The Threat of Airborne Transmission of Influenza A: H5N1

Introduction
By Morgan O'Connor
The evolution of the influenza virus has long been a threat to our society. From the first major pandemic in 1918 involving the H1N1 "Spanish flu," which killed an estimated 50 million people and infected about 500 million people worldwide to the Swine flu in 2009 that affected one in six Americans and caused a reported 43-89 million cases and 8870-18300 deaths worldwide, viral re-assortment and mutation have raised a serious concern in public health [1,2]. Globally, we are constantly trying to combat and prevent viral infection. With viruses everywhere, continuously undergoing re-assortment and rapid evolution, it is important to try to understand and predict the next outbreak. The mode of transmission should be taken into account with degree of severity, as the right combination can produce a severe problem for our society today: an easily transmittable (airborne) fatal virus could spontaneously emerge which renders our immune systems and modern medications defenseless.
Background
The influenza viruses are RNA viruses which are classified as genera of the family Orthomyxoviridae [3]. The three genera include influenza A, B and C. A distant relative of the parainfluenza viruses belonging to the Paramyxovirus family, influenza most commonly affects human respiratory tracts [3].
Among all three genera, influenza A is known to cause the most severe symptoms and be the most pathogenic to humans [4]. Influenza A originated from the natural hosts wild aquatic birds [4]. The disease was transmitted from birds to humans through handling or ingesting poultry. The H5N1 strain is the most well known avian strain, known as the "highly pathogenic avian influenza virus" (HPAI) [4]. It has been classified as a threat to humans and other animals, both a pandemic and panzootic.
When transmitted from one species to another or from reservoir to another host, for example, from birds to humans, influenza outbreaks can be dangerous [4]. Some of the serotypes which are known to have caused the most severe pandemics include: H1N1 which was the cause of the Spanish flu in 1918 (Fig 3) and Swine flu in 2009, H2N2 which was the source of the Avian flu in 1957 that was widespread in Asia [4]; H3N2 was the cause of the Hong Kong flu in 1968 and H5N1 which produced the Bird flu more recently in 2004 [4].

Influenza B is most often known to be pathogenic among humans, seals and ferrets but is much less common than influenza A [5]. A key reason why influenza B exhibits less infection than influenza A is due to its lack of mutation, as its mutation rate is about three times less than that of type A; this reduces the amount of heterogeneity of the virus [5]. Immune systems are therefore able to combat and produce effective defenses against type B, such as antibodies, from an early age that withstand throughout lifetime.Influenza C virus infects humans, dogs, and pigs [6]. It is much less common than types A and B. It is most often seen in young children, as their immune systems are not yet fully developed [6].
Currently, two FDA-approved antiviral medications exist known as Oseltamivir (Tamiflu) and Zanamivir (Relenza) [7]. Oseltamivir and Zanamivir both utilize the same chemical mechanisms in order to prevent influenza spread [7]. By inhibiting neuraminidase (NA), a surface protein, these antivirals are both effective against types A and B inhibiting the viral assembly process (Fig 4) [7]. Reported resistance to these treatments has thus so far been low [7]. However, researchers are aware that this number could easily increase, and are therefore trying to predict new mutations that could arise to help in search of new mechanisms for treatment and prevention. Resistance has also been reported to occur throughout treatment and even well after the final course of treatment [7].
Vaccinations for influenza are also available and are modified each season corresponding to the strain and serotype that has emerged most recently [8]. Forms of the vaccination made available are both injection and nasal spray [8]. The influenza vaccine usually consists of antigens from a couple different serotypes, and are always both from influenza types A and B [8]. The United States Center for Disease Control and Prevention (CDC) advises for everyone over the age of 6 months to receive a vaccination at the start of each flu season, November-March [8]. People who are most susceptible to infection and therefore most advised to receive the vaccine include those over the age of 60, living with serious illnesses and those who are exposed to children and illness, such as hospital workers [8].
Structure and Composition
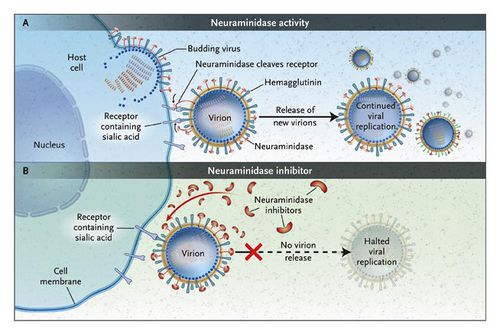
The size of the influenza particle is about 80-120 nanometers in diameter [9]. The structure is typically sphere-like however variations due to filamentous structures can form, most commonly observed in type C [9].There is a viral envelope which encases the central core, housing and protecting the RNA genome [9]. The envelope is made up of two types of glycoproteins, HA, hemagglutinin, and NA, neuraminidase (Fig 4) [9]. HA is a lectin or a carbohydrate protein that mediates the binding and entry processes of RNA into the host target cells [9]. Alternatively, NA is more involved in the release of the progeny assembled viruses from host infected cells (Fig 4) [9]. Mechanistically, NA cleaves the sugars which keep the mature viral particles attached to the host cell [9]. The HA and NA proteins are responsible for the type of antibody responses in the host cell and are also easy targets for inhibition of the virus used in antiviral drugs [9]. Influenza A viruses are classified into serotypes based on the forms of HA and NA and ultimately their antibody responses, for example H1N1. There are 16 different HA and 9 different NA [9].
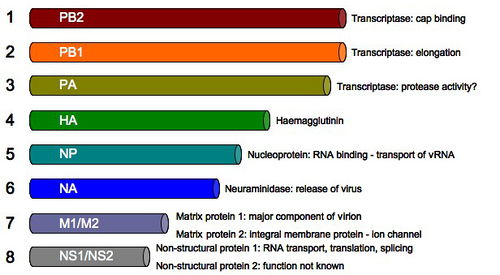
The genome of the influenza virus consists of 8 pieces of segmented negative sense RNA (Fig 5). Two strands encoding for the surface proteins: hemagglutinin (HA), neuraminidase (NA). PB2, the polymerase basic 2, encodes for the transcriptase involved in cap binding while PB1, the polymerase basic 1, is responsible for the transcriptase that facilitates elongation [9]. The PA encodes for a transcriptase for protease activity. The NP, nucleoprotein, is involved in the RNA binding and transport. M1/M2, or the matrix proteins 1 and 2, are involved in producing the vital proteins in influenza such as ion channels (Fig 5) [9]. The NS1 is involved in RNA transport, translation and splicing while the NS2 encodes for unknown functions [18].
The sizes of the strands also vary in length. In descending order the strand sizes are: PB2, PB1, PA, HA, NP, NA, M1/M2 and NS1/NS2 (Fig 5) [9]. For a virion to be complete, it must package all 8 RNA strand components. Therefore, there are many viruses that are assembled which are not viable.
Replication
The influenza virus, like all viruses, replicate within their host cells; the replication process begins as the virus binds to the cell through its HA surface proteins and the sialic acid sugars found on the surface of host epithelial cells (Fig 5) [10]. This binding typically occurs within the throat, nasal and lung regions in mammals, also within bird intestines [10]. A protease then cleaves HA as the virus is endocytosed into the cell, attaching to the vacuole membrane (Fig 5) [10]. Acidic conditions allow for M2 ion channels to open up within the membrane of the virus, as protons penetrate and acidify the core of the virus, releasing the vRNA and core proteins [10]. In amantadine drugs, another common antiviral drug, this M2 ion channel is blocked [10]. In the second stage of infection, the vRNA molecules, accessory proteins and RNA-dependent RNA polymerase enter freely into the cytoplasm and form a complex that is transported to the nucleus [10]. Within the nucleus, the RNA-dependent RNA polymerase transcribes a complementary positive sense vRNA; this vRNA can then follow either of two pathways, as it can be exported to the cytoplasm and translated or can remain in the nucleus (Fig 5) [10].
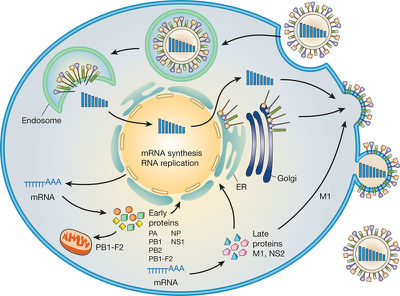
The newly synthesized viral proteins are secreted through the Golgi apparatus, while some are transported to the surface, i.e. the NA and HA surface proteins [10]; proteins can also be transported back to the nucleus to synthesize a new viral genome (Fig 5) [10].
For virion assembly to occur, synthesis of the negative sense vRNAs, which alone forms the genome of future viruses, as well as the RNA-dependent RNA polymerase and other viral proteins must be completed [10]. Once all parts are synthesized, the HA and NA molecules come together in the host cell membrane [10]. The vRNA along with the other viral core proteins exit the nucleus and enter the same membrane area as HA and NA, forming a protrusion ready to exit the cell (Fig 5) [10]. A mature viruses then bud from the cell, encased in a sphere of phospholipid membrane, obtaining the HA and NA in the surface membrane layer [10]. The viruses remain attached to the host cell membrane and the mature viruses finally separate as the NA cleaves sialic acid residues (Fig 5) [10]. After release of the fully formed viruses, the host cell ultimately dies [10].
The high mutation rate of the influenza virus which is the cause of the emergence of new strains leading to new pandemics, is largely due to the absence of an RNA proofreading enzyme [11]. This absence causes the RNA-dependent RNA polymerase that replicates the viral genome to make an error every ten thousand nucleotides, which is about the length of the vRNA [11]. This glitch causes a large majority of influenza viruses to be mutants. This creates large antigenic drift, or a change of antigens on the surface of the virus. The make-up of the genome which includes eight distinct segments of RNA also allows for re-assortment of RNAs with other viruses, when more than one type of influenza virus infects a single cell [11]. This type of re-assortment of RNA allows for antigenic shift which is defined as the shift from one antigen to a different one [11].
Transmission
Influenza has three main modes of transmission: direct, indirect and airborne. Direct transmission occurs when infected mucus through nose, eyes or throat, is passed into those of others. This involves direct contact of mucus membranes. Airborne transmission is produced through the respiratory tract, as aersols of an infected person are inhaled by a non-infected person. Liquid droplets small enough for inhalation are about 0.5-5μm in diameter [12]. On average, one sneeze can produce up to 40,000 droplets, the majority of which are too big for airborne transmission to occur [13]. The final mode of transmission is through indirect transmission, as transmission occurs through surface contact. Examples of common indirect contact include: skin to skin and skin to contaminated surface. Influenza can survive from up to 24-48 hours on sturdy non-porous surfaces [14]. However, on paper dry surfaces, it can only survive up to 15 minutes [14]. On skin, the virus can last up to 5 minutes [14]. In mucus, the viruses are viable up to 17 days, making direct contact with mucosal membranes the most dangerous source of infection [12]. Humidity and UV radiation are significant environmental factors that contribute to the transmission frequency, as low humidity and lack of sunlight in the winter facilitate viral survival [12].
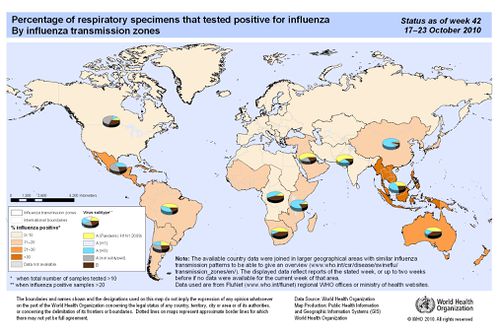
The shedding time of the influenza virus, or the duration of time in which an infected person might be infectious to others, begins about 24 hours before symptoms become overt [15]. The virus is released between 5 to 7 days, however cases may vary. Peak of infection is most often observed between day 2 and 3 [15]. Shedding time of the virus is also correlated with temperature of the host environment as hosts exhibiting fevers usually have higher transmission rates [15]. Children are also much more infectious than adults, as shedding time can last up to two weeks after initial infection [12]. These are all important elements when taking into consideration how influenza might affect specific as well as general populations at a given time; mathematical models have been generated to predict effects [12]. There is a wide range of transmission rates exhibited in various regions of the world, with different cultures, climates and health care all contributing to differences observed (Fig 6).
Evolution of the Influenza A virus
Evolution through small base change mutations and re-assortment events of the vRNA strands between human, swine and avian viruses have been observed and recorded throughout history of pandemics [16] (Fig 7). The dangers of re-assortment lie in the fact that each new sub-type produces a novel antigen to the immune system. The 1918 avian pandemic was caused by a mutation in avian viruses producing an adaptation to replicate in humans [16]. In 1957, the H2N2 was a product of re-assortment between avian and human viruses, with a change in the PB1, HA and NA segments [16]. The 1968 H3N2 was a re-assortment of again avian and human viruses with PB1 and HA, along with another avian virus in the NA [16]. The 1977 H1N1 Russian flu was basically a re-emergence of the H1N1 flu in 1918 [16]. The most recent 2009 H1N1 was a re-assortment of avian, swine and human RNA strands with PB1, HA, NP, NA, M and NS [16] (Fig 7).
Current Research
A study conducted by Fouchier et al. (2012), found that as few as 5 mutations in WT H5N1 are needed in order to create a fatal airborne transmission between ferrets, which have similar respiratory systems to humans (Fig 8) [5]. In a study conducted by Kawaoka et al. (2012), researchers instead produced an airborne H5N1 by simulating co-infection and re-assortment creating a hybrid with the H1N1 swine flu and then observed only 4 mutations were needed for virus to become airborne [17]. Although airborne transmission occurred in Kawaoka’s study, none of the ferrets in the study ultimately died, whereas all ferrets died in Fouchier’s study. In an extended study (Smith et al., 2012), researchers looked at data to generate a mathematical model for predicting the emergence of airborne H5N1 strain and found the mutations described in Kawaoka and Fouchier to already present in nature; more specifically, some strains of H5N1 already contain one or two of the mutations found in Kawaoka and in Fouchier and are therefore as little as up to only three mutations away from becoming airborne [18].

There has been a lot of controversy as to whether or not to release the information on the specific mutations that need to occur to transform H5N1 into an easily transmittable airborne disease. As recently as December 2011, the United States National Science Advisory Board for Biosecurity (NSABB) advised against releasing the scientific research, due to the potential threat of biological terrorism [19]. However, many scientists opposed withholding information stating that it would be more beneficial for the progression of research to share the data than to withhold for any fear of terrorism [20]. Altogether, this research is key not only in predicting and possibly preventing the next pandemic but also calls into question the idea of scientific censorship that is becoming increasingly more relevant in modern research and technology [19].
In February 2012, researchers and public health officials finally decided to fully disclose all research, publishing both studies [21]. Research was subsequently put on hold in order to allow information to disseminate and appease any growing concerns. At the end of March, the NSABB revised the papers. The methods and results were not altered, however in Fouchier’s paper, the overall biosafety was extended upon [21]. To date, the WHO has reported thus far a 60% fatality rate on H5N1 [20]. However, there have only been about 600 cases reported to hospitals, where symptoms are already severe enough to receive medical attention [20]. Though there has only been a small amount of incidence of H5N1, it is priority in research to predict any future behavior. Derek J. Smith, mathematician in modeling study of H5N1 outbreak of University of Cambridge stated: "We now know that we're living on a fault line. [Influenza] could do something. What we need to know is how likely is that is,” in Science press [20]. Fouchier was named Time magazine’s top 100 most influential people in the world in 2012 due to his groundbreaking research [20].
In one study conducted by Herfst et al. (2012) researchers sought to obtain an airborne transmissible A/H5N1 virus through ferrets and did so through first selecting a mutant A/H5N1 virus with specificity for α -2, 6-linked SA receptors; The specificity for α -2, 6- indicates a shift from avian intestine α -2, 3-linked SA receptors to human and ferret respiratory tracts [22]. Methodically, QuickChange multisite-directed mutagenesis kit was used to introduce the amino acid substitutions: N182K/Q222L/G224S in the HA of wild-type (WT) A/Indonesia/5/2005, resulting in A/H5N1HA N182K, A/H5N1HA Q222L, G224S and A/H5N1 HA N182K, Q222L, G224S [22].
In the first experiment, the three mutant viruses and A/H5N1 WT were inoculated into six ferrets for each group intra-nasally. To quantify shedding, throat and nasal swabs were collected daily and virus titers were quantified utilizing Madin Darby Canine Kidney (MDCK) cells [22]. Titers were collected after day 3 and day 7 [22]. Throughout the experiment, the WT inoculated viruses produced up to 10 times more titers than the highest producing mutant A/H5N1 HA Q222L, G224S, all after day 7; there were no significant differences after day 3 [22].
In the next experiment conducted in this study, another amino acid substitution, E627K, was made in the PB2 in A/Indonesia/5/2005 to increase specificity of host to humans, producing a recombinant A/H5N1 HA Q222L, G224S E627K [22]. There was not a significant difference in virus shedding with this additional amino acid substitution, both producing up to 1 x 104 50% tissue culture infectious doses (TCID50) [22].
Experimental set up involving the ferrets was then put into place. Four ferrets were housed into cages adjacent to those with four inoculated ferrets to test for airborne transmission [21]. The A/H5N1 HA Q222L, G224S E627K initially was not found to be airborne. To promote selection for an H5N1 that infects the ferret respiratory tract with the E627K mutation, one ferret was inoculated intra-nasally with WT and one with A/H5N1 HA Q222L, G224S E627K [21]. Throat and nose swabs were collected daily until day 4; nasal turbinates were used to inoculate the next ferret intra-nasally with passage 2 [22].
Procedure was done repeatedly until passage 6; at passage 6, sneezing was induced in the ferrets [22]. Sneeze samples were used for intra-nasal inoculation of ferrets for the passages of 7 through 10 (Fig 9). Ten passages were conducted to ensure sufficient adaptation of the virus in ferret respiratory system (Fig 9). WT virus turbinates exhibited a small range of titers from 1x105 to 1x107 TCID50/gram tissue throughout the course of 10 serial passages [22]. The ferrets inoculated with A/H5N1 HA Q222L, G224S E627K showed a slight increase in observed virus titers as passage number increased; the range was spread from 1x104 TCID50/gram tissue at the beginning of the experiment to about 3.2 x 105 to 1x106 TCID50/gram tissue in passage 10 [22]. Additionally, the nose swabs also increased throughout with peak shedding at passage 10. This suggests that A/H5N1 HA Q222L, G224S E627K had a greater amount of replication after repeated passages [22].
Sequencing of the viral quasi-species, or the mixture of viruses with a variety of mutations, was performed in the nasal washes after 10 passages of both of the WT and A/H5N1 HA Q222L, G224S E627K [22]. Mutations in the A/H5N1 HA Q222L, G224S E627K remained present in population throughout all passages [22]. The only additional amino acid substitution found in both WT and A/H5N1 HA Q222L, G224S E627K was T156A in HA, which removes a potential N-linked glycosylation site in HA [21]. This mutation was found in 99.6% of viruses in WT in 89% of the A/H5N1 HA Q222L, G224S E627K.The remaining 11% showed substitutions in the N154K which essentially has the same function as T156A [22].
In their final experiment researchers in this study wanted to test for airborne transmissible viruses present after passage 10 in the ferrets in both WT and A/H5N1 HA Q222L, G224S E627K [22]. They did this through obtaining nasal washes collected from ferrets at passage 10 which were then used to inoculate six ferrets intra-nasally, two with WT and four with A/H5N1 HA Q222L, G224S E627K (Fig 9). These inoculated ferrets were each placed into a cage connecting airways to an un-inoculated ferret placed in a neighboring cage (Fig 8). These cages allowed for airflow but no direct contact. Three out of four un-inoculated ferrets that shared airways with the infected A/H5N1 HA Q222L, G224S E627K ferrets were all found to become infected through aerosols though at lower levels than reported in Fouchier et al. (2012), and when compared to H1N1 2009 (Fig 10) [22]. Ferret 2 of the airborne infected A/H5N1 HA Q222L, G224S E627K contained highest amount of virus titers among ferrets and was used to inoculate and test once more for airborne transmission with 6 other ferrets, which confirmed results [22].
The genome sequences of viruses with airborne transmission all had Q22L, G224S and E627K mutations, only two additional amino acid substitutions both in HA [22]. H103Y which contributes to the formation of part of the HA trimer interface, and the previously mentioned T156A [22]. There were also many other mutations but none of which were consistently present throughout airborne mutant A/H5N1 populations [22].
Overall, this data suggests that there are only a few mutations that need to occur in nature in order for H5N1 to cause a new pandemic and through co-infection and re-assortment with other viruses, the diversity of viruses is rapidly increasing. Steps for prevention and treatment are currently being researched and the promise for new antiviral treatments is increasing as this information is becoming more widespread.
References
3) Urban, Marguerite A. MD. 2009. Influenza. The Merck Manual Home and Health Handbook.
8) Center for Disease Control and Prevention. 2020. Key Facts About Seasonal Flu Vaccine.
10) Smith AE, Helenius A. 2004. How viruses enter animal cells. Science 304 (5668): 237–42.
17) Kawaoka, Yoshihiro. 2012. H5N1: Flu transmission work is urgent. Nature. 482:155.
19) NIH. 2012. National Science Advisory Board for Biosecurity Findings and Recommendations.
25) Vainionpää R, Hyypiä T. 1994. Biology of parainfluenza viruses. Clin. Microbiol. Rev. 7 (2): 265–75.
Edited by student of Joan Slonczewski for BIOL 238 Microbiology, 2013, Kenyon College.
