Microbial production of recombinant chymosin
Introduction
Cheese production requires the use of a 323 aminoacids protein called chymosin (also known as rennin). Chymosin (36 kDA) is a proteolitic enzyme which is usually obtained from calf stomachs. The role of this enzyme is to coagulate the milk, which is very important for digestion of milk in young animals. Initially, chymosin is secreted as inactive prochymosin and then activated at low pH. Once it is active, it breaks milk protein k-casein in a specific point and effects clotting which is the first step in cheese production. By clotting the milk, a solid product called curd is obtained. Curd is then processed to make cheese[1,2].
The increasing world production of cheese, coupled with a decline in the number of slaughtered calves, has stimulated a search for alternative sources of chymosin. One of these alternative sources is the use of recombinant chymosin produced by microorganisms.
Recombinant chymosin is mainly produced by funghi, but it can also be produced using bacteria.
Chymosin mechanism of action
Milk consists of water, fat, protein, phosphate, lactose, citric acid and inorganics such as calcium phosphate. The protein component of milk can be divided into two groups, the casein fraction (αs1-casein, αs2-casein, β-casein and κ-casein) and the whey proteins (β-lactoglobulin, β-lactoalbumin, immunoglobulins and serum albumin). The casein proteins are the ones that will form the curd during cheese making, specifically κ-casein [3].
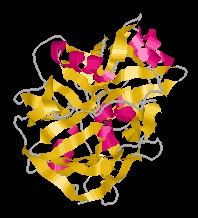
Chymosin, commonly known as rennin, is the main milk-coagulating enzyme that consists of a single polypeptide chain of 323 amino acids with intramolecular disulfide linkages. Preparations of calf rennet contain two forms of chymosin, A and B. The only difference between chymosins A and B is one amino acid in the polypeptide chain; the chymosin A contains an aspartic acid residue at position 286, whereas the chymosin B contains a glycine residue at the same position [2].
Like other acidic proteases from the gastric juice, chymosin is secreted as an inactive precursor, prochymosin. The precursor is stable at weakly alkaline pH whereas the active enzyme is rapidly denatured at pH values above 7. Below pH 5 prochymosin is converted into chymosin by a limited proteolysis during which a peptide segment is cleaved from the N-terminus. The proteolytic activity of chymosin has optimum pH about pH 3-5.
The milk-clotting activity of chymosin is due to the proteolysis of the κ-casein. κ-Casein consists of two parts: one part is hydrophobic and the other one is hydrophilic. In milk, this protein stabilizes the casein micelles against aggregation. During milk-clotting, a Phe-Met bond is hydrolysed, the hydrophilic part of the κ-casein is liberated and aggregation occurs[3]. Chymosin specifically recognizes the sequence from His 98 to Lys 111 and cleaves the peptide bond between Phe 105 and Met 106 in the κ-casein chain[4].
Production of recombinant chymosin
Recombination is the process by which genetic material is broken and then joined to a new genetic material. In this case, the chymosin gene from the cow is taken out of the genome and then introduced into a plasmid. The plasmid is then introduced into a microorganism which will start producing chymosin by transcribing and translating the gene from the plasmid.
The first step in this process is to obtain a piece of tissue from the calf stomach and isolate the DNA from the cells. After isolating the DNA it is necessary to amplify the chymosyn gene using PCR to increase the amount of gene fragments compared to other sequences of the genome. Another option is to isolate the mRNA directly and turn it into cDNA using the enzyme retrotranscriptase; this way we would have the DNA fragment without the introns as genes are found in prokaryotic microbes.
The second step is to introduce the gene sequence into plasmids that are then introduced in the target microbes. The gene is often introduced after a promoter sequence that we can control to increase its expression.
After introducing the gene in the plasmid and the plasmid in the microbe, it will start producing chymosin and the last step would be to purify it. The most common way of doing this is using a purification column in which chymosin will stick to the column due to its affinity for the matrix inside the column while the other proteins will not stick[2].
An important fact to be aware of while using recombination of genes from one species into a different one is the specific variation in codon usage. This problem usually causes low expression levels of the protein that we want to produce. To avoid this problem, the DNA sequence of the gene has to be modified and adapted to the codon usage of the species in which the chymosin will be produced, this is called codon-optimization[5].
Another important step in the production is the secretion of the protein by the microbial cells, the sequence of the chymosin can be also changed to improve its secretion. The modifications are often additions of secretory control regions of the microbe that is going to be used to produce the gene. One example of these strategies to improve production of chymosin is the addition in frame of the glucoamylase gente in Aspergillus, right after the prochymosin gene. Glucoamylase is a secreted protein that is secreted even with the chymosin protein attached to it. With this gene fusion a higher concentration of secreted prochymosin is obtained[6].
Some other useful tools to increase the production are the addition of transcriptional and translational control regions of the host species to the gene which may be not only specific promotors (as mentioned above) but also specific transcription terminators[7]. A common way to control the production (and increase it) is to add a promotor that is inducible by some sort of molecule. When we add that molecule (inductor) to the culture media the microbe will start transcribing our gene and producing chymosin. One example of inductor that is used when working in Escherichia coli is the molecule IPTG, which is a lactose analog and binds to the represor protein in the lac operon, thus inducing it. Using the lac operon promotor in front of the chymosin gene and the IPTC as inductor increases the production of the protein[8].
Microbes used in chymosin production
The most common microbes used for the production of recombinant chymosin are funghi, but this protein can also be produced using bacteria. Three of the most used microbes are Aspergillus niger, Kluyveromices lactis and Escherichia coli. These three species have really well known genetics and metabolic pathways, specially Aspergillus niger and Escherichia coli, which make them very useful for recombination experiments like chymosin production.
Aspergillus niger
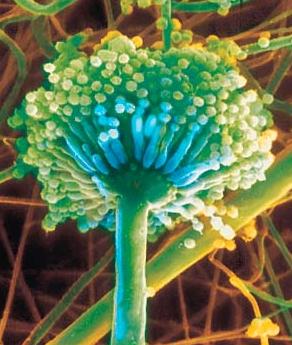
Aspergillus is the name used for a genus of molds that reproduce only by asexual means. Aspergillus species are common and widespread. They are among the most successful groups of molds with important roles in natural ecosystems and the human economy[9].
Aspergillus niger is one of the most common species of this genus[10]. It is found in a large variety of environments, but especially on vegetation, decaying organic matter, and soil. In humans, an Aspergillus niger infection is only a problem with people who have a weakened immune system. It most commonly causes lung infections, but it can also spread to other organs and open wounds[11].
A. niger first acquired practical importance in 1919, when its ability to produce citric acid was industrially exploited. Gluconic and fumaric acids have been produced with A. niger, although they are of less economic importance. However, since the 1960s, A. niger has become a source of a variety of enzymes that are well established as technical aids in fruit processing, baking, and in the starch and food industries. Gene technology has been successfully applied to improve production processes and to make use of A. niger as an expression system for foreign proteins[12]. That last part is the reason of using this microbe for the production of chymosyn, A. niger is used in a lot of different biotechnological processes, so the genetics are perfectly known. Another reason for using A. niger is that many of the enzymes produced by A. niger, such as citric acid, amylases, lipases, cellulases, xylanases and proteases, are considered GRAS (generally recognized as safe) by the United States Food and Drug Administration[12], so it is not considered a risk for human health.
The most used A. niger in chymosin production is Aspergillus niger var awamori. A glucoamylase-overexpressing mutant obtained from the NRRL 3112 strain was subjected to mutagenesis and selection in order to isolate two auxotrophic mutants pyrG (deficient in orotidine-5'- monophosphate decarboxylase; requiring uridine for growth) and argB (deficient in ornithine transcarbamylase; requiring arginine for growth). These strains were crossed and a double mutant pyrG argB was isolated from the progeny and designated as GC12. The purpose of these modifications was to provide selectable markers for the subsequent genetic manipulations. In the next step, the GC12 strain was modified by replacing its gene coding for aspergillopepsin A, an extracellular aspartic proteinase that can degrade chymosin and cause off-flavors in cheese, with the A. nidulans argB gene. A linear DNA fragment containing 5' and 3' flanking regions from the aspergillopepsin A gene and the A. nidulans argB gene was used to transform the GC12 strain. The transformants were selected for the loss of the arginine auxotrophy. The expression vector designated pGAMpR was constructed on the basis of pBR322 plasmid. The vector contains the A. awamori glucoamylase promoter and glucoamylase coding region fused in frame with bovine prochymosin cDNA and followed with the A. niger glucoamylase terminator. The vector also contains the pyr4 gene from Neurospora crassa which can complement the pyrG mutation in the host strain and allow for the selection of transformants in which the requirement for uridine was lost. The aspergillopepsin-deficient strain GCdeltaAP4 was transformed with the expression vector. The cells are grown aerobically under the proper conditions of pH, temperature, nutrient composition, etc. After chymosin reaches a desired level in the fermentation broth, the fermentation is stopped and the fungal cells are separated from the liquid. Chymosin is then recovered from the broth. For the recovery, the broth is filtered, followed by chromatographic purification and concentration of chymosin[13].
Kluyveromyces lactis
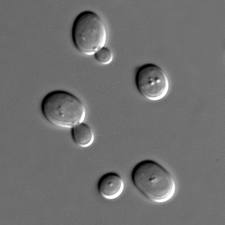
Kluyveromyces lactis is an ascomyceteous yeast (very similar to Saccharomyces cerevisiae) which has the ability to assimilate lactose and convert it into lactic acid, in fact, it has been isolated from milk and constitutes the predominant eukaryote during cheese production.
K. lactis is used for the production of beta-galactosidase and its fermentation properties are well understood. It shares most of the features that make Saccharomyces cerevisiae the best eukaryotic model organism (the existence of well-established genetics, the availability of fast and efficient transformation procedures and its efficient homologous recombination machinery) and it is also important that this microbe is used in other stages of the cheese production chain too. It is also considered a GRAS microbe[14,15].
It was found that chymosin could be produced in this host with good levels of secretion into the media. The chymosin gene is inserted into the K. lactis chromosome (usually by means of an expression vector derived from the pUC18 plasmid) and the yeast is grown by fed-batch fermentation (feeding of a growth limiting nutrient substrate to the culture). After fermentation, the yeast is killed by addition of benzoic acid and the chymosin is isolated by filtration[16,17].
Usually, an intermediate host is needed to introduce the prochymosin gene in K. lactis. The preprochymosin cDNA is cloned into pBR322 plasmid and amplified in E. coli. DNA encoding prochymosin produced by E. coli is then isolated and used for the construction of the expression vector on the basis of pUC18 plasmid that is then introduced in K. lactis[13].
Escherichia coli
Escherichia coli is a bacterium that is a common inhabitant of the human colon. It also lives in the intestine of many other animals, wild as well as domestic. It is a gram negative, rod-shaped gammaproteobacteria. It is the model organism for the study of bacteria. Normally, Escherichia coli does not cause disease although some strains frequently cause diarrhea, and it is the most common cause of urinary tract infections.
Escherichia coli is one of the most thoroughly studied of all living things. It is a favorite organism for genetic engineering since its genetics and metabolism are the best understood. Another advantage of studying Escherichia coli is that it can be grown very easily and inexpensively in a laboratory setting. The enzymatic properties of recombinant E. coli chymosin are indistinguishable from those of native calf chymosin[18] which is also important because the efficiency of the process of cheese production is the same.
The problem with E. coli, however, is that recombinant proteins are frequently synthesized as intracellular inclusion bodies and not secreted, increasing process costs considerably. After disintegration of cells, inclusion bodies are harvested by centrifugation. Another issue with E. coli is that it is not generally recognized as safe for human consumption[2].
One example of production in E. coli is the K12 JA198 strain. This strain was subjected to several genetic manipulations to construct the recipient strain for the expression plasmid carrying the prochymosin A gene. The expression plasmid was derived from the widely used cloning vector pBR322. cDNA coding for bovine chymosin A was previously cloned and characterized. The prochymosin gene was divided into three sections, each terminated by a unique restriction endonuclease recognition site. All three subcloned sections were assembled together in the correct order to reconstruct the prochymosin gene, which was inserted into pBR322 vector.
The production strain is grown in an aqueous solution containing carbohydrates, nitrogen, mineral salts and miscellaneous inorganic and organic compounds. The solid prochymosin is liberated from the producing organism by cell disruption and harvesting of "inclusion bodies" by centrifugation[13].
Conclusion
Due to the decrease in the number of slaughtered calves and the ethical problem that killing a huge amount of animals causes, it is important to have an alternative way of producing cheese that does not require the sacrifice of any animal. Moreover, microorganism production of recombinant chymosin offers an easy way of increasing the production of chymosin compared to the amount that can be obtained from young calves.
It is important to consider that some modifications may be needed in the gene sequence depending on the microbe that is going to be used for production of any recombinant protein due to the variation between species of the codon usage. Taking this into account may increase dramatically the production of the desired protein. It is also important to modify that sequence to achieve the higher production levels, and that is what current research is focusing on.
References
[9] Joan W. Bennett. "An Overview of the Genus Aspergillus".
[11] Aspergillus niger.
[16] Wésolowski-Louvel, M., K.D. Breunig, and H. Fukuhara, Kluyveromyces lactis. In: K. Wolf, editor. Non-conventional yeast in biotechnology. Heidelberg: Springer- Verlag, 1996: p. 139-201.
[17] Lachance MA. "Current status of Kluyveromyces systematics". FEMS Yeast Res. 2007 Aug;7(5):642-5.
Edited by Enrique Rodriguez, a student of Nora Sullivan in BIOL187S (Microbial Life) in The Keck Science Department of the Claremont Colleges Spring 2013.