Hepatitis B Virus X gene in Hepatocellular Carcinoma
A Viral Biorealm page on the family Hepatitis B Virus X gene in Hepatocellular Carcinoma
Hepatocellular carcinoma (HCC) is among the most frequently diagnosed cancers worldwide. For over forty years physicians and researchers have observed an association between HBV infection and development of HCC. Of the seven HBV genes, hepatitis b virus X protein (HBx, 17.5 kDa) is the smallest and required extensive efforts to characterize; its elusiveness is highlighted by its designation “X” [1]. The protein functions as a transcriptional transactivator of both viral and cellular transcripts and is capable of inducing cytoplasmic signal transduction pathways [1]. HBx has been shown to interact directly with a wide variety of biomolecules, including ssDNA, multiple transcription factors (TFs), proteins associated with apoptosis, TATA-binding protein, and an essential subunit of human RNA polymerase I, II, and III, RPB5. Interestingly, HBx was a more potent activator of transcription upon deletion of its N-terminal 1 to 50-amino-acid fragment, suggesting this domain may function as a negative regulatory element [1].
Virus Classification
Family: Hepadnaviridae
Genus: Orthohepadnavirus
Baltimore Classification: Class VII - double-stranded DNA viruses that replicate through a single-stranded RNA intermediate
Genome: partially double-stranded DNA, ~3.5 kb
Virus Overview
Hepatitis B virus is transmitted by mucosal or percutaneous contact with body fluids or blood from an infected individual [18]. This occurs most often through injection-drug use, sexual contact with an infected person, or mother-to-child transmission [18]. The infection localizes in the liver and has been linked to the development of several diseases including cirrhosis of the liver and hepatocellular carcinoma. The hepatitis B vaccine contains hepatitis B surface antigen (HBsAg) and is administered in a course of three injections. The vaccine induces serum levels of anti-HBsAg sufficient enough to activate the recruitment cytotoxic T cell and NK cell responses to areas of viral infection and wipe out the pathogen.
Hepatitis B Virus Genome
The hepadnaviral genome is packaged as relaxed circular (RC), partially double-stranded DNA about 3.5 kb in length. The (+) strand exists bound to DNA-binding proteins which inhibit its complete synthesis. In the nucleus, these proteins dissociate and viral polymerase extends the (+) strand, leaving the viral genome as covalently closed circular DNA (cccDNA). The virus benefits from a cccDNA genome because the DNA strand stores energy in supercoils and, as a result, is less reliant on host ATP-dependent helicase activity for DNA replication.
The HBV genome consists of four genes with four partially-overlapping ORFs: core (C), polymerase (P), surface antigen (S), and X. The core gene consists of the precore and core regions [17]. The presence of several in-frame translation initiation codons for S and C genes allows for the translation of related but functionally distinct proteins [17]. The C ORF encodes the viral nucleocapsid (HBcAg) at the core ORF and hepatitis B e antigen (HBeAg) at the precore ORF. The S ORF codes for the surface antigen, HBsAg [17].
Polymerase (pol) is a large protein and consists of three subunits: a reverse transcriptase, a terminal protein subunit which provides the active site for DNA synthesis, and the ribonuclease H which helps replication and breaks down pregenomic RNA [17]. Strand-specific synthesis during replication is controlled by two genomic elements, direct repeats 1 and 2 (DR1 and DR2) in the 5’ ends of the plus strand. Liver-specific expression of viral gene products is controlled by two viral enhancer elements, En1 and En2 [17].
Replication Cycle
To enter a cell, hepatitis B virus binds an unidentified receptor and is endocytosed. Utilizing duck HBV as a model infection system, several blotting techniques, and immunofluorescence microscopy, Breiner et al. (1998) showed that carboxypeptidase D (gp180), a cell surface receptor protein involved in intracellular trafficking to the Golgi, plays a role in viral entry via association with hepatitis B pre-S polypeptide [21].
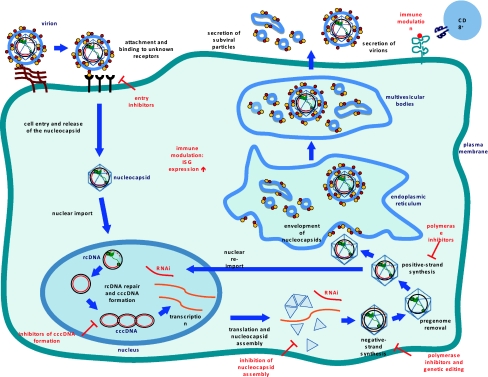
After uncoating of the nucleocapsid core, the viral genome enters the nucleus where the single-stranded gap region is repaired by viral polymerase, leaving the viral genome as cccDNA, the functional template for transcription by host RNA polymerase II. The fact that HBV mRNAs are subject to alternative splicing presents a challenge in the development of small molecule inhibitors targeting specific viral binding domains [19].
Viral DNA is transcribed and the resulting mRNA may be translated into a protein or saved as pre-genomic information. Pre-genomic mRNA (pgRNA) serves as the template for reverse transcription and translation of pol and core; pre-core mRNA levels determine expression of the pre-core protein product. As viral proteins are synthesized, the core packages pgRNA in response to a cis-acting stem loop denoted epsilon. After encapsidation, pgRNA is reverse transcribed by pol to partially double stranded relaxed circular (RC) DNA. The immature virion then interacts with proteins along the endoplasmic reticulum where it matures prior to lysing the cell membrane, triggering its release into the extracellular environment.
Hepatitis B Virus Protein X Interacts with Host and Viral Components
There have been numerous studies investigating the mechanisms of HBx-mediated cell proliferation and HCC development. Two studies in the 1990s demonstrated that HBx potentiates c-myc induced liver oncogenesis in a mouse model [3, 4]. HBx mediates the expression of a wide variety of host RNAs and genes, including transcription factors, the p53 tumor suppressor gene, oncogenes, ribosomal protein S27a [6], ribosomal RNAs [7], miRNAs, and proteins responsible for cell cycle regulation. In HCC cells, overexpression of HBx and the oncogene c-myc each increased rRNA levels; when cells were co-transfected with HBx and c-myc, an additive effect on both rRNA synthesis and cell division was observed [7]. Furthermore, in vitro analyses of HCC cells infected with HBx-knockdown HBV demonstrated the existence of particular HBx-dependent proliferative signaling events in this cancer [5].
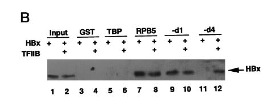
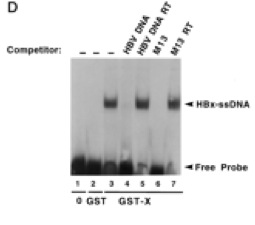
In 1996, it was confirmed by electromobility shift assays (EMSA) that HBx binds single-stranded DNA in a manner that is non-sequence-specific (Fig 4) [24]. A year later, researchers used GST-resin pull-down assays, far-Western blotting, and in vivo co-immunoprecipitation to show that HBx is capable of forming a trimeric complex with RPB5 and TFIIB, a general TF that facilitates transcription by interacting with the pre-initiation complex (Fig 5) [25]. A separate group used similar techniques to show interactions between TATA-binding protein and HBx. Collectively, these findings suggest that HBx enhances transcription efficiency of viral and host genes. Researchers moved outside the realm of general transcription events (RPB5, TBP, TFIIH) and looked for interactions between HBx and HCC-related signaling events.
HBx was shown to suppress apoptosis of hepatocytes via inducing survivin, disrupting the p53 gene, and activating c-jun N-terminal kinases [13, 12]. Each of these proteins have crucial roles in regulating the cell cycle; a hallmark of tumorigenesis is mutation of the p53 gene.
In infected cells, HBx induces overexpression of a cellular target, HBXIP, which functions as a regulator of centrosome dynamics and cytokinesis – critical aspects of mitosis [14]. Furthermore, it was reported that HBx activates src and the ras/raf/ERK (extracellular signal-regulated kinase) pathway, which leads to proliferation of quiescent cells [12]. Additionally, Kumar et al. demonstrated that HBx is significantly involved in evasion of the innate immune response [30]. The authors found that HBx binds a protein (IPS-1) capable of mediating IFN-β release. HBx-IPS-1 binding decreased the ability of cells to produce and secrete active IFN-β. In the context of immune system evasion, HBV has evolved much like how a tumor evolves.
More recently, Sung et al. (2009) performed the first ChIP-on-chip analysis of HBX-host interactions [22]. The core hypothesis of the study, summarized in Fig 6, was that HBx directly binds transcription factors that target genes involved in cancer development. The group first showed that HBx co-immunoprecipitates with transcription factors E2F1, SMAD4, and YY1. These TFs activate target genes involved in regulation of the cell cycle and apoptosis. It was shown that HBx interacts with the promoter of five target genes using anti-HBx antibodies and primers specific for the selected promoter regions. To verify that E2F1, SMAD4, and YY1 directly regulate transcription of target genes, the three TFs were knocked-down with siRNA and the transcript levels of their respective target genes were determined by RT-qPCR. Indeed, the experiment verified that each TF is responsible for regulating mRNA levels of target genes, and HBx has an affinity for these TFs which regulate cancer development. The authors plugged their biological finding into the REACTOME database using the NextBio search engine to identify "core pathways that may be deregulated by HBx through its binding to the promoters of target genes." Six pathways were identified, including the Jak-STAT signaling pathway. This is especially interesting in light of a body of literature which identifies the Jak-STAT pathway as overactive in individuals with HCC. This notion is highlighted in an article which appeared in Gastroenterology, "Ubiquitous Activation of Ras and Jak/Stat Pathways in Human HCC," (Calvisi, et al., 2006).
Collectively, these findings suggest that HBx is a key mediator of HCC development in HBV-infected individuals.
Overview of Interactions between HBV and Host miRNAs
Since the first report of a miRNA possessing antiviral properties in 2005 [15], researchers have begun to explore the possibility of interactions between host miRNAs and HBV. These interactions have provided mechanistic insights regarding the role of HBx in host gene expression and HCC development in HBV-infected individuals.
Several cellular transcription factors have an affinity for HBV enhancer elements and thus are crucial for efficient viral replication (Fig 8). A 2011 review reported C/EBP and CREB as TFs necessary for efficient viral replication. C/EBP is “CAAT enhancer-binding protein,” designated as such because of its affinity for the CAAT promoter motif, and CREB is cAMP-response element binding protein. Since publication of the review, Hu et al. reported peroxisome proliferator-activated receptor alpha (PPARA) as another host TF that binds viral enhancer elements and induces transcription of viral genes.
HBx-miRNA interactions are identified by several techniques. Hu et al. (2011) were interested in identifying a host miRNA that is capable of regulating HBV replication. The initial screen they employed was a three step process. First, the group selected 64 miRNAs from an expression microarray that are known to be involved in cell differentiation, viral infection, and cancer. For each miRNA, they co-transfected HCC cells with HBV DNA plasmid + miRNA or control plasmid + miRNA. Then, HBsAg levels for each of the 64 cell populations were assayed by ELISA to determine the ability of each miRNA to inhibit HBV replication (Fig 9).
In an effort to elucidate the precise host-virus interactions which result in HBx-regulated miRNA expression, Ren et al. overexpressed HBc, HBs, and HBc, and HBx proteins separately in the presence of overexpressed Drosha in HCC cells. They found decreased Drosha activity only in the presence of HBx. Inhibition of HBx by shRNA knockdown corresponded with increased Drosha activity. Considering Drosha is a key protein in the processing of functional miRNAs, this study provides strong evidence that HBx alters host gene expression through a complex network of interactions. Ren et al. provided a base of support for further study of HBx-miRNA interactions in HCC development. In 2011, Wu et al. showed that HBx down-regulates the miR-16 family in malignant hepatocytes. This family of RNAs targets various cyclins that are responsible for activating cyclin-dependent kinases, making these miRNAs key regulators of the G1/S cell-cycle checkpoint [2]. Also in 2011, Potenza et al. demonstrated that has-miR-125a-5p, a microRNA expressed in hepatocytes, down-regulates the expression of HBV S gene and, the corollary, levels of secreted HBsAg. The authors speculate that the in vivo effect of introducing this miRNA to localized sites would be decreased viral load. It has been shown that experimental knockdown of HBsAg leads to suppression of HBV replication [23].
X. Wei et al. (2012) investigated HBx-mediated expression of a miRNA (miR-101) that targets DNA methyltransferase DNAMT3A (Fig 11) [8]. The ability of HBx to down-regulate activity of a DNA methyltransferase suggests hepatitis B virus can influence epigenetic modifications to alter host gene expression. Evidence of miR-101 down-regulation in HBV-positive HCC tumor tissue suggests that miR-101 plays a tumor-suppressive role in HCC development [8]. The authors reported DNAMT3A targets several tumor-suppressor genes like RASSF1, APC, and CDKN2A. To summarize, HBx mediates increased expression of DNMT3A by decreasing miR-101 levels, thus decreasing expression of tumor-suppressor genes like RASSF1, APC, and CDKN2A (Fig 12). It is likely that these events contribute to the development of HCC in HBV infected individuals.
Current and Potential HBV Therapies
The seven drugs currently approved for the treatment of chronic hepatitis B in the US and Europe include IFN-α, pegylated IFN-α, and nucleotide analogues (NUCs) lamivudine, adefovir, entecavir, telbuvudine, and tenofovir [10]. In general, interferons are capable of inhibiting viral replication and stimulating an anti-viral immune response (mobilizing NK cells activity). NUCs inhibit synthesis of the viral genome by HBV polymerase by competing with natural NTPs for polymerase uptake. The lack of a hydroxyl group on NUCs prevents polymerase from forming an additional covalent bond, effectively halting DNA synthesis. Tenofovir, an adenosine analogue, is a common component of HAART regimens used to treat HIV infected individuals.
Despite extensive efforts to characterize HBx, a successful inhibitor of the viral protein has not been developed. Crystal structures of HBx bound to target TFs would be useful in developing small molecule inhibitors via rational drug design. For example, according to a 2012 issue of HBV Journal Review, researchers at UC-Boulder have identified two small proteins that help inhibit cell death, Bcl-2 and Bcl-xL, as the “bona fide” host targets of HBx [9]. The group determined that a result of HBx interaction with these proteins is increased intracellular calcium levels, which triggers necrosis in C. elegans through Ca2+ dependent proteases [18]. By utilizing the genomically simple C. elegans model, researchers hope to develop drugs that bind the Bcl-2/Bcl-xL recognition motif of HBx to reverse the negative effects of the viral oncoprotein without disrupting the normal function of Bcl-2/Bcl-xL.
In 2009, Deng et al. published promising results of an in vivo lentiviral therapy capable of decreasing viral load in mice [26]. The vector was cleverly designed to target the transcribed version of the HBV cccDNA genome. This pre-sliced RNA genome serves as both the immature template for translation and is stored as pre-genomic information. The authors used two shRNA duplexes "targeting the overlap region encoding the core antigen and the polymerase (pol)/surface antigen region (HBsAg) of the HBV genome" [26]. Decreased viral load in lentivirus-treated mice was confirmed by immunohistochemical staining, ELISA for HBsAg in serum, qPCR analysis of total HBV DNA in serum, and RT-qPCR analysis of HBV core-mRNA in serum (Fig 12).
In light of recent research regarding the role of miRNAs involved in HBV-related HCC, future studies could aim towards developing methods to restore miRNA functions in cases where HBx down-regulates these important and potentially antiviral sequences. It is possible that restoring proper miRNA function in HBV infected individuals will reduce their risk of developing HCC and even decrease the tumor burden in already-infected individuals. If Drosha function is indeed significantly inhibited in the presence of HBx [16], then researchers could investigate a method of restoring the activity of this crucial enzyme.
References
[1] Bouchard, MJ, and RJ Schneider. "The Enigmatic X gene of Hepatitis B Virus."Journal of Virology. 78.23 (2004): 12725-12734.
[2] Wu, G, F Yu, et al. "Hepatitis B virus protein downregulates expression of the miR-16 family in malignant hepatocytes in vitro." British Journal of Cancer. 105. (2011): 146-153.
[3] Kim CM, Koike K, Saito I, Miyamura T, Jay G. “HBx gene of hepatitis B virus induces liver cancer in transgenic mice.” Nature. 1991;351
[4] Terradillos O, Billet O, Renard CA, Levy R, Molina T, Briand P, Buendia MA. The hepatitis B virus X gene potentiates c-myc-induced liver oncogenesis in transgenic mice. Oncogene. 1997;14 (4:395–404.
[5] He, Yan, Hui-qing Sun, et al. "Knockdown of HBx by RNAi inhibits proliferation and enhances chemotherapy-induced apoptosis in hepatocellular carcinoma cells." Medical Oncology. 27. (2009): 1227-1233.
[6] Fatima , G, G Mathan, et al. " The HBx protein of hepatitis B virus regulates the expression, intracellular distribution and functions of ribosomal protein S27a." Journal of General Virology. (2011): 706-715.
[7] Shukla, SK, and V Kumar. "Hepatitis B virus X protein and c-Myc cooperate in the upregulation of ribosome biogenesis and in cellular transformation." FEBS Journal. (2012): 3859-3871.
[8] Wei, Xufu, and Tingziu Xiang . "miR-101 is down-regulated by the hepatitis B virus protein and induces aberrant DNA methylation by targeting DNA methyltransferase 3A." Cellular Signaling . (2012)
[9] Kukka, Christine. “Researchers Uncover New Ways to Prevent Viral Replication" HBV Journal Review. 9.11 (2012): 2-3
[10] Grimm, Daniel, Robert Thimme, et al. "HBV life cycle and novel drug targets.” International Journal of Hepatology. 5. (2012): 644-653.
[11] Qadri, I, H Maguire, et al. "Hepatitis B virus transactivator protein X interacts with the TATA-binding protein." Proceedings of the National Academy of Sciences. 92. (1995): 1003-1007.
[12] YC, Kim, and Song KS. " Activated ras oncogene collaborates with HBx gene of hepatitis B virus to transform cells by suppressing HBx-mediated apoptosis." Oncogene. 20. (2001): 16-23.
[13] Marusawa, H, S Matsuzawa, et al. "HBXIP functions as a cofactor of survivin in apoptosis suppression." EMBO Journal. 22.11 (2003): 2729-2740.
[14] Fujii, Ryoji, Changjun Zhu, et al. " HBXIP, Cellular Target of Hepatitis B Virus Oncoprotein, Is a Regulator of Centrosome Dynamics and Cytokinesis." Journal of Cancer Research. (2006)
[15] Lecellier, CH, P Dunoyer, et al. “A cellular microRNA mediates antiviral defense in human cells.” Science. 308. (2005): 557-560
[16] Ren, Min, Dongdong Qin, et al. "Correlation between hepatitis B virus protein and microRNA processor Drosha in cells expressing HBV." Antiviral Research. 94. (2012): 225-231.
[17] Liang, TJ. “Hepatitis B: The Virus and Disease." Hepatology. 49. (2009): S13-S21.
[18] Division of Viral Hepatitis. United States. Center for Disease Control. Viral Hepatitis Surveillance. 2010
[18] Geng, Xin, Brian L. Harry, et al. “Hepatitis B virus X protein targets the Bcl-2 protein CED-9 to induce intracellular Ca2+ increase and cell death in Caenorhabditis elegans." PNAS. 109.45 (2012): 18465-18470.
[19] Lee, GH, S Wasser, et al. " Hepatitis B pregenomic RNA splicing—The products, the regulatory mechanisms and its biological significance." Virus Research. 136.1-2 (2008): 1-7
[20] Wang, Wen, LJ Zhao, et al. “Application of HBx-induced anti-URGs as early warning biomarker of cirrhosis and HCC." Cancer Biomarkers. (2012): 29-39.
[21] Breiner, K, S Urban, et al. "Carboxypeptidase D (gp180), a Golgi-Resident Protein, Functions in the Attachment and Entry of Avian Hepatitis B Viruses." Journal of Virology. 72.10 (1998): 8098–8104.
[22] Sung, WK, Y Lu, et al. "Deregulated Direct Targets of the Hepatitis B Virus (HBV) Protein, HBx, Identified through Chromatin Immunoprecipitation and Expression Microarray Profiling." Journal of Biological Chemistry. 284.33 (2009)
[23] Potenza, N, U Papa, et al. "Human microRNA hsa-miR-125a-5p interferes with expression of hepatitis B virus surface antigen." Nucleic Acids Research. 39.12 (2011): 5157–5163.
[24] Qadri, I, ME Ferrari, et al. "The hepatitis B virus transactivator protein, HBx, interacts with single-stranded DNA (ssDNA). Biochemical characterizations of the HBx-ssDNA interactions." Journal of Biological Chemsitry. 271.26 (1996): 15443-50.
[25] Lin, Y, T Nomura, et al. "Hepatitis B Virus X Protein Is a Transcriptional Modulator That Communicates with Transcription Factor IIB and the RNA Polymerase II Subunit 5." Journal of Biological Chemistry. 272.11 (1997): 7132–7139.
[26] Deng, L, G Li, et al. "Hepatitis B virus inhibition in mice by lentiviral vector mediated short hairpin RNA." BMC Gastroenterology . 9.73 (2009)
[27] Beck, Juergen, and Michael Nassal. "Hepatitis B virus replication." World Journal of Gastroenterology. 13.1 (2007): 48-64.
[28] Cheng, Alfred A.L., Jun Yu, et al. "COX-2 mediates hepatitis B virus X protein abrogation of p53-induced apoptosis."Biochemical and Biophysical Research Communications. (2008)
[29] Liu, Wan-Hsin, Shiou-Hwei Yeh, et al. "Role of microRNAs in hepatitis B virus replication and pathogenesis." Biochemica et Biophysica Acta. (2011): 678-685.
[30] Kumar, M, SY Jung, et al. "Hepatitis B virus regulatory HBx protein binds to adaptor protein IPS-1 and inhibits the activation of beta interferon." Journal of Virology. 85.2 (2011): 987-95