Human Metapneumovirus
By Marta Hamilton
Introduction
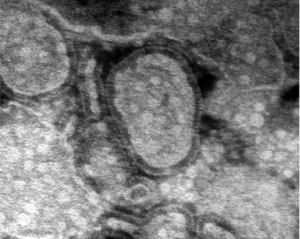
Respiratory tract infections, both upper and lower, can be caused by any number of factors, including several types of microbes (Crowe et al, 2004). Depending on the cause, respiratory tract infections can be difficult to treat and can increase the length of a patient’s hospital stay by weeks, with severity ranging from relatively minor to lethal. New causes of respiratory tract infections are still being discovered, and one such cause was recently identified.
A new virus has recently been characterized which is believed to cause both upper and lower respiratory tract infections, particularly in young children and the elderly (Boivin et al, 2002). First discovered in 2001 by Bernadette van den Hoogen and colleagues, while searching for the root cause of several respiratory infections in children, human metapneumovirus (HMPV) has since been extensively characterized and studied (Schildgen et al, 2011). HMPV was identified as a member of the paramyxovirus family, which contains many pathogenic viruses (Baker et al, 1999). HMPV was initially discovered when screens for known viruses were insufficient to discern the cause of infection in several patients (van den Hoogen et al, 2002). The isolates from these individuals were injected into monkeys, and HMPV was isolated (van den Hoogen et al, 2002).
Another common cause of respiratory tract infections, especially in children, is respiratory syncytial virus (RSV) (Crowe et al, 2004). This virus also has very high homology with HMPV, and the two share many genes which code for similar proteins (Peret et al, 2002). They share almost all of their genes, but RSV has two additional genes which code for NS1 and NS2 proteins (Peret et al, 2002). In addition to this, some of the genes which they share are present on their respective genomes in different orders (van den Hoogen et al, 2002). This could potentially be indicative of a need of slightly different concentrations of the proteins which are encoded by these genes, which could in turn indicate a difference in functionality between the two viruses (Schildgen et al, 2011). The two viruses also tend to cause similar symptoms in infected individuals, which is part of why HMPV was only discovered so recently (Crowe et al, 2004).
Another reason that HMPV could have only been isolated so recently is that it hasn’t been a distinct species for a very long time, according to phylogenetic studies which have been done on the evolution of the virus (van den Hoogen et al, 2002). One of the closest relatives of HMPV is avian metapneumovirus (AMPV), and the two are thought to have diverged approximately 200 years ago (de Graaf et al, 2008). Both viruses have four distinct strains which have been identified: A-D in AMPV, and A1, A2, B1, and B2 in HMPV. The two lineages of HMPV, A and B, diverged ~100 years ago, and then the sublineages began to differentiate about 50 years ago (Bayon-Auboyer et al, 2000, de Graaf et al, 2008).
Over the years, the varying strains of HMPV have altered frequency (Schildgen et al, 2011). For instance, HMPV A1 has only been rarely detected since 2004 but was much more common in the preceding decade (Schildgen et al, 2011). It has been hypothesized that the different strains become more prominent at different times to maintain susceptibility in the host population, rather than a single strain remaining as the primary infecting agent for so long that the host begins to develop immunity to that particular strain (Schildgen et al, 2011). The most common strain of HMPV tends to vary every decade or so (Schildgen et al, 2011). In 2004, a group in Germany thought that they had isolated a new strain of HMPV, but they could not culture it successfully, nor could they find it again after their initial study (Schildgen et al, 2011). No groups since have managed to relocate or isolate this supposed strain, indicating that perhaps it was never a novel strain at all (Schildgen et al, 2011).
AMPV-C is thought to be most closely related to the various strains of HMPVs. At the time of HMPV’s discovery, only avian strains of metapneumovirus had been characterized (van den Hoogen et al, 2002). HMPV is actually the first non-avian metapneumovirus to be identified and characterized, and as such, expands the metapneumovirus subfamily to include viruses which can be pathogenic to humans (van den Hoogen et al, 2002). AMPVs share high levels of sequence identity with HMPVs, as do HMPVs and RSVs (de Graaf et al, 2008). However, since AMPVs are only pathogenic to avian animals, there have not been extensive studies on the similarities between HMPVs and AMPVs, except from a phylogenetic standpoint (van den Hoogen et al, 2002). Much more is known about the relationship between HMPVs and RSVs, both of which infect humans, and both of which can cause very similar symptoms with similar levels of severity (Crowe et al, 2004).
Shortly after the initial discovery of HMPV, it underwent a period of study where it was genotyped and characterized fairly quickly. van den Hoogen et al, many of whom were on the same team that discovered the virus in the initial study in the Netherlands, presented the full genome of the virus in 2002. Every open reading frame and protein - several of which are discussed below - was examined in this study, and a phylogenetic analysis was done to investigate the evolution of HMPV (van den Hoogen et al, 2002). After this study concluded, attention began to turn to the medical knowledge which could be gained from the study of HMPV, as well as ways to treat the virus and prevent infections from becoming extremely severe in the target groups.
The genome of HMPV is composed of a single negative-sense single-stranded RNA which is approximately 13,000 nucleotides in length and has eight genes. Its eight genes code for proteins, three of which are surface proteins which are crucial in the propagation of the virus (van den Hoogen et al, 2002). All paramyxoviruses contain negative-sense single-stranded RNA and an envelope, so this is a characteristic of the family to which HMPV belongs (Baker et al,1999). According to several studies, HMPV is believed to replicate in a manner similar to other negative-sense single-stranded RNA viruses, but the latter half of HMPV's replication mechanism has yet to be examined for this specific virus (Baker et al, 1999, Schildgen et al, 2011). Some of the beginning of the mechanism has been examined, and thus far, it has followed the expected path for metapneumoviruses and paramyxoviruses (Baker et al, 1999, Schildgen et al, 2011).
There is plentiful current research available on HMPV, since it was discovered so recently and more knowledge surrounding the virus has incredibly useful medical applications. As such, a large portion of the research which has been undertaken in the past several years surrounds the virus' ability to cause disease, including its typical target groups, symptoms, and potential methods of treatment, including some evidence that it may be possible to produce a vaccine (Lèvy et al, 2013, Boivin et al, 2002). There have also been several case studies which examine patients infected with HMPV, and the mechanisms and patterns of infection which HMPV tends to follow have been observed (Dokos et al, 2013).
Susceptible Groups
Interestingly, HMPV seems to target primarily children under the age of five (van den Hoogen et al, 2002). Specifically, symptoms are seen between the ages of six months and five years (Bach et al, 2004). Until later studies were performed, HMPV was only identified in children under the age of five. However, shortly after van den Hoogen et al (2002) performed their second study dealing with the virus, another group began to examine the symptoms and effects of the virus. This group determined that not only were young children at a higher risk for severe HMPV infection, so were elderly patients (Boivin et al, 2002). This then makes elderly patients the second group identified as a specific risk group for HMPV. Healthy adults are very rarely infected with HMPV, but it has been shown that the virus functions better in situations where the individual infected is immunocompromised (Boivin et al, 2002). The third group at risk, then, consists of individuals who have compromised immune systems, for one reason or another. Immunocompromised individuals can include those who have autoimmune diseases (such as HIV/AIDS), individuals who have cancer, or those who have suppressed immune systems for other reasons, such as people who are on immunosuppressants following an organ transplant.
Given the above information, the three major target groups for HMPV are young children, elderly individuals (over the age of 65), and those who are immunocompromised. The illnesses which result after infection occurs also vary based on age group. Children under the age of five tend to suffer more from pneumonitis and bronchiolitis, while patients over the age of sixty-five tend to suffer from bronchitis and pneumonitis (Boivin et al, 2002). The patients who have lowered immune function also suffer most commonly from pneumonitis (Boivin et al, 2002). Boivin et al (2002) also focused on more severe cases than did van den Hoogen et al (2002), providing valuable insight into the degrees of severity which can be related to HMPV infection. The virus also does not spread from the respiratory tract, but infects solely this area of the human system. In addition, HMPV has never spread in any of the model organisms which have been used for its study, indicating that something about the virus causes it to preferentially infect only the respiratory tract (Crowe et al, 2004).
HMPV readily infects the above groups of people; however, it has a fairly low infection rate among healthy individuals, which means that it is rarely the primary cause of ailment among this group (Boivin et al, 2002, Crowe et al, 2004). In fact, healthy individuals usually only express HMPV symptoms when some other ailment has caused the immune system’s function to decline (Boivin et al, 2002). This, in turn, means that HMPV can be a cause of hospitalization, but usually only when the patient is already afflicted with some other illness. There is a chance of coinfection with HMPV and it can coinfect with several other viruses and infectious agents (Bach et al, 2004). HMPV can exacerbate an illness which is already present without being the primary cause of illness (Bach et al, 2004). In addition, in immunocompromised individuals, HMPV infections can be several times more severe than in healthy individuals, and can, in some cases, be fatal (Dokos et al, 2013).
Symptoms
Many of the symptoms exhibited by those infected with HMPV are similar to those seen in individuals infected by RSV, and can include everything from minor coughing to bronchitis and pneumonia, with degrees of severity ranging from relatively minor to lethal (van den Hoogen et al, 2002, Boivin et al, 2002). The high degree of similarity between the symptoms could be one of the most prominent reasons why HMPV was not discovered until 2001 (Schildgen et al, 2011). Fortunately, tests have since been designed to screen for HMPV, which allows for differentiation between HMPV and RSV in clinical settings (Bach et al, 2004). Common illnesses recorded in HMPV target groups were examined by Boivin et al in 2002, where it was discovered that one of the most common resulting illnesses following HMPV infection is pneumonitis.
Perhaps one of the most interesting pieces of knowledge about HMPV is that, by the age of twenty-five, almost everyone has been exposed to the virus, and many have been infected with it, regardless of geographic location (Schildgen et al, 2011). However, as stated previously, in healthy adults, the virus has little to no noticeable effect and doesn’t last long in the system (Boivin et al, 2002). Infections tend to begin to be noticeable at an age of around six months, but if no symptoms of HMPV infection have been observed by the age of five, it is unlikely that the individual will suffer from HMPV-related illnesses unless the individual becomes immunocompromised later in life (Crowe et al, 2004). However, HMPV has been linked to some chronic respiratory conditions, particularly asthma (Schildgen et al, 2011). It has been demonstrated that individuals who are infected with HMPV early in life (before five years of age), have an increased risk of developing asthma later in life and may experience a more severe presentation of asthma than those who do not have HMPV-related asthma (Schildgen et al, 2011).
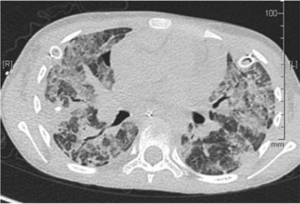
One case study examined the incidence of a ten-year-old girl who was admitted to the hospital after a stem cell transplant. The patient had cancer before the transplant and became infected with HMPV after, when her immune system was compromised as a result of the procedure (Dokos et al, 2013). Within a month of her admission to the hospital, the HMPV infection worsened enough to be lethal, as examined by Dokos et al, (2013), who took several scans of her lungs (Fig. 2). Had this infection occurred in an individual with a healthy, functional immune system, it likely would have only presented as a minor cough, which shows how HMPV can take advantage of a compromised immune system to thrive in a host’s respiratory tract (Boivin et al, 2002).
Vaccine Development
There has been some evidence that a vaccine for HMPV was created using virus-like particles (VLPs), rather than an attenuated form of the virus or an inactivated virus (Lèvy et al, 2013). According to this study, the VLPs are better for making a vaccine for this particular virus because, while the attenuated or inactivated vaccines can create an immune response in the test organism or patient, it does not have quite as good of a chance of lasting over a long-term time period (Lèvy et al, 2013). Because of this, the authors of this study chose to test the effects of VLPs, despite the fact that VLPs are harder to create for enveloped viruses (which all members of the paramyxovirus family are) (Lèvy et al, 2013). VLPs also have the advantage of being less dangerous to patients with suppressed or compromised immune systems because the VLP contains none of the genetic material of the virus, just a protein coat. This makes VLPs particularly applicable to the study of this virus, given that it primarily targets those who have immune systems which are not functioning properly (Boivin et al, 2002, Lèvy et al, 2013). The purpose of the VLPs is to mimic the surface proteins found on the surface of the HMPV virion (Lèvy et al, 2013).
HMPV has three glycoproteins which can be found on the surface of the virion: a G protein, an F protein, and an SH protein (Peret et al, 2002). The G protein serves the function of helping the virion attach to a host cell. The G protein is the protein which differs the most between the lineages of HMPV (van den Hoogen et al, 2002, Peret et al, 2002). The F protein facilitates the fusion of the virion membrane with the host membrane, allowing the virus to deposit its genetic material inside of the host cell (Peret et al, 2002). After that, the virus' genetic material is transcribed into positive-sense RNA before more proteins are made. The majority of HMPV's mechanism of reproduction has not been studied. However, it is likely that the mechanism will be similar to that of other paramyxoviruses (van den Hoogen et al, 2002, Peret et al, 2002). The third surface protein is the SH protein, the function of which has yet to be discovered (Schildgen et al, 2011). There is a significant difference in the length of the SH protein in HMPV and RSV, however, indicating that both viruses are still functional, even with their different versions of the protein. Studies were performed in which the SH protein was removed from HMPV, but it did not affect the ability of the virus to propagate or act as a pathogen (Schildgen et al, 2011).
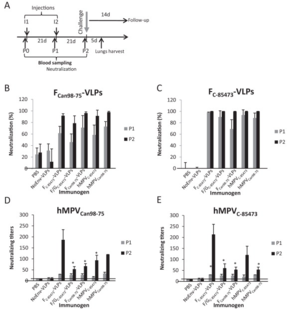
These three glycoproteins are mimicked on the surface of a VLP to trick the host immune system into developing a response to HMPV (Lèvy et al, 2013). The study was performed using a mouse model, where the mice were first injected with VLP for a prescribed amount of time (Lèvy et al, 2013). Blood samples were analyzed for antibody production, and at the end of the study, the mice were infected with HMPV (Lèvy et al, 2013). They were then compared to control mice (which had never been injected with VLPs) to see if there was an immune response difference (Lèvy et al, 2013). The mice which had been injected with VLPs were much more capable of resisting the HMPV infection (Fig. 3). To see if the treatment would still work across different strains of HMPV, the group had two test groups, each injected with a VLP for a different strain of HMPV (Lèvy et al, 2013). These groups were then split into two again, and of each group, half was infected with the same strain as their VLP, while the other half was infected with the strain which was different from their VLP (Lèvy et al, 2013). In both cases, mice which had been exposed to VLPs fared significantly better against HMPV than mice which had not been treated with any VLPs (Lèvy et al, 2013).
References
Edited by student of Joan Slonczewski for BIOL 238 Microbiology, 2013, Kenyon College.
