Pediatrics, U.S. National Library of Medicine, Feb. 1978, pubmed.ncbi.nlm.nih.gov/634667/.
- ↑ 10.0 10.1 Winram, Scott B., et al. “Characterization of Group B Streptococcal Invasion of Human Chorion and Amnion Epithelial Cells In Vitro.” Infection and Immunity, vol. 66, no. 10, 1998, pp. 4932–4941., doi:10.1128/iai.66.10.4932-4941.1998.
- ↑ 11.0 11.1 Goldenberg, Robert L., et al. “Intrauterine Infection and Preterm Delivery.” New England Journal of Medicine, vol. 342, no. 20, 2000, pp. 1500–1507., doi:10.1056/nejm200005183422007.
- ↑ 12.0 12.1 Lamy, Marie-Cécile, et al. “CovS/CovR of Group B Streptococcus: a Two-Component Global Regulatory System Involved in Virulence.” Molecular Microbiology, vol. 54, no. 5, 2004, pp. 1250–1268., doi:10.1111/j.1365-2958.2004.04365.x.
- ↑ 13.0 13.1 Jiang, Sheng-Mei, et al. “Variation in the Group B Streptococcus CsrRS Regulon and Effects on Pathogenicity.” Journal of Bacteriology, vol. 190, no. 6, 2008, pp. 1956–1965., doi:10.1128/jb.01677-07.
- ↑ 14.0 14.1 Rajagopal, Lakshmi, et al. “Regulation of Cytotoxin Expression by Converging Eukaryotic-Type and Two-Component Signalling Mechanisms in Streptococcus Agalactiae.” Molecular Microbiology, vol. 62, no. 4, 2006, pp. 941–957., doi:10.1111/j.1365-2958.2006.05431.x.
- ↑ 15.0 15.1 15.2 Pritzlaff, Craig A., et al. “Genetic Basis for the Beta-Haemolytic/Cytolytic Activity of Group B Streptococcus.” Molecular Microbiology, vol. 39, no. 2, 2001, pp. 236–248., doi:10.1046/j.1365-2958.2001.02211.x.
- ↑ Lembo, Annalisa, et al. “Regulation of CovR Expression in Group B Streptococcus Impacts Blood-Brain Barrier Penetration.” Molecular Microbiology, vol. 77, no. 2, 2010, pp. 431–443., doi:10.1111/j.1365-2958.2010.07215.x.
- ↑ Gonzalez, M. R., et al. “Bacterial Pore-Forming Toxins: The (w)Hole Story?” Cellular and Molecular Life Sciences, vol. 65, no. 3, 2007, pp. 493–507., doi:10.1007/s00018-007-7434-y.
- ↑ Giaever, I., and C. R. Keese. “Micromotion of Mammalian Cells Measured Electrically.” Proceedings of the National Academy of Sciences, vol. 88, no. 17, 1991, pp. 7896–7900., doi:10.1073/pnas.88.17.7896.
- ↑ Nizet, V, et al. “Group B Streptococcal Beta-Hemolysin Expression Is Associated with Injury of Lung Epithelial Cells.” Infection and Immunity, vol. 64, no. 9, 1996, pp. 3818–3826., doi:10.1128/iai.64.9.3818-3826.1996.
- ↑ Spellerberg, Barbara, et al. “Thecylgenes OfStreptococcus Agalactiaeare Involved in the Production of Pigment.” FEMS Microbiology Letters, vol. 188, no. 2, 2000, pp. 125–128., doi:10.1111/j.1574-6968.2000.tb09182.x.
- ↑ 21.00 21.01 21.02 21.03 21.04 21.05 21.06 21.07 21.08 21.09 21.10 21.11 21.12 21.13 21.14 21.15 21.16 Whidbey, Christopher, et al. “A Streptococcal Lipid Toxin Induces Membrane Permeabilization and Pyroptosis Leading to Fetal Injury.” EMBO Molecular Medicine, vol. 7, no. 4, 2015, pp. 488–505., doi:10.15252/emmm.201404883.
- ↑ Taxman, Debra J., et al. “Inflammasome Inhibition as a Pathogenic Stealth Mechanism.” Cell Host & Microbe, vol. 8, no. 1, 2010, pp. 7–11., doi:10.1016/j.chom.2010.06.005.
- ↑ 23.00 23.01 23.02 23.03 23.04 23.05 23.06 23.07 23.08 23.09 23.10 23.11 23.12 23.13 23.14 23.15 23.16 23.17 23.18 23.19 23.20 23.21 23.22 23.23 23.24 23.25 23.26 23.27 23.28 23.29 Vornhagen, Jay, et al. “Group B Streptococcus Exploits Vaginal Epithelial Exfoliation for Ascending Infection.” Journal of Clinical Investigation, vol. 128, no. 5, 2018, pp. 1985–1999., doi:10.1172/jci97043.
- ↑ Patras, Kathryn A., et al. “Group B Streptococcus CovR Regulation Modulates Host Immune Signalling Pathways to Promote Vaginal Colonization.” Cellular Microbiology, vol. 15, no. 7, 2013, pp. 1154–1167., doi:10.1111/cmi.12105.
- ↑ Hirsch, Emmet, et al. “A Model of Intrauterine Infection and Preterm Delivery in Mice.” American Journal of Obstetrics and Gynecology, vol. 172, no. 5, 1995, pp. 1598–1603., doi:10.1016/0002-9378(95)90503-0.
- ↑ Elovitz, Michal A., et al. “A New Model for Inflammation-Induced Preterm Birth.” The American Journal of Pathology, vol. 163, no. 5, 2003, pp. 2103–2111., doi:10.1016/s0002-9440(10)63567-5.
- ↑ Santi, Isabella, et al. “CsrRS Regulates Group B Streptococcus Virulence Gene Expression in Response to Environmental PH: a New Perspective on Vaccine Development.” Journal of Bacteriology, vol. 191, no. 17, 2009, pp. 5387–5397., doi:10.1128/jb.00370-09.
- ↑ Sitkiewicz, Izabela, et al. “Transcriptome Adaptation of Group B Streptococcus to Growth in Human Amniotic Fluid.” PLoS ONE, vol. 4, no. 7, 2009, doi:10.1371/journal.pone.0006114.
- ↑ 29.0 29.1 Lamouille, Samy, et al. “Molecular Mechanisms of Epithelial–Mesenchymal Transition.” Nature Reviews Molecular Cell Biology, vol. 15, no. 3, 2014, pp. 178–196., doi:10.1038/nrm3758.
- ↑ Costa, Liana Cristina, et al. “Expression of Epithelial-Mesenchymal Transition Markers at the Invasive Front of Oral Squamous Cell Carcinoma.” Journal of Applied Oral Science, vol. 23, no. 2, 2015, pp. 169–178., doi:10.1590/1678-775720140187.
- ↑ Streuli, Charles H. “Integrins as Architects of Cell Behavior.” Molecular Biology of the Cell, vol. 27, no. 19, 2016, pp. 2885–2888., doi:10.1091/mbc.e15-06-0369.