Thalassiosira pseudonana
Classification
Eukaryota; Bacillariophyta; Coscinodiscophyceae; Thalassiosirales; Thalassiosiraceae [Others may be used. Use NCBI link to find]
pseudonana
|
NCBI: Taxonomy |
Thalassiosira pseudonana
Description and Significance
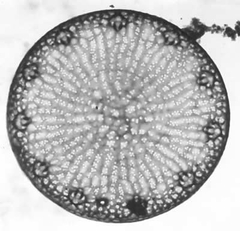
Thalassiosira pseudonana is a marine centric diatom. Diatoms are unicellular, eukaryotic, phytoplankton that display a unique evolutionary history and provide major ecological contributions in marine environments. Diatoms are capable of photosynthesis, having acquired plastids through secondary endosymbiosis of primary endosymbionts, including plants and, green algae, red algae, and glaucophytes. Having evolved 91.5 million years ago during the Upper Turonian period [1], analyses of these organisms display long-term contributions to deposits of diatomite, carbon cycling, global climate, and petroleum reserves. Today diatoms continue to have major ecological implications by playing a fundamental role in global carbon cycling and global climate. As a result of the ecological importance of diatoms, T. pseudonana was the first diatom to undergo full genome sequencing. In addition, T. pseudonana have elaborate silicified cell wall nanostructures that may contribute to future study of silica nanotechnology. T. pseudonana diatoms display a unique combination of metabolic processes including genes for nitrogen fixation, the urea cycle, carbon fixation, iron uptake, and photosynthesis [3].
Genome Structure
The genome of T. pseudonana was sequenced by E. Armbrust et. al using a whole-genome shotgun approach. The nuclear genome of T. pseudonana is 34.5 million bp, accounting for a predicted total of 11,242 protein-encoding genes and encodes 24 pairs of chromosomes, totaling 34.5 Mb. The plastid genome is 128,813 bp accounting for 144 protein-encoding genes, and the mitochondrial genome is 43,287 bp with 40 protein-coding genes. The most abundant domain in T. pseudonana is protein kinase 1 [3]. The high number of kinase-encoding domains throughout the genome suggests that phospho-relay-based signal pathways are commonly used by T. pseudonana [3]. T. pseudonana differ from many eukaryotic species and has a relatively low reliance on receptor kinases and leucine-rich receptor (LRR) containing receptors and that the major transcription factor is the heat-shock family.
T. pseudonana and all other species of diatoms, have evolved from previously non-photosynthetic eukaryotes that underwent secondary endosymbiosis by engulfing a photosynthetic eukaryote having previously evolved through primary endosymbiosis. Analysis of the T. pseudonana genome reveals that diatoms align with almost half of their closest homologs in plant, red algal, and animal genomes [3]. Specifically, the genome of T. pseudonana displays homology with 182 proteins of the red algae, C. merolae, 865 proteins of the plant, A. thaliana, and 806 proteins with the animal, M. musculus [3]. In addition, the genome displays homology with cyanobacteria and high levels of gene transfer between genomes during endosymbiosis establishment [3] Comparisons of nucleomorph genomes to the diatom nucleus show that there were multiple gene transfers from red algal plastid genomes to red algal nuclear genomes to diatom nuclear genomes [3]. The genome does not display any recent transfers from plastid or mitochondrial genomes to the host nuclear genome; however, a gene (psb28) encoding a photosytem II was found in both the plastid and nuclear genomes, suggesting a plastid-to-nucleus transfer in in progress [3].
Cell Structure, Metabolism and Life Cycle
The frustule, or cell wall, of T. pseudonana is composed of hydrated silicon dioxide and a small amount of organic matter [3]. The frustule of diatoms can withstand extreme force, in some cases displaying resistance up to 720 µN [4]. The rigidity of the frustules has resulted from an evolutionary co-arms race between diatoms and their predators, including copepods and euphausiids. These predators have silica-lined mandibles and gizzards lined with rows of sharp teeth used to crush open the hard frustule shells of diatoms [4]. Diatom frustules have evolved resistance to high external force as a result of the selective pressures posed by the architecture of their predators.
T. pseudonana is capable of metabolizing multiple forms of nitrogen. T. pseudonana contains multiple transporter genes for inorganic forms of nitrogen including, nitrate, ammonium, phosphate, sulfate, and salicylic acid [3]. T. pseudonana also contains genes that allow it to take up organic forms of nitrogen and to catabolize amino acids [3]. Not present in any other eukaryotic photoautotrophs, T. pseudonana contains all enzymes necessary for a complete urea cycle. The reason for carrying out this process is not fully understood, but T. pseudonana have been shown to carry out the urea cycle and direct the byproducts produced at various steps toward the initiation and mediation other pathways, such as the synthesis of long-chain polyamines required for frustule formation [3].
Carbon fixation is an important metabolic process carried out by T. pseudonana due to its role in global geological carbon cycling. Interestingly, genome analysis indicates that carbon fixation occurs in the cytoplasm of this organism, rather than in the plastid [3]. Both glycolysis and gluconeogenesis, are completely localized to the cytoplasm [3]. The mechanism by which carbon dioxide is transferred to Rubisco during the first step of carbon fixation remains unknown, but the decarboxylating enzymes necessary for the delivery of carbon dioxide to Rubisco are found in the cytoplasm [3].
T. pseudonana evolved photosynthesis capabilities through secondary endosymbiosis of a photosynthetic eukaryote. Analysis of the genome of T. pseudonana reveals that diatoms absorb red and blue light, but not green light [3]. These diatoms possess homologs for cryptochromes, which absorb blue light, and homologs for phytochrome, which absorbs red light [3]. Blue and red light are found most frequently at the water surface whereas green light penetrates deep within ocean water, suggesting that diatoms only encode photoreceptors for blue and red light to maintain close proximity to the water surface [3].
T. pseudonana have genes for a high-affinity iron-uptake system [3]. In total, these diatoms were shown to possess genes for two ferric reductases, a multi-copper oxidase, and two iron permeases, that can deliver Fe3+ to cells by reduction of ferrous iron. In addition, genome contains genes for metal homeostasis and metal detoxification [3].
Ecology
T. pseudonana is a diatom found in marine ecosystems, including oceans and freshwater [3]. Through the processes of carbon fixation and photosynthesis, diatoms play a major role in the flux of atomspheric carbon dioxide in oceans [3]. Diatoms contribute about 40% of organic carbon produced in the ocean each year (45 to 50 billion metric tons) and play a fundamental role in global carbon cycles [2]. The role of diatoms in global carbon cycling is so extreme that the following has been said about diatoms: "Their role in global carbon cycling is predicted to be comparable to that of all terrestrial rainforests combined" [3].
T. pseudonana control biogenic silica processing to a great extent, such that all silicon atoms entering the oceans are incorporated into diatom frustules 40 times before entering the ocean floor [3]. The process of generating and maintaining frustules controls biogenic silca cycling as diatoms take up silicic acid from sea water to build their frustules [6]. In 1995, an average of 6.1 ±2.0 teramoles of silicic acid were predicted to leak into the world's oceans each year [3]. Diatoms match this input by producing a predicted 7.1 ± 1.8 teramloes of biogenic silica each year [3]. Diatom uptake of silicic acid for frustule formation dramatically enhances cell growth preventing CaCO3 from being released to marine sediments [6]. Diatoms enhance their growth at a cost to coccolithophorids, which require CaCO3 for cell growth [6]. This limitation on coccolithophorid growth decreases the carbonate pump, ultimately increasing the levels of atmospheric CO2[6]. After diatom cell death biogenic silica that is not dissolved in ocean waters settles in the marine sediment, available for the carbonate pump [7].
References
1. Sinninghe Damste, Jaap S., Muyzer, Gerard, Abbas, Ben, Rampen, Sebastian W., Masse, Guillaume, Allard, W. Guy, Belt, Simon T., Robert, Jean-Michel, Rowland, Steven J., Moldowan, J. Michael, Barbanti, Silvana M., Fago, Frederick J., Denisevich, Peter, Dahl, Jeremey, Trindade, Luiz A.F., Schouten, Stefan. "The Rise of the Rhizosolenid Diatoms." Science. 23 April 2004. Vol. 304, pp. 584-587.
2.Nelson, DM, Treguer, P, Brezinski MA, Leynaert, A, Queginer, B. "Production and Dissolution of Biogenic Silica in the Ocean - Revised Global Estimates, Comparsion with Regional Data and Relationship to Biogenic Sedimentation." Global Biogeochemical Cycles. September 1995. Vol. 9, pp. 359-372.
3. Armbrust, E. Virginia, Berges, John A., Bowler, Chris, Green, Beverly R., Martinez, Diego, Putnam, Nicholas H., Zhou, Shigou, Allen, Andrew E., Apt, Kirk E., Brzezinski, Mark A., Chaal, Balbir K., Chiovitti, Anthony, Davis, Aubrey K., Demarest, Mark S., Detter, J. Chris, Glavina, Tijana, Kapitonov, Vladimir V., Kroger, Nils, Lau, Winnie W.Y., Lane, Todd W., Larimer, Frank W., Lippmeier, J. Casey, Lucas, Susan, Medina, Monica, Montsant, Anton, Obornik, Miroslav, Parker, Micaela Schnitzler, Palenik, Brian, Pazour, Gregory J., Richardson, Paul M., Rynearson, Tatiana A., Saito, Mak A., Schwartz, David C., Thamatrakoln, Kimberlee, Valentine Klaus, Vardi, Assaf, Wilkerson, Frances P., and Rokhsar, Daniel S. "The Genome of the Diatom Thalassiosira pseudonana: Ecology, Evolution, and Metabolism." Science. 4 October 2004. Vol. 306, pp. 79-86.
4. Hamm, C.E., Merkel, R., Springer, O., Jurkojc, P., Maier, C., Smetacek, V. "Architecture and material properties of diatom shells provide effective mechanical protection." Nature. Vol. 421, pp. 841-843.
5. Paasche, E. "Silicon and the ecology of marine plankton diatoms. I. Thalassiosira pseudonana (Cyclotella nana) grown in a chemostat with silicate as limiting nutrient." Marine Biology. Vol. 19, pp. 117-126.
6. Matsumoto, Katsumi, Sarmiento, Jorge L., Brezinski, Mark A. "Silicic acid leakage from the Southern Ocean: A possible explanation for glacial atmospheric pCO2." Global Biogeochemical Cycles. September 2002. Vol. 16, pp. 51-56.
7. Treguer, Paul, Nelson, David M., Van Bennekom, Aleido, J., DeMaster, David J., Leynaert, Aude, Queginer, Bernard. "The Silica Balance in the World Ocean: A Reestimate." Science.' 21 April 1995. Vol. 268, pp. 375-379.
Author
Page authored by Kayla Mitman, student of Prof. Jay Lennon at Indiana University.
