Schizosaccharomyces pombe
A Microbial Biorealm page on the genus Schizosaccharomyces pombe
Classification

Higher order taxa
Superkingdom: Eukaryota, Domain: Fungi, Phylum: Ascomycota, Subphylum: Dikarya, Subphylum: Taphrinomycotina, Class: Schizosaccharomycetes, Order: Schizosaccharomycetales, Family: Schizosaccharomycetaceae, Genus: Schizosaccharomyces (1)
Species
Schizosaccharomyces pombe (1)
Also known as: Schizosaccharomyces malidevorans (1)
Description and significance
The fission yeast Schizosaccharomyces pombe is a unicellular eukaryote that is rod shaped. They measure approximately 2 to 3 microns in diameter and 7 to 14 microns in length (2). S. pombe is usually found in sugar-containing fermentations of alcohol from the subtropical regions (5).
Even though its origin dates back to quite a long time ago, it was not widely known before the 1890’s. It was discovered in 1893 when a group working in a Brewery Association Laboratory in Germany was looking at sediment found in millet beer imported from East Africa that gave it an unsavory acidic taste (8). P. Lindner was the first to describe Schizosaccharomyces pombe. He chose as its epithet the Swahili word for beer, pombe. It was identified as yeast, and it became known as the fission yeast because it reproduces by means of fission unlike its relative Saccharomyces cerevisiae. The name Schizosaccharomyces was assigned to it because Schizo- means “different,” which had been previously used to describe other fission species (8).
The sequencing of its genome was significant since S. pombe is a single-celled living archiascomycete fungus that shares many features with cells of more complicated eukaryotes (13). Researchers have identified fifty genes of S. pombe associated with human diseases including cystic fibrosis, hereditary deafness, and diabetes (13). Researchers state that the largest group of human disease-related genes are those implicated in cancer. There are 23 such genes, and they are involved in DNA damage and repair, checkpoint controls, and the cell cycle. All these processes are involved with maintaining genomic stability (13). These discoveries are important because it will allow researchers to find out more about the evolution of one-celled and multi-celled eukaryotic organisms compared to others such as bacteria, which do not have nucleated cells. Further analyses and comparisons should reveal which genes define eukaryotic cells and the transition from one-celled to multi-celled organisms (13).
Genome structure
S. pombe was the sixth organism with a nucleus to have its genome sequenced. S. pombe has a small and compact genome. Its genome contains 13.8 million base pairs distributed among chromosomes I (5.7 Mb), II (4.6 Mb) and III (3.5 Mb) and 4,824 genes. It has the lowest number of protein-coding genes yet identified in a free-living eukaryotic cell (13). The 3 chromosomes are linear, but circular chromosome formation has been observed in fission yeast mutants. If the S. pombe lose the simple repeats at the very ends of the chromosomes as well as much of the subtelomeric TAS elements, it will fuse end to end to generate circular chromosomes (14). The three centromeres are 35, 65, and 110 kb long for chromosomes I, II, and III respectively. This leaves about 12.5 Mb of unique sequence (13).
A total of 4,730 introns is distributed among 43% of S. pombe genes, with 15 being the largest number of introns found within a single gene. Of genes that have two or more introns, 614 have two introns, 324 have three, 148 have four, 70 have five and 40 have six (13). Therefore, the number of genes having an extra intron decreases by about half, as intron number increases from two to six per gene (13). These observations may be relevant when we study mechanisms by which introns are generated and removed. The large number of introns in S. pombe provides opportunities for alternative splicing to generate protein variants, which could have regulatory roles as well as increasing the range of protein types present in the cell (13).
The usefulness of S. pombe for investigating the functions of genes related to human disease is assessed. Protein-coding genes of S. pombe were identified that generate products with similarities to proteins coded by 289 genes that are mutated, amplified, or deleted in human disease (13). A total of 172 S. pombe proteins have similarity with members of this data set of human disease proteins. The largest groups of human disease-related genes are those implicated in cancer. There are 23 such genes and they are involved in DNA damage and repair, checkpoint controls, and the cell cycle, all processes in maintaining genomic stability (13). The cell cycle and checkpoint background of S. pombe make it a good model organism for studying these particular cancer disease pathways. Other categories represented in S. pombe are those involved in metabolic (12 genes), neurological (13 genes), cardiac (1 gene) and renal (1 gene) disease (13).
Cell structure and metabolism
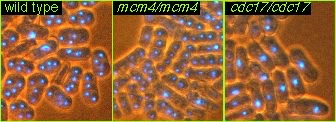
S. pombe is a rod-shaped fission yeast. If the cells have a mutation that prevents the normal progression of the cell cycle, they become very elongated. This occurs because the cells can continue to grow but are unable to divide. These mutants are called cdc mutants, for cell division cycle (6).
S. pombe is a chemoorganotroph, so it uses organic compounds as a source of energy and does not require light to grow. These fission yeasts can grow under both aerobic and anaerobic conditions. Fission yeasts are facultatively fermentative and exhibits aerobic fermentation in the presence of excess sugar (9). Alcohol dehydrogenase (ADH) catalyzes the reduction of acetaldehyde to ethanol in the last step of alcohol fermentation. This reduction is coupled with the oxidation of NADH and provides the NAD+ essential for the glyceraldehyde-3-phosphate oxidation in glycolysis. Therefore, ethanol production is important to maintain the redox balance in the cytoplasm (9). For a while, it has been widely assumed that S. pombe does not contain a mitochondrial ADH isoenzyme, and therefore does not have ethanol-dependent respiratory activity in the mitochondria (4). Ethanol-dependent respiratory activity is generally attributed to the presence of mitochondrial ADH isoenzymes. However, it was shown recently using genetic knockout strains that S. pombe does exhibit mitochondrial ADH activity, but the physiological function of yeast mitochondrial ADH enzymes is unclear (4).
S. pombe has evolved as a natural inositol auxotroph. Inositol is essential for the growth of all eukaryotic cells because it is a precursor of a major membrane phospholipid, sphingolipids, and glycosylphosphatidyl-inositol. These phosphorylated metabolites of inositol play an important role in the signal transduction pathways (7). It was discovered that S. pombe might have evolved as a natural inositol auxotroph because the natural environment of S. pombe contains a significant amount of phytic acid, which can be utilized as a source of inositol under very specific conditions. However, more research needs to be done in this area (7).
Ecology
S. pombe is found in alcoholic beverages, and therefore, it does not play a role in the environment. However, it is one of the yeasts that play an important role in the ecology of Kombucha fermentation (12). Kombucha is a traditional fermentation of sweetened tea, involving a symbiosis of yeast species and acetic acid bacteria. The study of yeast ecology in Kombucha fermentation reveals that S. pombe works with other yeast species such as B. bruxellensis, C. stellata, T. delbrueckii, and Z. bailii. S. pombe and other yeasts would ferment the sugar in the tea medium to ethanol, and then ethanol is oxidized by the acetic acid bacteria to produce acetic acid. S. pombe is suited to the Kombucha environment because of its ability to tolerate high-sugar foods (12). Isolates of S. pombe, T. delbrueckki, and Z. bailii exhibit tolerance up to 60% glucose concentration and are commonly associated with alcoholic fermentation for wine and champagne production. As the fermentation progressed, species with low acid tolerance decreased in population (12). Species such as S. pombe, with moderate tolerance to acidic conditions, die off after day 10. In general, Kombucha fermentation is initiated by osmotolerant species of yeast, which are capable of growing in the presence of high concentrations of sugar. The process is then succeeded and ultimately dominated by acid-tolerant species (12).
Pathology
The fission yeast Schizosaccaromyces pombe is a harmless, rapidly growing eukaryote. Therefore, there are no pathologies associated with this particular organism (8).
Application to Biotechnology
As of now, S. pombe is not used in any applications to biotechnology. However, it remains an important model used in researching cell division in molecular genetics and cell biology (4).
Current Research
A total of 437 human full-length cDNAs were isolated by microarray analysis of liver and/or gastric cancer tissues were evaluated for their relevance to cancer using S. pombe. Over-expression of 161 human cDNAs in S. pombe caused growth inhibition and/or morphological changes, which can be considered cancer-related phenotypes. Sixteen genes that caused growth defects were chosen to validate their oncogenic properties (3). They were highly expressed in liver and/or gastric cancer cell lines. When the mouse embryonic fibroblast cell was transfected with the genes, the proliferation rates of the cells increased by 32% to 120%. Therefore, the recent study demonstrated that S. pombe can be a powerful tool in the screening of human genes relevant to cancer (3).
S. pombe serves as a model system for studying the role of actin cytoskeleton, since it has simple actin cytoskeletons and is genetically tractable (11). However, fission yeast actin before had not been isolated in its native form and characterized. Therefore, biochemical assays of fission yeast actin-binding proteins or myosin have been performed using rabbit skeletal muscle actin that may interact with the fission yeast actin-binding proteins in a manner that is different from fission yeast actin (11). Recently, a novel method for isolating functionally active actin from fission yeast cells has been reported. The establishment of this new method will enable reconstruction of physiologically relevant interactions between the actin and fission yeast actin-binding proteins or myosins and contribute to clarifying the function of actin cytoskeleton in cellular activities (11).
Current research has also determined that a high-electric field application of S. pombe cells under hyperosmotic conditions improved the electro-induced release of protein and cytoplasmic enzymes. A cell suspension was pulsed at 12.5 kV/cm for 10 ms in a batch system and immediately postincubated at 30 degrees Celsius (10). The total protein release rate from cells increased in the hypertonic solutions of 1.5-2.5 M sorbitol. The protein release rate in 2.0 M sorbitol greatly increased up to 4.5 h and then gradually increased after 7.5 h (10). The hyperosmotic condition during and following single electric pulse application improved the release of large cytoplasmic enzymes. However, the amounts of released proteolytic enzymes were significantly lower than the ones obtained by conventional methods. This new discovery can be useful for the scale-up of large enzyme extraction processes from S. pombe(10).
References
2. http://www.ebi.ac.uk/2can/genomes/eukaryotes/Schizosaccharomyces_pombe.html
Edited by Anna Jan, student of Rachel Larsen
Edited by KLB
