Klebsiella pneumoniae
A Microbial Biorealm page on the genus Klebsiella pneumoniae
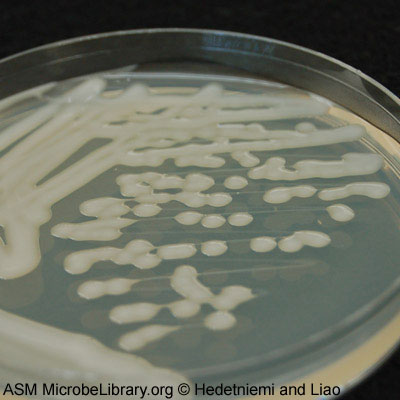
Classification
Higher order taxa
Domain: Bacteria;
Phylum: Proteobacteria;
Class: Gammaproteobacteria;
Order: Enterobacteriales;
Family: Enterobacteriaceae;
Genus: Klebsiella;
Species: K. pneumoniae
|
NCBI: Taxonomy |
Species
Klebsiella pneumoniae
|
NCBI: Complete genome here |
Description and significance
K. pneumoniae is a gram negative bacterium. It is facultative anaerobic. It is rod-shaped and measures 2 µm by 0.5 µm. In 1882, Friedlander C. Uber first discovered Klebsiella to be a pathogen that caused pneumonia (8). Many hospital cases around the world have been linked to K. pneumoniae. Therefore, more studies of the strains were important and performed. The bacterium was isolated and sequenced from a patient in 2004. K. pneumoniae is commonly found in the gastrointestinal tract and hands of hospital personnel (3). The reason for its pathogenicity is the thick capsule layer surrounding the bacterium. It is 160 nm thick of fine fibers that protrudes out from the outer membrane at right angles (6) (5). Another site on the human body that this bacteria can be found is the nasopharynx. Its habitat is not limited to humans but is ubiquitous to the ecological environment. This includes surface water, sewage, and soil (4). The frequent occurrence of resistant K. pneumoniae infections continues to spark great interest in research on how to control these resistant infections, possibly by preventing them with vaccines (7).
Genome structure
The complete genome was determined in 2006 at the Genome Sequencing Center at Washington University in St. Louis. The genome was named Klebsiella pneumoniae subsp. pneumoniae MGH 78578. It includes one chromosome of 5.3 Mbp. The GC content is 57%. There are five plasmids, pKPN3, pKPN4, pKPN5, pKPN6, and pKPN7. Respectively, each plasmid length is 0.18 Mbp, 0.11 Mbp, 0.089 Mbp, 0.0043 Mbp, and 0.0035 Mbp. The DNA is circular. In total, the genome encodes for about 5,300 genes. Of these 5,300 genes, 5,184 are protein genes, 111 are RNA genes, and 3 are pseudogenes. All RNA genes are located in the chromosome, while 1 pseudogene is encoded in chromosome, pKPN3 and pKPN4, respectively [1]. The sequence of K. pneumoniae genome was found to be closely related that of Escherichia coli K-12 (1).
Cell structure and metabolism
Cell Structure: K. pneumoniae contains a capsule around its cell. Known as K antigen, it is to protect the bacteria from phagocytosis. "K. pneumoniae strains of serotypes 01:Kl, 01:K10, and O1:K16, which have only the K antigen exposed at the cell surface, resist complement-mediated killing by impeding complement activation. It is also clear that purified capsular polysaccharides (K antigen) from nine different serotypes (able or unable to mask the LPS) were unable to activate complement (19)." In 1992, K. pneumoniae could be determined apart from other species of Klebsiella. Two oligonucleotide probes and the hydroxylated fatty acid C14:0-2OH are distinctive of K. pneumoniae (2). he capsule is divided into two layers, an inner and an outer layer, differentiated by the arrangement of bundle fibers that make up the capsule itself. The bundles fibers of the inner layer stand perpendicular to the surface of the outer cell membrane while the outer layer has fibers thinly arranged to form “a fine network structure” [7]. In addition to the capsule, the pathogenic strains of K. pneumoniae have been shown to synthesize type 3 fimbriae [6].
Metabolism:
Nitrogen fixation (nif) is unique to K. penumoniae. Enterobacteriaceae don't have the characteristic of nitrogen fixation. But, K. pneumoniae can take atmospheric nitrogen gas and reduce it to ammonia and amino acids. This was discovered by analysis of hisD-linked nif genes and hisD-unlinked nif genes (9). It was later found that the structural gene for glutamine synthetase (G.S.), glnA, and a closely related glnG regulates the nif genes for nitrogenase (10). Another metabolic product of K. pneumoniae is 1,3-propanediol. It is created by anaerobic fermentation (a two-step fermentation from glucose to glycerol to 1,3-propanediol) or mild aerobic fermentation (11). K. pneumoniae uses the operon nifLA to regulate the nitrogenase synthesis [5]. NifA acts as a transcriptional activator, while nifL inhibits the production of nifA in the presence of oxygen. In addition to these genes, other nif genes have been shown to also regulate the activity of nitrogenase [5]. In K. pneumoniae, there are approximately 20 nif genes that collectively regulate the metabolic activity. Due to its metabolic specificity, K. pneumoniae will only use nitrogen fixation in the absence of oxygen as oxygen can damage part of the nitrogenase enzyme [5].
Ecology
K. pneumoniae is ubiquitous as it is found in mammals and ecological environment. It has pathogenic effects worldwide. There is evidence of community-acquired and hospital-acquired infections in countries such as Taiwan and South Africa. Community-acquired K. pneumoniae has been found, in some places, to be associated with alcoholism. There are a large number of infections acquired when it affects different organs of the body. It can affect the liver, urinary tract, lungs, to name a few (18).
Pathology
K. pneumoniae is an important cause of human infections (Also see Description and significance). Infections or diseases are usually nosocomial or hospital-acquired. In 1998, K. pneumoniae and K. oxytoca accounted for 8% of nosocomial bacterial infections in the United States and in Europe. Diseases include urinary tract infections, pneumonia, septicemias, and soft tissue infections (3). The diseases caused by K. pneumoniae can result in death for patients who are immunodeficient. Differences in the diseases are determined by the different virulence factors. For example, mucoid phenotype varies as the strains for mucoid vary (14). CPS and LPS O side chain are two of the most important virulence factors of K. pneumoniae (7). They serve to protect the bacterium from phagocytosis by the host. Treatment is done by antibiotics such as clinafloxacin (13). But, there are an increasing amount of antibiotic-resistance strains. Ciprofloxacin is an antibiotic that is becoming less effective (12).
In California sea lions (Zalophus californianus) an isolate of the phenotypic characteristic hypermucoviscosity (HMV) of the bacteria Klebsiella pneumoniae has been found in a total of 25 cases. The HMV phenotype of K. pneumoniae was isolated from cases in which the sea lions had suppurative pneumonia and pleuritis; as well it was isolated from sea lions with abscesses. This is the first incidence of a pathogen that could be transmitted from marine animals to humans. Therefore, it is of great importance that marine mammals should be screened for pathogenic bacteria that could cause health problems in humans. Furthermore, K. pneumoniae HMV is starting to become more prevalent in marine coastal mammals as the primary pathogen. As a result, further studies of K. pneumoniae HMV are required to further improve and determine the extent of our understanding of this pathogen and its effects on epidemiology.(21).
Klebsiella pneumoniae is a very common pathogen that is encountered by many health care providers. Other than being a hospital-acquired pathogen that causes several infections such as urinary tract, nosocomial pneumonia and intraabdominal infections, Pneumoniae has been identified as a community-acquired infection with fluctuating prevalence. Strong correlation has been established between the demographic and geographic distribution among world populations and the incidents of community-acquired infections caused by K. penumoniae. K. pneumoniae has been considered a respiratory pathogen that causes Pneumoniae, the sysmptoms include: toxic presentation with sudden onset, high fever, and hemoptysis. Diagnosis through chest radiograph looks for abnormalities such as bulging interlobar fissure and cavitary abscesses. Over the years the contribution of K. pneumoniae to the total community-acquired cases of Pneumoniae has severely declined, while its contribution to other disease states increased. Community Acquired K. pneumoniae has been responsible for increased number of bacteremic liver abscess cases, especially in central and far Asia. Patients with Klebsiella liver abscess also showed higher rates of occurrence for the following complications: pulmonary emboli or abscess, brain abscess, pyogenic meningitis, endophthalmitis, prostatic abscess, osteomyelitis, septic arthritis, or psoas abscess. It is important to note that the rates of infections caused by community-acquired K. pneumoniae vary among world populations. For example: the rate of meningitis caused by K. pneumonia increased more than two folds in some hospitals in central Asia, while the same infection caused by K. pneumoniae only accounted for less than 2% in a hospital in the United States.(24).
K. pneumoniae is pathogenic and is responsible for a large number of infections every year. The most common pathogenic strains of K. pneumoniae are the K1 and K2 capsular serotype [6]. In addition, a study done on the K1 and K2 serotypes discovered that the mucoid phenotype was present in greater than 94% of K. pneumoniae infections [6]. The main virulence factor in pathogenic K. pneumoniae strains is aerobactin (produced by K1 and K2 serotypes), which is also the main virulence factor in pathogenic E. coli strains [6]. In a recent study, hypervirulent strains were responsible for the deaths of five patients in China in 2013 [6]. Most cases of K. pneumoniae infections were the result of patient to patient transfer and were hospital acquired. Common symptoms of K. pneumoniae infections in the blood include fever, chills, rash, and light-headedness [7]. Infections in the lungs can lead to meningitis and can result in breathing problems. The most common treatment for K. pneumoniae infections are antibiotics but recent strains have been found to resist many common antibiotics including carbapenem.
Current Research
(1) Currently, Steven Clegg PhD at the University of Iowa is conducting a research of identifying regulation of genes that encode for fimbriae of enteric bacteria. K. pneumoniae is being used by taking the genes that encode for fimbriae and inserting it into and E. coli plasmid. Since E. coli fimbriae can adhere to mucosal surfaces of eukaryotes, investigation can be conducted of the regulation of expression, proteins, and amino acids. Understanding the growth of K. pneumoniae on eukaryotes or mucosal surface will enlighten the way of understanding biofilms of this bacterium inside of the human body. Infections and prevention of those infections can be determined as well (15).
(2) An outbreak of IMP-1 β-lactamase-producing Klebsiella pneumoniae occurred in Japan hospitals and was acquired for investigation. Enteric bacteria have evolved a metallo-β-lactamase (MBL) resistance. The bacteria were once susceptible to β-lactam antibiotics except penicillins, but now developed resistance to expanded-spectrum β-lactams, because of extended-spectrum β-lactamases (ESBLs). Tests were done on antibiotics to determine the resistant and susceptible ones. All species are resistant to β-lactams including carbapenems such as imipenem and meropenem. The ones susceptible are those that make MBL. Patients with the bacteria at the hospital would be given levofloxacin antibiotic of the imipenem-resistant strain. It resulted in susceptibility. However, MBL-producing K. pneumoniae infections in this hospital were caused by a device-related or healthcare-associated infection. What is concluded so far is the list of sensitive drug against IMP-1 β-lactamase-producing Klebsiellan pneumoniae. Its isolation led to the solution of proper handling at the hospital because infections are derived from growth on hospital devices during patient care (16).
(3) Since the number of effective antibiotics is declining, scientists have measured the effectiveness of anti-virulence molecules on K. pneumoniae. This molecule is antibacterial; it doesn't prevent growth in vitro but prevents biosynthesis of the Klebsiella capsule and lipopolysaccharides, a the two important virulence factors. D1 and D41 (related triazines) and D0 (inactive triazine) measured their minimal inhibitory concentration. Growth on either D1 or D41 but not on D0 identified the extent of conservation of the virulence molecule. Further tests can be done and extended to human use in order to find a alternative in anti-virulence if antibiotics don't work (17).
(4) K. pneumoniae is one of the top organisms causing infections in hospitalized patients. These organisms can cause serious bloodstream infections and other intrusive infections that can potentially lead to fatality. A study in a Jamaican hospital was conducted that focused on multidrug resistant (MDR) strains of K. pneumoniae over a 5 year period. These MDR strains include resistance to broad spectrum cephalosporins and other extensively used antibiotics. This study led to the conclusion that K. pneumoniae showed endemic persistence of select clones, as opposed to pandemic persistence. This led to the hypothesis that the transmission of the organism was largely due to patient-to-patient contact and to health care worker-to-patient contact in the hospital. (20).
(5) Since the 1980s there has been a presence of individuals with community acquired Klebsiella pneumonia infections resulting in primary liver abscesses. These liver abscesses are caused by the capsular serotype K1 and the gene magA. The majority of these cases have appeared in Taiwan and South East Asia. Therefore researchers believe that there is an increase genetic susceptibility in people of this descent. The K antigen is a capsular polysaccharide and is considered to be an important virulence factor of K. pneumonia. There are 77 serological K antigen types and studies from Taiwan have revealed that K1 is the primary serotype causing these liver abscesses. Liver abscesses caused by the K1 and the magA gene are associated with the new phenotype that presents with features that allow it to be distinguished from traditional liver abscesses. These features include community acquired origin, absence of underlying hepatobiliary diseases and the presence of liver abscesses with other invasive complications. The liver abscesses of this strain are less harmful than other bacterial liver abscesses. The magA phenotype is observed more frequently in Klebsiella pneumonia strains that are liver invasive rather than those that are not. The prevalence of the magA gene was highest amongst strains from northern Taiwan which suggests a geographic variation. Researchers observed 98% of bacterial isolates from liver abscesses carried that magA gene. This gene is believed to be involved in exopolysacharide biosynthesis. These exopolysacharides can protect the bacteria and contribute to its virulence. All isolates containing the magA gene were all serotype K1 which leads researchers to believe that the K1 capsular serotype is an important virulence factor. These particular infections have only been recognized since the 1980s which leads researchers to believe that a recent mutation in the genome of Klebsiella pneumonia caused by acquisition of the magA gene has increased the pathogenicity of this strain, with a special affinity to genetic features in those with South East Asian descent. (22)
The advent of noscomial infections across the globe has lead research concerning the effect of Klebsiella pneumoniae on the immune system. Cytokines IL-17 and INF-γ are the critical for host defense against pulmonary K. pneumonaie infection. These cytokines are regulated by IL-23 and IL-12 respectively. These cytokines are responsible for the pro-inflammatory response in defense against K. pneumoniae infection.(23)
(6) In 2006, the Korean Medical Journal published an article on Klebsiella pneumoniae bacteremia, specifically examining the implications of community versus nosocomial-acquired bacteremia. The study found that nosocomial-acquired bacteremia tends to associate most commonly with neoplastic conditions, as well as having a mortality rate more than twice that of community-acquired bacteremia. Furthermore, 33.3% of the nosocomial isolates demonstrated resistance to cephalosporin, and 22% to ciprofloxacin. Invasive medical procedures increased the likelihood of nosocomial infection, and previous usage of cephalosporin and ciprofloxacin increased the chances of resistance development. The study also showed that diabetes mellitus and chronic liver disease were most commonly associated with community-acquired bacteremia. These isolates displayed lower levels of resistance to cephalosporin and ciprofloxacin (3.7% and 4.2% respectively). (25)
(7) It has been observed that there are many organisms that are able to produce extended spectrum β-Lactamases (ESBL’s), with Klebsiella pneumoniae being the most common species to produce ESBL’s. Bacteria that are able to produce ESBL’s are not susceptible to treatment with third generation cephalosporins, which poses a problem in effectively treating patients who are infected with an ESBL producing organism. In Eastern Europe as many as 50% of K. pneumoniae isolates are able produce ESBL’s. The study presented in this article was conducted to observe the affects that different antibiotic treatments had on infections due to K. pneumoniae. This was an observational study by Paterson et al (2004) on bacteremia caused by K. pneumonia in 65 patients who survived at least 14 days after the onset of illness. Types of therapy used were: carbapenem monotherapy, quinolone monotherapy, cephalosporin monotherapy, β-Lactam/β-Lactamase inhibitor combination, and amino glycoside monotherapy. This study found a significantly higher mortality rate among patients who were treated with non-carbapenem-β-Lactam antibiotics, and quinolones. The mortality rate among patients that were treated with carbapenem antibiotics was 3.7%. It was found that over 98% of ESBL producing organisms are still susceptible to either imipenem or meropenem, both of which are carbepenems. The study concluded that the treatment of choice for severe infections caused by ESBL-producing organisms should be carbapenems. (26)
(8) Isolates of Klebsiella pneumoniae have been producing extended-spectrum β-lactamases, rendering them resistant to many classes of antibiotics such as β-lactam/β-lactamase inhibitor combinations, quinolones, and aminoglycosides. This limits the choice of treatment to carbapenems for the treatment of serious infections. However, there has been an increasing spread of carbapenem-resistant Klebsiella pneumoniae strains (KPC-Kp). This leaves limited therapeutic options against KPC-Kp (tigecycline and colistin). However, they are even starting to find KPC-Kp strains resistant to Collistin, and Tigecycine does not work as well against blood infections. Due to this reason, there is a need for the development of new antimicrobial agents that can fight off the multi-drug resistant strains of Klebsiella pneumoniae. Researchers are currently within the early developmental stages of a “neoglycoside” known as ACHN-490. This is supposed to be the next generation of aminoglycosides that is supposed to be able to bypass the mechanisms of resistance within the bacteria. The research team evaluated how effective ACHN-490 was against 102 multidrug-resistant Klebsiella pneumoniae strains (25 of those strains being KPC-Kp). The strains were selected based on resistance against 3 or more antibiotics. They tested the strains’ susceptibility to ACHN-490 and various other antibiotics. The results showed that ACHN-490 significantly lowered concentrations of Klebsiella pneumoniae more than all of the other aminoglycosides. The dilution concentration of strains against ACHN-490 were even lower than that of the drug with the least resistance in their armamentarium. This shows that ACHN-490 is a promising future aminoglycoside alternative for treatment against multidrug-resistant strains of Klebsiella pneumoniae - including KPC-Kp strains. (27)
(9) Another area of research surrounding "Klebsiella pneumoniae" is learning about its presence in different regions of the world and understanding each of the different strains of the bacteria. One study looked at approximately 450 cases of "Klebsiella pneumoniae" infection worldwide to determine the effects of regionality (specifically different environments and different patients). The authors of this study concluded that patient characteristics were the main variation factor in each of the infections rather than the environment of the region [29]. Regions with lower socioeconomic standards and high prevalence of other diseases tended to have the strains with the highest virulence. Strains found in South Africa and Taiwan had the highest lethality in mice, however the researchers could not draw conclusive evidence toward the causes of in the differences between the strains [29]. Another study done on a hypervirulent strain of "Klebsiella pneumoniae" in China set out to investigate the genes pertaining to hypervirulence. The researchers found a new sequence type (ST1797) that contains the genes that encodes for drug resistance and transmission. The strain studied (K1 hvKP) was particularly dangerous, killing all five patients that were infected [30].
(10) The last area of current research is dedicated to understanding how Klebsiella pneumoniae functions, especially pertaining toward how the microbe survives in a host’s body. Researchers are interested in learning how Klebsiella pneumoniae can live in a hostile environment (inside a host with an immune system). The lungs and respiratory tract contain an immune system defense mechanism called antimicrobial peptides that break down and kill foreign bacteria [31]. Research found that Klebsiella pneumoniae have a protective bacterial polysaccharide capsule that limits the AP surface interaction with the bacteria itself. As the APs slowly breakdown the protective capsule, Klebsiella pneumoniae can regulate its formation of the polysaccharide layer and increase its production in the presence of APs [31].
References
(7) [https://www.ncbi.nlm.nih.gov/pubmed/26358222 Lee, W.H., H.I. Choi, S.W. Hong, K.S. Kim, Y.S. Gho, and S.G Jeon. 2015. Vaccination with Klebsiella pneumoniae-derived extracellular vesicles protects against bacteria-induced lethality via both humoral and cellular immunity. Experimental & Molecular Medicine 47(9):183.
(9) Friedlander C. Uber die scizomyceten bei der acuten fibrosen pneumonie. Arch Pathol Anat Physiol Klin Med 1882. 87:319-24.
(16) Cleggs, S. "Adherence of enterobacteria to eucaryotic receptors" 2007.
(26) Kang, Chaol-In. "Community-Acquired versus Nosocomial Klebsiella Pneumoniae Bacteremia: Clinical Features, Treatment Outcomes, and Clinical Implication of Antimicrobial Resistance." J Korean Medical Science (2006): 816-22. Academic Search Premier. Web. 20 Apr. 2010. <http://www.ncbi.nlm.nih.gov/pmc/articles/PMC2721989/pdf/jkms-21-816>.
(29) Yu, V., D. Hansen, W. Ko, A. Sagnimeni, K. Klugman, A. Gottberg, H. Goossens, M. Wagener, V. Benedi. 2007. Virulence Characteristics of Klebsiella and Clinical Manifestations of K. pneumoniae Bloodstream Infections. Emerging Infectious Diseases (13): 986-993
(30) Zhang, R., D. Lin, C. Wai-chi, E.W. Chan, D. Gu, G.X. Chen, S. Chen. 2016. Emergence of Carbapenem-Resistant Serotype K1 Hypervirulent Klebsiella pneumoniae Strains in China. Antimicrobal Agents and Chemotherapy 60:709-711.
(31) Campos, M. A. 2004. Capsule Polysaccharide Mediates Bacterial Resistance to Antimicrobial Peptides. Infection and Immunity 72 (12): 7107–7114.
Edited by Bernadette Miszczyszyn / Katherine Ramsey, students of M. Glogowski at Loyola University
Edited by Lauryn Samelko / Ashley Morawski, students of M Glogowski at Loyola University
Edited by James Tasch / Ani Michl (Loyola University Chicago)
Edited by Allyson Flores, student of Rachel Larsen
Edited by KLB
Edited by Viral Patel/Mary Ayers (Loyola University Chicago)
Edited by Havil Maddela and Bahjat AlAdili students of M Glogowskiat Loyola University
Edited by Emily Coffey, Hailey Wouters, Victoria Giampaoli students of M Glogowskia Loyola University
Edited by Jennifer Baltes and Monica Colunga, students of M. Glogowski at Loyola University
Edited by Kelsey Patel and Kulin Panchal, students of M Glogowskiat Loyola University Chicago
Edited by Nicholas Gentile, Justin Goucher, Ryan Kobayashi, and Makenna Kobrin, students of Dr. Osborne at Boston University