Pseudomonas entomophila
A Microbial Biorealm page on the genus Pseudomonas entomophila
Classification
Higher order taxa
Bacteria, Proteobacteria, Gammaproteobacteria, Pseudomonadales, Pseudomonadaceae, Pseudomonas4
Genus and Species
Pseudomonas entomophila7
|
NCBI: Taxonomy |
Description and significance
Pseudomonas entomophila is a Gram-negative bacteria found in soil, aquatic, and rhizosphere environments. It was first isolated from the species Drosophila melanogaster as once ingested by this insect, it was found to cause lethality in the fly larvae and adults. Pseudomonas entomophila is significant because it is the first known Pseudomonas strain to be pathogenic in Drosophila melanogaster, even though the type III secretion system that is usually present in pathogens is absent.3 Pseudomonas entomophila's genome encodes insecticidal toxins, diffusible hemolytic activity, lipases, extracellular proteases, and potential adhesions which cluster with type I or II secretion system proteins. Being relatively harmless to plant life, Pseudomonas entomophila may be used in future insecticides.1
Genome structure
Pseudomonas entomophila's DNA is a single circular chromosome consisting of 5,888,780 nucleotides and its replicon type is a chromosome. 5,169 coding sequences have been discovered in its genome along with 107 RNA encoding genes, with 3,466 of those genes having predicted functions.5
Cell structure and metabolism
As mentioned previously, Pseudomonas entomophila is a gram-negative bacteria that has flagella. Its structural proteins consist of three proteins, PSEEN0141, PSEEN2177, and PSEEN3946, involved in adhesion to host surfaces and promotion of colonization. Its metabolism includes the pentose phosphate pathway, the Entner-Doudoroff pathway, the tricarboxylic acid cycle, and an incomplete Embden-Meyerhof-Parnas pathway due to the absence of 6-phosphofructokinase. Its genome encodes for hydrolytic activities, lipases, proteases, a set of 19 uncharacterized hydrolases involved in the degradation of polymers found within the soil.2 This bacterium's genome also encodes for a putative hydrogen cyanide (HCN) synthase, leading researchers to believe that Pseudomonas entomophila can produce HCN through cyanogenesis.8
Ecology
Pseudomonas entomophila are found in diverse environments such as the soil, the water, the rhizosphere, and the pathogenic interaction with Drosophila melanogaster. D. melanogaster maintain a hostile environment for microbes by secreting antimicrobial factors such as lysozymes and other digestive enzymes. However, Pseudomonas entomophila's genome contains four catalases, two superoxide dismutases, three hydroperoxide reductases, and eleven glutathione-S-transferases, which are involved in resistance against these oxidative stress produced by D. melanogaster to prevent the proliferation of microbes in the gut.2
Pathology
Pseudomonas entomophila's presence within the gut initiates a local and systemic immune response in the insect. Its virulence factor causes lethality within 1-2 days of high dose ingestion among Drosophila melanogaster larvae and adults. Not only is Pseudomonas entomophila lethal to Drosophila melanogaster, it is also lethal to other insects, meaning it is an entomopathogenic bacterium.6
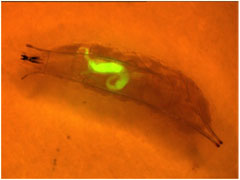
Previous studies have placed more emphasis on insecticidal toxins by this bacterium as they can possibly be used as biological control agents.6 Pseudomonas entomophila is highly toxic to insects, but its genome is absent of genes encoding enzymes able to breakdown plant cell walls, confirming its non-pathogenesis in plants. Pseudomonas entomophila have multiple genes involved in virulence including: PSEEN2485, PSEEN2697, and PSEEN2788, all of which are insecticidal toxins.2 This bacterium also secretes a diffusible hemolytic activity which is believed to be involved in pathogenicity in insects as the bacterial hemolysins act as exotoxins and cause the cells to rupture by attacking the blood cell membranes.2 Its genome also holds protease genes: three serine protease genes (PSEEN3027, PSEEN3028, PSEEN4433) and an alkaline protease gene (PSEEN1550).2
Protease genes contribute to the virulence among many different species, but in Drosophila melanogaster, Pseudomonas entomophila secrete a protease called AprA that reduces the amount of antimicrobial peptides produced in the intestines, therefore, inhibiting the intestines from ridding themselves of these harmful bacteria.6 However, Pseudomonas entomophila which cannot secrete the AprA protease are still found to be pathogenic, suggesting that this bacterium is has multi-factorial pathenogenesis.6 As previously mentioned, this bacterium is able to undergo cyanogenesis. HCN production was found to be regulated by the oxygen-regulator, ANR, which controls the hcnABC gene cluster that makes the HCN.8 This suggests that cyanide is a virulence factor used by this bacterium for its pathogenicity.8 Pseudomonas entomophila virulence is under the control of the GacS/GacA system which regulates secondary metabolite production, protein exudation, and virulence factors in gammaproteobacteria.6
Application to Biotechnology
Pseudomonas entomophila's characteristic pathogenicity in insects, combined with its non-pathogenicity in plants suggests great potential for a biological insecticidal in agriculture. It can be used to strictly target insects while being harmless to plant life. Also, its complete genome represents a type of template for further studies in pathogenic properties and host-pathogen interactions.2
Current Research
Today, sequencing of the entire genome of Pseudomonas entomophila is being done to obtain information on the various proteins present that are involved in metabolism, transport, regulation, and structure. Also being studied are the toxins and virulence factors produced by Pseudomonas entomophila to survive in different environments within its given ecology.2
In addition, Pseudomonas entomophila's pathogenic relationship mainly with Drosophila melanogaster as well as other insects from other orders is being researched along with local and systemic immune response, followed by lethality upon ingestion among Drosophila melanogaster adults and larvae. With this, scientists are studying the toxins, proteases, and hemolysins that contribute to its virulence and pathogenic nature.1
As previously stated, Pseudomonas entomophila have the potential for biological benefits as insecticides within agriculture. Their non-pathogenic behavior among plants is seen from its lack of enzymes within its genome capable of degrading plant cell walls. And although they are harmless to plants, they house multiple proteins and toxins in their genome that serve as pathogens leading to virulence among certain insects and factors that eliminate competition among neighboring microbes within soil, aquatic, and rhizosphere environments. This leads scientists to see the advantages of using this organism in insecticides3.
References
1. PubMed=16699499; [ NCBI , Israel , Japan ]
5. Kyoto Encyclopedia of Genes and Genomes (KEGG)
7. Comprehensive Microbial Resource (CMR) "Pseudomonas entomophila L48 Genome Page"
Edited by Jason Kim, student of Rachel Larsen and Kit Pogliano
Edited by Lauren Drewry, of Randolph-Macon College
KMG