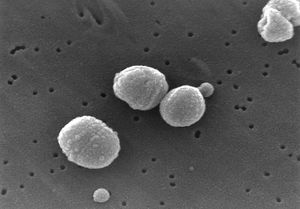
Streptococcus pneumoniae
A Microbial Biorealm page on the genus Streptococcus pneumoniae
Classification
Higher order taxa
Bacteria; Firmicutes; Bacilli; Lactobacillales; Streptococcaceae
Species
Streptococcus pneumoniae
Description and significance
Streptococcus pneumoniae are Gram-positive bacteria in the shape of a slightly pointed cocci. They are usually found in pairs (diplococci), but are also found singly and in short chains. S. pneumoniae are alpha hemolytic (a term describing how the cultured bacteria break down red blood cells for the purpose of classification). Individual bacteria are between 0.5 and 1.25 micrometers in diameter. S. pneumoniae do not form spores and are non-motile, though they sometimes have pili used for adherence. (5) S. pneumonia are found normally in the upper respiratory tract, including the throat and nasal passages. (2) They are mesophillic, living optimally at temperatures between 30 and 35 degrees Celsius.
S. pneumoniae was isolated in 1881 by Louis Pasteur. The species was then known as pneumococcus due to its role in the disease, pneumonia. It was termed Diplococcus pneumonia in 1926 due to its propensity to exist in pairs of cells, and renamed Streptococcus pneumoniae in 1974 because of its formation of chains in liquid.
S. pneumoniae played a significant role in the history molecular genetics, being the subject of the experiments that gave birth to the field. In 1928 Frederick Griffith transformed live, harmless S. pneumoniae into a deadly strain by combining them with an extract from heat-killed, virulent S. pneumoniae. Further, in 1944, Avery, MacLeod, and McCarthy proved that genetic material is DNA by showing that the transforming factor in Griffith's experiments was not protein, but DNA. (7)
Genome structure
The genome ofS. pneumoniae D39, the virulent strain used in the revolutionary experiments of Griffiths and McCarthy, consists of a single circular chromosome with about 2 million base pairs. Its GC content is 39.7%, and it codes for 1914 proteins and 73 RNAs. (2) Several surface proteins have been identified that may serve as vaccine candidates. Several different strains of Streptococcus pneumoniae have been identified, possibly accounting for differences in virulence and the presence of antigens. (1)
S. pneumoniae has shown a significant increase in antibiotic resistance over the past 20 years. This is likely due to its use of a natural transformation system used for genetic exchange. It can also develop resistance to antibiotics through mutation and natural selection. S. pneumoniae has a relatively fast growth rate and reaches great cell densities in infections environments, conditions which favor natural transformation and natural selection toward antibiotic resistance. (5)
Growing Resistance of Streptococcus pneumoniae
To treat bacterial infections caused by Streptococcus pneumoniae, the original antibiotic treatment was penicillin. Penicillin tends to interfere with the synthesis of peptidoglycan in bacterial cell walls. Peptidoglycan is a major component of bacterial cell walls and is a complex molecule made of sugars and polypeptides that form a tough, strong lattice that ends up surrounding bacterial cells. Penicillin prevents peptidoglycan from linking in the last stages of bacterial cell wall synthesis. This weakens the cell wall and causes the bacterium to burst open because of osmotic pressure. Penicillin is known to prevent infection by inhibiting the growth of bacteria because it directly kills bacteria. Since peptidoglycan is not found in human cells, once penicillin is injected into the human body it becomes selectively toxic and does not harm us in the process. Overtime, penicillin-resistant variants of S. pneumoniae have spread globally causing it to be a pervasive and problematic pathogen worldwide (Appelbaum, 2002). Resistance to penicillin involves modified structures of penicillin-binding proteins of S. pneumoniae, where these binding proteins allow peptidoglycan synthesis to occur. This resistance can often be overcome with high doses of penicillin, but overtime, this approach will not always necessarily work (Van der Poll et al. 2009). Penicillin resistant Streptococcus pneumoniae is noted as a major global health threat because S. pneumoniae is a prevalent bacterial pathogen of pneumonia and a major cause of meningitis and other associated diseases (Stanek et al. 2011). Although examples of infrequent antibiotic resistance in S. pneumoniae had been reported since the mid-1960, treatment of pneumococcal infections was predetermined until the early 1990’s because the resistance to β-lactam agents was low, therefore penicillin treatment could be relied upon in few cases (Morrissey and Tillotson, 2004).
Bacterial infections, such as Streptococcus pneumoniae, put a serious strain on the United States healthcare system with roughly thousands of deaths and hospitalizations every single year (Huttner et al. 2011). Penicillin resistance in S. pneumoniae has an effect on mortality rates of patients hospitalized with pneumonia. Many studies have shown that the mortality rates in patients infected by the penicillin-non susceptible S. pneumoniae were around 19.4%, and those infected by the penicillin-susceptible S. pneumoniae had a mortality rate around 15.7%. Penicillin resistance is therefore strongly associated with an increased mortality rate. Pneumonia caused by S. pneumoniae is most prevalent in poorer countries, therefore increasing the need for an economical and effective treatment, as well as understanding the mechanisms of this association is needed. Bacterial resistance to antibiotics depends particularly on the total usage of antibiotics, different selective power for bacteria depending on the antibiotic (Granizo et al. 2000). Many populations have showed that the increasing use of penicillin is associated with increased resistance in the pneumococci strain. Thus, increasing dosage comes with risks of promoting higher resistance.
The two primary antibiotics used to treat pneumococci infections are penicillin, as well as erythromycin, an antibiotic that is very similar in its effects to penicillin. These two antibiotics treat various bacterial infections and are most active towards gram-positive bacteria. Penicillin and erythromycin fall into the categories of β-lactam (antibiotic containing β-lactam ring in molecular structure) and macrolide (antibiotics containing a lactone ring in molecular structure) drugs. The primary cause of Streptococcus pneumoniae resistance to erythromycin is the consumption of macrolides taken twice a day. Macrolide consumption inhibits the growth of bacteria and are often prescribed to treat common bacterial infections, hence consumption throughout the day. There is rising speculation that another possibility of this streptococcal resistance is to whether the same macrolide consumption could also increase penicillin resistance (Granizo et al. 2000). Global consumption of antibiotics and the different capabilities of antibiotics to select resistance must be taken into account when studying antibiotic impact on bacterial populations. These in total suggest that the overuse of certain antibiotics is more likely to be related to the increase in drug-resistance stains of Streptococcus pneumoniae (Granizo et al. 2000).
Cell structure and metabolism
Streptococcus pneumoniae is characterized by a polysaccharide capsule that completely encloses the cell, and plays a key role in its virulence. The cell wall of S. pneumoniae is composed of peptidoglycan, with teichoic acid attached to every third N-acetylmuramic acid, and is about 6 layers thick. Lipoteichoic acid is attached to the membrane via a lipid moiety, and both teichoic and lipoteichoic acid contain phosphorylcholine. Two choline residues may exist on each carbohydrate repeat, which is important to S. pneumniae because the choline adheres to choline-binding receptors located on human cells. (5)
S. pneumoniae contains more than 500 different surface proteins. A notable group is the family of choline-binding proteins (CBPs). Twelve of these are bound to the choline moiety of the cell wall and assist in attaching various functional elements onto the surface of the cell. Among the CBPs are found important determinants of virulance, including PspA (protective antigen), LytA, B, and C (autolysins), and CbpA (adhesin). (5)
S. pneumoniae lacks catalase and ferments glucose to lactic acid, like most other streptococci. However, unlike most other streptococci, it does not display an M protein and it hydrolyzes insulin. (5) S. pneumoniae gets a significant amount of its carbon and nitrogen through extracellular enzyme systems that allow the metabolism of polysaccharides and hexosamines, as well as cause damage to host tissue and enable colonization. (1)
S. pneumoniae has the ability to enzymatically disrupt and disintegrate itself. This enzyme, an autolysin, causes a culture of S. pneumonia to undergo an autolysis, which kills the entire culture during the stationary phase (the phase in which growth slows due to exhaustion of available nutrients and buildup of toxins). With initial growth beginning in optimal conditions, autolysis usually begins within 18-24 hours, with colonies collapsing in the centers. (5)
Colonization by Streptococcus pneumoniae
Nasopharyngeal colonization involves colonization of the upper respiratory tract or the nasopharynx, which is the upper part of the throat behind the nose (Bogaert et al. 2004). The upper respiratory tract of the human body is home for many bacterial species. For the human body, the process of nasopharyngeal colonization of Streptococcus pneumoniae becomes established during the early stages of life. The mechanisms associated with nasopharyngeal colonization and colonizing species is the adherence to the epithelial lining (membrane tissue composed of many layers of cells) of the upper respiratory tract (Bogaert et al. 2004). Nasopharyngeal colonization provides important information about many pneumococcal diseases and its prevention. This type of colonization is not only mandatory for many diseases, but is crucial in regards to the spread of strains of pneumococci (Bogaert et al. 2004). The risk of colonization with strains of pneumococci are high under circumstances of the crowding of individuals, as well as the high rate of transmission of penicillin resistant S. Pneumoniae between children and their parents (Hoshino et al, 2002). Therefore, the introduction and usage of highly effective vaccines in young children is suggested for future prevention of invasive pneumococcal diseases. Infants and young children tend to acquire S. pneumoniae in the upper respiratory tract because of their low immunity (Hoshino et al, 2002). As a result, many studies agree that one way to prevent pneumococcal disease is to focus on the prevention of nasopharyngeal colonization (Bogaert et al. 2004). Some research suggests that the prevention of nasopharyngeal colonization in the early stages of life may reduce certain diseases caused by S. pneumoniae. There are difficulties that are associated with this prevention as this may lead to sensitivity towards other serotypes and other pathogenic species. In this case, this leads to a different array of diseases not taken into account for (Bogaert et al. 2004).
Ecology
Streptococcus pneumoniae is found in the respiratory tracts of mammals. While it is part of the normal flora of this environment, going unnoticed when present in small densities, S. pneumoniae essentially becomes a pathogen when its population becomes too large.
S. pneumoniae shares its mucosal microenvironment with another pathogen, Haemophilus influenzae. In vitro, competition between S. pneumoniae and H. influenzae leads to the former killing off the latter by attacking it with hydrogen peroxide. Tested individually in vivo, both bacteria are able to successfully grow; however, when both are present in the same environment, H. influenza rapidly outcompetes S. pneumoniae. Research shows that the inflammatory response elicited by H. influenza causes the clearing of the other pathogen, S. pneumoniae. (6)
Pathology
Streptococcus pneumoniae is known to cause bacteremia, otitis media, and meningitis in humans, though it is best known for causing pneumonia, a disease of the upper respiratory tract that causes illness and death all over the world. (5) Symptoms of pneumonia include a cough accompanied by greenish or yellow mucous, fever, chills, shortness of breath, and chest pain. The bacteria enter the body most commonly via inhalation of small water droplets. Very young children and the elderly are the most prone to catching pneumonia.
The virulence factors of S. pneumoniae include a plysaccharide capsule that prevents phagocytosis by the host's immune cells (5), surface proteins that prevent the activation of complement (part of the immune system that helps clear pathogens from the body), and pili that enable S. pneumoniae to attach to epithelial cells in the upper respiratory tract. (4)
The polysaccharide capsule interferes with phagocytosis through its chemical composition, resisting by interfering with binding of complement C3b to the cell's surface. Encapsulated strains of S. pneumoniae are found to be 100,000 times more virulent than unencapsulated strains during invasion of mucosal surfaces. (5) Virulence is a harmful quality possessed by microorganisms that can cause disease, and the characteristics of encapsulated strains are essentially the key to virulence. Being encapsulated is advantageous for bacteria, since the capsule is in most cases antiphagocytic, meaning it prevents phagocytosis. It is also antigenic as it stimulates the production of an antibody when introduced into the body. Encapsualted S.pneumoniae strains have caused large conjunctivitis outbreaks in schools, military training facilities, and at other locations worldwide (Valentino et al., 2014). However, recent studies of epidemiologically unrelated conjunctivitis cases found that most cases were caused by encapsulated strains.
Pili are long, thin extracellular organelles that are able to extend outside of the polysaccharide capsule. They are encoded by the rlrA islet (an area of a genome in which rapid mutation takes place) which is present in only some isolated strains of S. pneumoniae. These pili contribute to adherence and virulence, as well as increase the inflammatory response of the host. (4)
Application to Biotechnology
S. pneumoniae is noted as a major cause of infections and mortality in patients with upper and lower respiratory tract infections. Streptococcus pneumoniae is mainly found in the lab for the purpose of creating a vaccine against it. There are 90 different capsular types of S. pneumoniae, so a vaccine that is based on polysaccharide alone will not work. The most feasible vaccines against this bacteria work against different highly prevalent subgroups. In 1945, there were 4 targeted subgroups, which rose to 14 in the 1970's, and finally to the current vaccine that targets 23 subgroups. These 23 targeted types represent 85-90% of the virulent types, and the vaccine has an efficiency of around 60%. However, utilization of this vaccine remains very low considering the number of S. pneumoniae infections around the world. (5)
The study of pneumococcal proteins has been initiated to assess their possible role in future pneumococcal medications. This suggests that pneumococcal proteins can be used as a carrier protein in a vaccination or antibiotic by inducing protection against the remaining serotypes, preventing serotype replacement of S. pneumoniae. Carrier proteins have the potential to transport specific substances across the cell membrane by active transport.
Drug development could be an alternative approach to combat pneumonia in addition to vaccines. The use of gemifloxacin, which is a recent member of the quinolone class of antibacterial agents, was approved by the United States Food and Drug Administration for the treatment of community-acquired pneumonia and other bacterial damage (Morrissey and Tillotson, 2004). Gemifloxacin shows excellent activity in vitro, as well as in vivo against the respiratory pathogen S. pneumoniae because of its strong affinity for DNA gyrase and topoisomerase IV (the dual target action system) (Morrissey and Tillotson, 2004).
A different approach in regards to a class of compounds is the use of opsonin . Opsonin is an antibody that binds to foreign microorganisms or cells, making them more susceptible to phagocytosis. Some protection against pneumococcal infections is resolved by opsonin. This is thought to be among the major mechanisms associated with the immune system that protects the host against numerous pneumococcal infections (Vitharsson, 1994).
Another potential candidate is the bacterium Lactococcus lactis, which produces the S. pneumoniae surface protein A (PspA). Exposure to this antigen may stimulate antibody production effective against S. pneumoniae, offering an economical alternative to current options. (3) Due to a continuing increase in S. pneumoniae's antibiotic resistance, the search for a better vaccine is ongoing, and studies show that research on the Lactococcus lactis bacteria for use as a vaccine is promising. Its production of the pneumococcal surface protein PspA makes it a good candidate for a mucosal vaccine, which could be administered through the nose instead of an injection. Studies show that the lactococcal vaccine offers better protection against respiratory infection by S. pneumoniae than injections of similar amounts of recombinant PspA administered by injection. There is considerable potential to develop a vaccine with L. lactis, for use against S. streptococcus and more. (3)
Current Research
Streptococcus pneumoniae remains actively virulent in today's world with an increasing resistance to antibiotics. The genome sequences for a few different strains have just been mapped, including the capsulated, virulent strain D39 and the unencapsulated, avirulent strain R6. Comparisons of the different strains expose modern mutations and information about regulation, metabolism, and virulence in the bacteria. (2)
References
1. Hervé, T., et al. "Complete Genome Sequence of a Virulent Isolate of Streptococcus pneumoniae". Science. July 20, 2001. Volume 293. p. 498-506.
http://www.sciencemag.org/cgi/content/abstract/293/5529/498
2. Lanie, J., Ng, W., Kazmierczak, K., Andrzejewski, T., Davidsen, T., Wayne, K., Tettelin, H., Glass, J., Winkler, M. "Genome sequence of Avery's virulent serotype 2 strain D39 of Streptococcus pneumoniae and comparison with that of unencapsulated laboratory strain R6". Journal of Bacteriology. 2007. Volume 189. p. 38-51.
http://www.ncbi.nlm.nih.gov/sites/entrez?cmd=Retrieve&db=PubMed&list_uids=17041037&dopt=Abstract
3. Hanniffy, S., Carter, A., Hitchin, E., and Wells, J. "Mucosal Delivery of a Pneumococcal Vaccine Using Lactococcus lactis Affords Protection against Respiratory Infection". The Journal of Infectious Diseases. 2007. Volume 195. p. 185-193.
4. Barocchi, M., Ries, J., Zogaj, X., Hemsley, C., Albiger, B., Kanth, A., Dahlberg, S., Fernebro, J., Moschioni, M., Masignani, V., Hultenby, K., Taddei, A., Beiter, K., Wartha, F., von Euler, A., Covacci, A., Holden, D., Normark, S., Rappuoli, R., Henriques-Normark, B. "A pneumococcal pilus influences virulence and host inflammatory responses". Proceedings of the National Academy of Sciences of the United States of America. 2006. Volume 103. p. 2857-62.
http://www.ncbi.nlm.nih.gov/sites/entrez?cmd=Retrieve&db=pubmed&dopt=Abstract&list_uids=16481624
5. Todar, K. "Streptococcus pneumoniae: Pneumococcal pneumonia". Todar's Online Textbook of Bacteriology. 2003.
http://textbookofbacteriology.net/S.pneumoniae.html
6. Lysenko, E., Ratner, A., Nelson, A., Weiser, J. "The role of innate immune responses in the outcome of interspecies competition for colonization of mucosal surfaces". PLoS Pathogens. 2005. Volume 1. p. 1.
http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pubmed&pubmedid=16201010
7. Lederberg, J., Gotschlich, E. "A Path to Discovery: The Career of Maclyn McCarty". PLoS Biology. 2005. Volume 3. p. 341.
http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=1250295
8. Tleyjeh IM, Tlaygeh HM, Hejal R, Montori VM, Baddour LM. "The impact of penicillin resistance on short-term mortality in hospitalized adults with pneumococcal pneumonia: a systematic review and meta-analysis". 2006. Volume 42. p. 788-97.
http://www.ncbi.nlm.nih.gov/sites/entrez?cmd=retrieve&db=pubmed&list_uids=16477555&dopt=Abstract
Bogaert, D., de Groot, R., Hermans, P.W.M. 2004. Streptococcus pneumoniae colonisation: the key to pneumociccal disease. Lancet Infectious Diseases. 4:144-154.
Bogaert, D., Hermans, P.W.M., Adrian, P.V., Rumke, H.C., de Groot, R. 2004. Pneumococcal vaccines: an update on current strategies. Vaccines. 22: 2209-2220.
Granizo, J.J., Aguilar, L., Casal, J., Garcia-Rey, C., Dal-Re, R., Baquero, F. 2000. Streptococcus pneumoniae resistance to erythromycin and penicillin in relation to macrolide and β- lactam consumption in Spain (1979-1997). Journal of Antimicrobial Chemotherapy. 46: 767-773.
Huttner, B., Samore, M. 2011. Outpatient Antibiotic Use in the United States: time to “Get Smarter”. Clinical Infectious Diseases. 53(7): 540-543. Streptococcus pneumoniae. 2015. Centers for Disease Control and Prevention.
Van der Poll, T., Opal, S.M. 2009. Pathogenesis, treatment, and prevention of pneumococcal pneumonia. Lancet. 374:1543-1556.
Stanek, R.J., Maher, M.B., Norton, N.B., Mufson, M.A. 2011. Emergence of a Unique Penicillin-Resistant Streptococcus pneumoniae Serogroup 35 Strain. Journal of Clinical Microbiology. 49(1): 400-404
Morrissey, I., Tillotson, G. 2004. Activity of gemifloxacin against Streptococcus pneumoniae and Haemophilus influenzae. Journal of Antimicrobial Chemotherapy. 53: 144-148
Hoshino, K. et al. 2002. High Rate of Transmission of Penicillin-Resistant Streptococcus pneumoniae between Parents and Children. Journal of Clinical Microbiology. 40 (11): 4351-4359
Appelbaum, P.C. 2002. Resistance among Streptococcus pneumoniae: Implications for Drug Selection. Clinical Infectious Diseases. 34: 1613-1620
Valentino, M.D. et al. 2014. Unencapsulated Streptococcus pneumonia from conjunctivitis encode variant traits and belong to a distinct phylogenetic cluster. Nature Communications. 5:5411
Vitharsson G. et al. 1994. Opsonization and antibodies to capsular and cell wall polysaccharides of Streptococcus pneumoniae. J Infect Dis. 170(3):592-9.