Prochlorococcus marinus
A Microbial Biorealm page on the genus Prochlorococcus marinus
Classification
Higher order taxa
Bacteria (Domain); Cyanobacteria (Phylum); No Class Listed; Prochlorales (Order); Prochlorococcaceae (Family); Prochlorococcus (Genus)
|
NCBI: Taxonomy |
Species
Prochlorococcus marinus
Description and significance
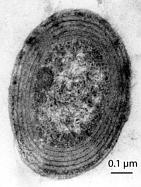
Prochlorococcus marinus was fist discovered and classified by Sallie W. Chrisholm in 1988. [l]
Prochlorococcus marinus is the dominate photosynthetic organism in the ocean. It accounts for up to 60% of the ocean’s chlorophyll in many regions such as the subtropical Pacific and is vastly numerous, often occurring at 10^5 cells/ml, abundant over the range of 40 °N to 40 °S The primary production and carbon sequestering capabilities in the open, oligotrophic, ocean are performed by these tinniest of cyanobacteria. P.marinus ranges in size from .6 to .eight um and have genomes of range l.6 to 2.6 Mbp. Most strains however, have genomes sized around 1.7 or 2.4 Mbp. Many experts understand such a condensed genome to be representative of the minimal genomic requirements for photosynthetic microbes.[l][2]
All P. marinus utilize divinyl chlorophyll b and a2 as their light harvesting photochemicals. Its identification is still greatly in flux as many potential systems (subspecies, ecotypes, multiple gene sequence comparison) of classification have merit and support for varying taxonomy. The primary and predominate distinction between types of Phroclorococcus marinus is the ratio of their chl b to chl a2. Strains fall into 2 categories: High B/A isolates that have adapted to low iridescence light levels and low B/A isolates which flourish under high iridescences of light. [l][2][6]
There is only one species in the genus. Thus, Prochlorococcus is synonymous with Prochlorococcus marinus.
Genome structure
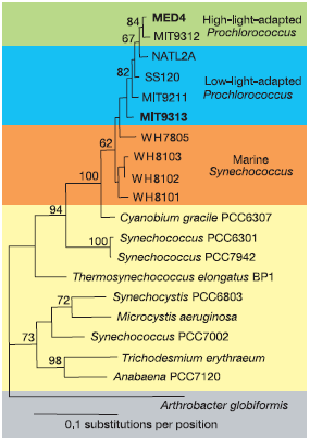
The strains of P.marinus all have circular DNA. Genome size ranges from l.6 to 2.7Mbp. Most strains however, have genomes sized around 1.7 or 2.4 Mbp. Many strains of Prochlorococcus have been sequenced. Its small genome and abundance in the accessible open ocean allow many studies to be done on its genome. [2]
All isolated strains are either low light adapted or high light adapted which corresponds to a high chl b to chl a2 ratio and a low ratio, respectively. The high B/A ratio, low light strains and the low B/A ratio, high light strains have do not have distinct genome sizes: all prochlorococcus have ~1.7 or 2.4 Mbp but this division does not always fall along the niche divide. The high light clade can be fairly divided into to subgroups based on size of genome. Members of the high light clade are very similar genetically. The divisions are less clear and diversity is more multifaceted in the low light clade. MED4 is often used as a reperesentitive high light strain with l.66 Mbp, 1,716 coding genes and a 30.8 G+C%. MIT9313 is most cited low light strain with 2.41 Mbp, 2,275 coding genes and a 50.6 G+C%. [2][4]
The 16S rDNA sequences of the Prochlorococcus ecotypes correlate with their physiology. Strains of the low-B/A ecotype are phylogenetically very closely related (99% identity in 16SrDNA sequence). Strains of the high-B/A ecotype have a lower degree of identity in their 16S rDNA sequence (97 to98%) High-B/A Prochlorococcus strains also have a higher degree of sequence identity to Synechococcus ( a member of the same phylum and Prochlorococcus’ closest relative) than the low-B/A ecotype. Some high-B/A Prochlorococcusstrains and marine cluster A Synechococcus strains are not well resolved using 16S rDNA sequences. Thus, the sequences of the internal transcribed spacer (ITS) region between 16S-23S rRNA is utilized to further characterize these isolates. [2][5][6]
Genomic data supports the evolutionary model describing a common ancestor of Cyanobacteria leading to organisms with differing light harvesting methods(the major difference between Prochlorococcus and Synechococcus) From there, high light and low light clades evolved by a mix of riding themselves of unnecessary genes, and adapting to their niches.In both MED4 and MIT9313 over 30% of the genes have hypothetical or unknown function. [3][4][5]
Cell structure and metabolism
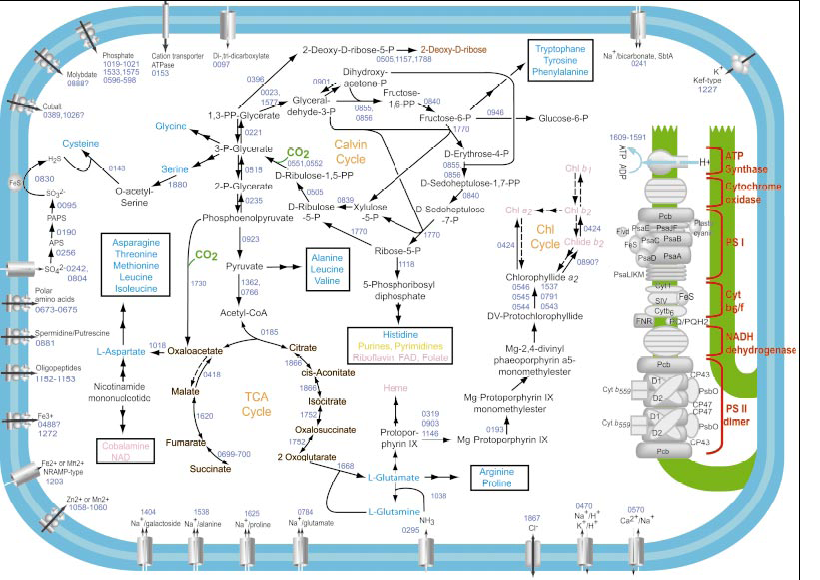
Such a compressed genome, especially of the very small, high light adapted strains, is lacking many basic functions previously considered important for the survival of cyanobacteria. Small genome Prochlorococcus MED4 is oval shaped and has no motility. It has very few transporters, mostly Sodium symporters. It has lost the ability to use nitrogen that is not in ammonia form. It lacks signal transduction capabilities and most cell and DNA repair mechanisms. It uses the Calvin Cycle to fix carbon but is lacking key TCA cycle enzymes making partial heterotrophy impossible. A consistent theme allowing such frugality of genome is the nonspecific nature of some enzymes in processes that normally require a dedicated protein. For example, a dedicated tyrosine aminotransferase is missing, and the last step in the biosynthesis of Phe, Tyr, and Trp is catalyzed by the nonspecific aromatic acid aminotransferase HisC. [2][3][4][6]
Such small cell size has two distinct advantages: to increase the in vivo absorption coefficient by reducing the package effect and to increase the cell surface to volume ratio and thereby improve nutrient uptake.[3]
The genus’ set of chlorophylls and method of light harvesting is unique amongst cyano bacteria. Cryptochrome like proteins PMM0425and PMM1360 are the genes that are most upregualted by increase and blue light and increased intensity. Researchers currently hold those genes, which do not have homologs in other cyanobacteria, to be responsible for Prochlorococcus’ light sensing ability.[5] Prochlorococcus has adapted to take advantage of relatively stable niches allowing it to sacrifices flexibility for efficiency and consistency. The picture to the right is the entire schematic layout of metabolism of a strain found in the Sargasso Sea. SS120 is a rarer, small genome but low light adapted strain.
Ecology
Prochlorococcus marinus is ubiquitous in open oceans over the range of 40 °N to 40 °S. The strains are adapted to different niches in that range bases on temperature, toxic metal content and depth. The distinct difference in ecotypes is those strains living in the first hundred meters of the ocean and those living in the second hundred meters. The light intensity differs one thousand fold from the surface to the low end of Prochlorococcus’ range. Prochlorococcus utilizes blue light which has the smallest coefficient of degradation over distances in the ocean. Still, high light and low light strains are adapted enough that the low light strains are inhibited at the surface and high light strains cannot survive with the levels of iridescences [5 µmol Q/m/m/s] where low light strains flourish. Additionally, high B/A isolates are intolerant to copper while low B/A strains that live on the surface where copper is more present, do not show this intolerance. [1][3][6]
P. marinus shares its niches with Synechococcus. There are examples of opposite seasonal cycles of these cyanobacteria. Prochlorococcus are at maximum in stably stratified, nutrient depleted waters in late summer and early fall when Synechococcus is at a minimum. When there is significant mixing of the water column during winter in the Gulf of Aqaba off the Red Sea, Prochlorococcus is undetectable but during the summer while the water is stratified, Prochlorococcus dominates small Synechococcus populations. Areas of the permanently stratified subtropical Pacific where water never mixed below the euphoic zone are consistently dominated by Prochlorococcus. This incompatibility to inconsistent substrate levels of mixing oceans is a result of a minimal genome without adaptive capability.[2][3]
Prochlorococcus also have capabilities evolved after their phylogenic departure from Synechococcus. Prochlorococcus have iron-related genes that are missing in Synechococcus, which may further explain its dominance in the iron-limited equatorial Pacific. Another example of divergence: MIT9313 has a 41.8-kilobase cluster of surface polysaccharide genes which has a lower percentage G+C composition (42%) than the genome as a whole, implicating lateral gene transfer. Thus, Prochlorococcus strains are not merely Synechocococcus that have lost or given up many of their genes.[6][7][8]
Application to Biotechnology
Prochlorococcus and its unique characteristics will help scientists understand the true essentials for autotrophy in microbes. It was ignored until 1988 and now, seeing its significance, many groups are studying it in order to understand the major role Prochlorococcus plays in global carbon cycles. It holds the power to fix carbon dioxide very quickly and efficiently, in large batches with very few requirements. Those qualities by themselves make Prochlorococcus a great target for climate remediation study. As of yet however, there have been no industrial uses of Prochlorococcus marinus.
Current Research
-One large point of research is the possibility of seeding the ocean with iron and producing massive blooms of cyanobacteria thus increasing the uptake of carbon by the oceans. The proposal to manage global warming by fertilizing vast areas of open ocean with iron firstgained prominence in the late 1980s, when oceanographer John Martin declared "Give me half a tanker of iron and I'll give you an Ice Age" [9]
Prominent scholars such as Sallie W. Chisholm, the discoverer of Prochlorococcus, have rebuked this idea but there remain many who are experimenting with ways to use iron to produce more carbon fixers in the oceans. See Chisholm’s paper on the subject at: http://web.mit.edu/chisholm/www/publications/fefert.pdf
-Prochlorococcus provides a simple way to study global Nitrogen availability. It is widespread, a simple microbe and is intolerant to much change in nutrient availability. Thus, changes in N levels will have widespread and observable effects on Prochlorococcus. A recent paper on the subject: Global gene expression of Prochlorococcus ecotypes in response to changes in nitrogen availability Molecular Systems Biology 2, (03 Oct 2006)
-P. marinus’ small size and frugal genome make it an ideal organism to study the effects of cyanophages, viruses that infect cyanobacteria. Without sexual reproduction devices, these bacteria have presumably obtained much of their genetic diversity from inter and intra species horizontal gene transfer. A recent paper on the subject of diversity in Prochlorococcus due to viruses: Viruses in the sea Curtis A. Suttle Nature 437, 356 - 361 (14 Sep 2005)
--Prochlorococcus’ simplicity makes it a good organism to study ocean cell mechanics. A recent paper on cyanobacterial proteins KaiA, KaiB and KaiC, as the authors call them: “dancing proteins” Shifting nanoscopic clock gears Susan S Golden, Vincent M Cassone, Andy LiWang, Nature Structural & Molecular Biology 14, 362 - 363 (01 May 2007)
References
1. Chisholm, S. W. et al. A novel free-living prochlorophyte abundant in the oceanic euphotic zone. Nature 334, 340–343 (1988): [1]
2. Rocap, G et al. Genome divergence in two Prochlorococcus ecotypes reflects oceanic niche differentiation Nature 424: 1042-1047. (2003) [2]
3. Gabrielle Rocap 2000 Genentic Diverstiy in the Marien Cyano Bacteria Prochlorococcus and Synechococcus. Ph. Thesis, Massachusetts Institute of Technology
4.Dufresne, A. et all. Genome sequence of the cyanobacterium Prochlorococcus marinus SS120, a nearly minimal oxyphototrophic genome. PNAS August 19, 2003 vol. 100 no. 17 p10020–10025: [3]
5. Chisholm, S.W. et al. Genome-Wide Analysis of Light Sensing in Prochlorococcus JOURNAL OF BACTERIOLOGY, Nov. 2006, p. 7796–7806 : [4]
6. Chisholm, S.W. et al.Resolution of Prochlorococcus and Synechococcus Ecotypes by Using 16S-23S Ribosomal DNA Internal Transcribed Spacer Sequences APPLIED AND ENVIRONMENTAL MICROBIOLOGY, Mar. 2002, p. 1180–1191 Vol. 68, No. 3
7 Campbell, L. et al. Annual variability of phytoplankton and bacteria in the subtropical North Pacific Ocean at Station ALOHA during the 1991–1994 ENSO event. Deep Sea Res. I 44, 167–192 (1997).
8. Mann, E. et al. Copper toxicity and cyanobacteria ecology in the Sargasso Sea. Limnol. Oceanogr. 47, 976–988 (2002).
9. Chisholm S.W., Iron fertilization of the Oceans: Reconciling Commercial Claims with Published Models Unpublished: [5]
Edited by Stephen Hubbard, student of Rachel Larsen
