Salmonella enterica NEU2011
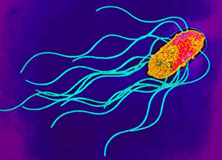
Transmission Electron Micrograph of Salmonella enterica. Photogragh from Todar's Online Textbook of Bacteriology (1).
Classification
Higher Order Taxa
Bacteria; Proteobacteria; Gammaproteobacteria; Enterobacteriales; Enterobacteriaceae
Genus & Species
Salmonella enterica
Description and Significance
Salmonella enterica is a gram-negative, rod-shaped, flagellated bacterium (figure 1) that is of interest due to its ability to cause infectious disease in humans and animals.(2). Human salmonellosis, S. enterica infection, occurs in about 1.3 million people per year, an estimated 30% of all food borne illness, causing about 500 deaths and has an estimated cost of $2.4 billion dollars per year (3). Salmonella sp. are facultative, being capable of both aerobic respiration for the production ATP as well as fermentation in the absence of oxygen. Dr. Daniel E. Salmon was the first person to identify Salmonella species over 100 years ago, currently there are approximately 2,300 serotypes known. S. enterica subspecies enterica serovar thyphi is a subspecies that is responsible for typhoid fever in humans by invading the gastrointestinal tract. Much of the current understanding of cellular and molecular functions in a cell has been made possible by using Salmonella. For example, S. enterica subspecies enterica serovar Typhimurium, was the first bacteria used to observe transduction by Zinder and Lederberg in 1952. Since then, a vast number of discoveries have been made using Salmonella ranging from gene regulation mechanisms, to cell surface antigen interactions with immune cells (1), (4).
Identification Characteristics
- PADase- negative
- Tryptophan deaminase- negative
- Urease- negative
- Indole- negative
- Lactose- negative
- Citrate-positive: can grow on Simon's Citrate Agar
- Produces acid and gas for glucose
- H2S produced from thiosulfate (1)
Genome Structure
The color scheme (explained in the key) shows what is specific to this serovar(blue) and what is specific to Salmonella(green) and what is found on all genomes (red). The outermost circle is the forward strand of the coding orientation and the inside circle is the reverse strand. The inner black circle is the G+C content; the purple, yellow innermost circle is the GC bias. The position of the origin of replication(ORI) and terminus(TER) are shown (6).
All serovar subtypes have as much as a 1% difference amongst each other in regards to net base pair differences.The genome of S. enterica differs from that of Escherichia coli by ~10%. The genome of S. enterica serovar Typhi Ty2 is almost 4.8 megabases long and is comprised of 52.05% G-C base pairs. There are 4323 genes, all of which encode for functioning proteins. There are 78 tRNA genes contained within the genome along with 22 rRNA genes.(5) Figure 2 shows a diagram of the genome of S. enterica serovar Typhimurium made up of 4,857 kilobases and a 94 kilobase virulent plasmid specific to the LT2 strain (6).
The genome can include plasmids but all subtypes have one main circular chromosome. (7)
The genome contains some outlying genes which are mostly unrelated to the genes contained within the base genome; some of these genes contain increased G-C base pairs which suggests that they were acquired more recently than most of the genome through a horizontal gene transfer. An example of these potentially newer genes would be the Salmonella pathogenicity island genes (abbreviated to "SPI", the two primary SPIs are known as SPI-1 and SPI-2) which affect the survival of S. enterica in eukaryotic cells. The study of S. Typhi and its single nucleotide polymorphisms (SNPs) has shown that it may be as young as 30,000 years old which would put its divergence far downfield from the divergence between S. enterica and Escherichia coli. (7)
Cell Structure and Metabolism
S. enteric in a gram-negative, peritrichously flagellated, faculatively anaerobic, bacilli.
Antigens
Salmonella species are characterized by three antigens present on the cell, as shown in figure 3.
These are the antigens used in serologic typing. Picture from Textbook of Diagnostic Microbiology (11).
- H antigen: This is the flagellar antigen. It occurs in either phase 1 or phase 2 and can alternate which is expressed (8),(9).
- O antigen: This is the somatic antigen. It is present on the outer membrane and its specificity is determined by the nature of the repeating units in the outer O-polysaccharide chain (8).
- Vi antigen: This is the virulence antigen. It is a capsular polysaccharide overlays the O antigen. It is not possessed by all serovars. This capsule is not necessary for infection, increase the infectivity by making it less detectable by the body’s immune system (10).
Fimbriae
S. enterica also possesses type I fimbriae, typical of other enteric bacteria, which aid in binding to epithelium cells (12). They are shorter than flagella but have a similar peritrichous distribution. They are made up of major and minor protein subunits that form a cylinder around a hollow core (13). Not all strains of S. enterica express fimbriae at the same time, but the fimbriaeted ones appear to be more virulent (11).
Siderophores
S. enterica also has the ability to to extract iron from host proteins by siderophores, which are made by the cell when iron concentration is low. There is also evidence that suggests siderophores contribute to virulence (14).
Metabolism
This bacteria exhibits mixed-acid heterofermentation of glucose to produce ATP. This produces CO2 and H2 with various acid end-products, such as formate, acetate, lactate, and succinate (15). S. enterica can also feed on maltose and maltodextrins through a type I, ATP-binding cassette(ABC) transporter. This system is encoded on the maltose regulon. Maltose is imported into the cell by crossing the outer membrane through a homotrimeric maltoporin, the maltose then binds to maltose-binding protein, which along with a protein with pore-forming subunits and an ATP-binding subunit is tranlocated across the inner-membrane where is can be used by the cell (16).
Ecology
Salmonella is ingested, and eventually invades the cells of the intestines. (17).
The main habitat where Salmonella is found is in the intestines of animals and humans (figure 4). Typical vectors of Salmonella enterica include chicken including their eggs, swine, dairy and beef cattle, and sometimes even insects, rodents, and other farm animals. S. enteric as well as other Salmonella species can live in the intestinal tracts of animals as part of the microbial flora without causing any disease to the animal but they will be passed out in the feces which can infect other animals, humans, vegetables and fruits. The life cycle of Salmonella enterica and related species follows that of a typical fecal to oral life cycle; the vast majority of the reproduction occurring in the intestines of the host or vector with a growth optimum of 37 degrees Celcius(3). Thus, it is also likely to find them inhabiting soil, water, and some plants due to improper sewage treatment of human waste or animal excretion into the environment. Reproduction of Salmonella scarcely occurs in the natural environment; however, cells can survive weeks in water samples and even longer in soil samples if the conditions are favorable (1). Salmonella enterica may be found in many foods due to fecal contamination by infected or carrier animals or humans (18). Most commonly, food that is grown in soil that is treated with manure to increase the fertility of that land can be contaminated with S. enteric as well as other pathogenic microbes. It was found that S. enterica will survive in both aerobic and anaerobic manure samples, thus it is not affected by aeration of manure compared to other pathogenic microbes such as E. coli (19). The increased cases of Salmonellosis have been linked to improper pasteurization of dairy products, improperly cooked ground beef and pork, and contamination from water runoff from farms (3).
Pathology
Scanning electron micrograph taken of S. enterica serovar Typhimurium invading human intestinal epithelial cells. (20).
Infectious diseases caused by S. enterica depends largely on secreted proteins and adhesion from both fimbrial and non-fimbrial sources to create a biofilm and contact with the host cell (21). Bacterial cells enter the cells that line the intestine, and the host cell’s membrane ruffles as a response to the initial attachment of the bacteria, this can be seen in figure 5. Ruffling is related to a triggering reaction that results in macropinocytosis. The vesicles of macropinocytosis are fairly large, providing an efficient route for non-selective endocytosis of solute macromolecules (22). The entrance of the bacterial cells briefly harms the microvilli on the cell surface. Following this disruption, white blood cells flood the mucosa which alters the homeostasis between absorption and secretion in the body (23).
S. enterica has several virulance factors. It has an enterotoxin, endotoxin, and cytotoxin. Diarrhea and vomiting are caused by the enterotoxin, which causes the cells to release vast amounts of fluid into the lumen. The endotoxin is comprised of the toxic lipopolysaccaride part of the outer layer of the cell envelope. This toxin results in a fever by causing the host cells to release endogenous pyrogens. The cytotoxin is responsible for interferring with protein synthesis in the host cell and causes an efflux of calcium ions, which rise and dislocate the cytoplasm (12). The infectious cells can then be transferred to the liver or spleen, where they multiply (8). S.enterica either returns to the host’s intestinal tract, or can be defecated. In addition to food contamination, further infection often occurs from infested feces present in polluted water, soil, or other unclean environments.
S. enterica is the cause of many cases of abdominal complications like gastroenteritis. When the balance between absorption and secretion is altered, initial symptoms include abdominal cramps, diarrhea, fever, and nausea. Most cases caused by food contamination resolve themselves within one week’s time, but more serious cases often need fluoroquinolone or cephalosporins, both are broad-spectrum antibiotics that can respectively kill the organism by stoping the cell wall from forming (24).
Watch a video of Salmonella invading the human intestanal tract.
Current Research
Much research is currently being done on drug-resistant, multidrug-resistant, and quinolones-resistant strains of Salmonella. Quinolones are classified as any class of antibiotics used in treating various infections, primarily gram-negative infections (25). A great deal information about the molecular mechanism of S. enterica resistance has recently come from sequencing the QRDR area of the gene gyrA (26). The gyrA gene encodes a subunit of DNA gyrase, and mutations in the QRDR area of the gyrA gene causes various resistant mechanisms to arise. Quinolones resistance commonly appears at low levels by gaining a preliminary resistance mutation, which can progress to stronger resistance levels to include resistance to second generation quinolones (first generation fluoroquinolones), and further on to second and third generation fluoroquinolones (27). This demonstrates how the Salmonella bacterium seems to discover ways to evolve and find new ecological roles, as we attempt to eradicate the bacterial pathogens. (28).
In addition to resistance studies, there is also investigation on faster detection of S. enterica. The two main mechanisms for detection are PCR based and culture based. PCR is beneficial in that it has the ability to be sensitive and specific. The indicated region is the only one that is coded for, so it can be done much quicker and give better specificity in that coded region. Culture based methods are more time consuming, but it codes the entire sample. In animals and their feed, the culture based methods have been found to be more successful when tracing areas because a detailed report can be compiled (29). In humans, the common method of detection via blood culture takes days and is not as effective as PCR suggested methods if treatment is needed immediately. This year, Zhou and Pollard have developed a sensitive blood culture PCR technique that can identify Salmonella enterica serovar Typhi in the same day with precise diagnosis (26).
Cool Factor
One of the interesting things that is being done with Salmonella is that researchers are using Salmonella to treat cancer in some mouse models. Some related research with S. Typhimurium is looking at infecting various cancer cells with Salmonella in order to target the cells as for immune response. Salmonella infection is more prone in some types of cancer than infection in the same tissue yet non-cancerous tissue and this difference leads directly to a selectable trait. The Salmonella tag allows for Salmonella-specific T cells to target the cancer cells and destroy them if they contain any Salmonella trace. This particular experiment used a melanoma model in mice for studying its effects; success rates for complete remission ranged from 50% to 100% in the experimental group and show strong signs of potential in the future for application to other diseases and work with other microorganisms. (30)
This research has widespread potential implications as things like novel cancer treatments can be pursued using this new advanced screening technique. What makes S. enterica so interesting is that it can tell cancer cells apart from non-cancerous cells while not letting the immune response take over and consume all of the same material.
References
11. Mahon CR, Manuselis G. Enterobacteriaceae. In: Mahon CR, Manuselis G, editors. Textbook of Diagnostic Microbiology. Philadelphia: W.B. Saunders; 1995.
12. Madigan M, Martinko J, Dunlap P, Clark D. Brock Biology of Microorganisms. San Francisco (CA): Pearson Education; 2009.
17. Aloe Vera Discovery (Internet). Forever Living products.(cited 2011 April 9).
26. Jimenez A, Moreno Y, Gonzalex A. Mendez-Vilas A, editor. Current Research Topics in Applied Microbiology and Microbial Biotechnology. Hackensack: World Scientific; 2009.
Edited by Abigail Bootes, Matthew Brennan, Alex Birch, and Danielle Laramee for BIOL2321 Microbiology, 2011, Northeastern University