Pseudomonas stutzeri
Classification
Domain Bacteria; Phylum Proteobacteria; Class Gammaproteobacteria; Order Pseudomonadales; Family Pseudomonadaceae NCBI
Species
Pseudomonas stutzeri
|
NCBI: [1] |
Description and Significance
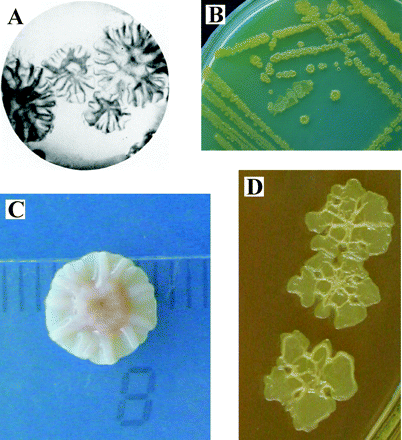
Pseudomonas stutzeri is a gram-negative bacterium that is known for its diverse metabolism. Individual P. stutzeri are rod-shaped and have a single polar flagellum. Cells are approximately
1-3um long and 0.5um in diameter. Colonies are disc shaped with ridges radiating from the center.
The presence of P. stutzeri in virtually all environments has led to it being called "almost universal." Soil and marine waters are two environments where P. stutzeri can be found. In soil, the organism has been found in the rhizosphere of cordgrass and other commonly grown plants including wheat, barley, and rice. In marine environments, P. stutzeri resides in the water column and sediment.
P. stutzeri's habitation of diverse environments and extensive range of metabolic functions makes it a very significant organism. As an organism that has some strains that are denitrifiers and others that are nitrogen fixers, P. stutzeri plays an interesting role in the nitrogen cycle. Nitrogen fixing P. stutzeri have been suggested to be significant in agricultural production and denitrifying P. stutzeri have been used as model organisms for denitrification. The organism's ability to metabolize a wide range of substrates may make it applicable in bioremediation and wastewater treatment.
Genome Structure
Three clinical strains of Pseudomonas stutzeri have been completely sequenced, providing a great deal of information about the content of the genome. The genome is encoded on a singular chromosome which is circular. No plasmids were detected in any of the three sequences. However, it is well known that P. stutzeri engages in natural transformation, which involves the uptake of plasmid DNA from the environment. The length of the chromosome of one particular strain was 4,547,930 bp, with the two other strains both being slightly longer. Genes for denitrification were found in two strains, but were absent in a third. As most strains of P. stutzeri are denitrifiers, the location and approximate length of the denitrifying genes on the chromosome has been studied. A cluster of about 30kbp, containing 33 genes, encodes information related to the structure of nitrogen oxide reductive enzymes, assembly processes, and electron donation. Other genes not located in this cluster also may play smaller roles in denitrification. All three strains contained genes encoding for benzoate and catechol degradation, as well as chemotaxis.
As P. stutzeri can be found in a wide variety of habitats, some mechanism must be present to allow the organism to adapt to a changing environment. Natural Transformation provides this bacterium with access to an abundance of naked environmental DNA that can be taken into the cell and integrated in the genome. DNA can be taken up not only from other members of the same species, but across species as well. This allows P. stutzeri to quickly adapt to changing environments and contributes to its vast distribution throughout the world.
Cell Structure, Metabolism and Life Cycle
Pseudomonas stutzeri are generally chemoorganoheterotrophs. This means that their energy source is dissolved organic molecules such as sugars, starches, amino acids, acetate, and pyruvate. It also means that they use organic molecules are their carbon source, rather than fixing and reducing their own carbon dioxide.
P. stutzeri are also denitrifiers. In other words they can use nitrate as an electron acceptor in their metabolism in place of oxygen. Nitrates undergo a four-step process in the cell, being converted to nitrite, nitric oxide, nitrous oxide, and finally nitrogen gas. P. stutzeri are used as a model organism for studying denitrification.
However, P. stutzeri, being found in a wide variety of habitats, is also a very metabolically diverse species. A wide variety of metabolisms have been observed in strains linked phylogenetically into the P. stutzeri species. Examples follow:
Some strains are able to use Thiosulfate as an energy source, producing H2SO4. Such strains are technically chemolithoautotrophic, since they use an inorganic compound as an energy and electron source.
Others have been shown to fix nitrogen in environments such as the roots of rice. In fact, some strains have even been shown to grow diazotrophically (in the absence of any nitrogen source other than N2 gas).
Other strains are able to oxidize either phosphite or hypophosphite in the absence of any other phosphorous source. Still other strains found in high-metal environments such as mines and polluted soil have been shown to resist those metals, often by converting them to an insoluble form. There are records of resistance of silver, zinc, and nickel ions, as well as tellurite and selenite.
It should be noted that despite this metabolic diversity (as well as genetic diversity), some common phenotypic characters are observed in all cases, and the species can be keyed out in the lab by those characteristics.
Ecology and Pathogenesis
The versatility of P. stutzeri's metabolism allows it to grow in a variety of conditions. It can be found in the soil and rhizosphere, where strains are known to be nitrogen fixers, thereby participating in a symbiosis with plant root systems
Some strains of P. stutzeri are of marine origin, and contribute to the nitrogen cycle by degrading nitrogen oxides to atmospheric nitrogen. Some strains have even been found present near hydrothermal vents in the Marianas Trench, the deepest part of the ocean.
P. stutzeri has also been found in wastewater, where its unique ability to degrade some anthropogenic compounds may have relevance to bioremediation.
P. stutzeri is an opportunistic pathogen. It has been isolated from clinical samples and has been found to cause infection in humans, though rarely. Many species of Pseudomonas have been shown to cause skin infections (Ecthyma gangrenosum) including P. stutzeri. Other cases include infections of prosthetic bone replacements. All but two cases of P. stutzeri have been treated successfully with antibiotics. The two cases that antibiotic treatment failed resulted in death of the patient. However, it was unclear whether the bacterial infection was the ultimate cause of death, as all patients infected with P. stutzeri had other health conditions prior to infection.
References
Author
William Baughman, Kathleen Balaze, and Andrew Bruce, students of Prof. Jay Lennon at Michigan State University.